Impact Factor
ISSN: 1449-1907
Int J Med Sci 2021; 18(14):3150-3157. doi:10.7150/ijms.61771 This issue Cite
Research Paper
PD-L1 correlated gene expression profiles and tumor infiltrating lymphocytes in pancreatic cancer
1. Department of Nuclear Medicine, Renji Hospital, School of Medicine, Shanghai Jiao Tong University, Shanghai 200127, China.
2. Institute of Clinical Nuclear Medicine, Renji Hospital, School of Medicine, Shanghai Jiao Tong University, Shanghai 200127, China.
3. Department of Cardiology, Gongli Hospital of Shanghai Pudong New Area, Shanghai 200135, China.
4. Department of Bioinformatics and Biostatistics, School of Life Science and Technology, Shanghai Jiao Tong University, Shanghai 200240, China.
5. Shanghai Key Laboratory of Molecular Imaging, Shanghai University of Medicine and Health Sciences, Shanghai 201318, China.
*These authors contributed equally to this work.
Abstract
Objective: To study the expression and clinical value of PD-L1 gene in pancreatic cancer, and to predict the role of PD-L1 gene in the development of pancreatic cancer.
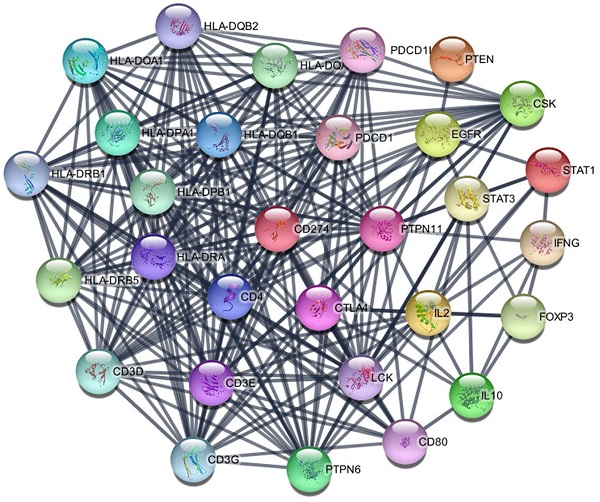
Methods: The pancreatic cancer datasets were downloaded from the Cancer Genome Atlas (TCGA) and the Oncomine to obtain the PD-L1 gene expression profile and clinical information. Bioinformatics methods were used to analyze the correlation between the expression level of PD-L1 gene in pancreatic cancer and clinicopathological indicators, as well as its influence on prognosis. GSEA and WGCNA analysis was performed to predict the possible pathways of PD-L1 gene regulation in pancreatic cancer. TIMER and MCP-counter were used for PD-L1 with immune infiltration. The genes interact with PD-L1 were also investigated by STING and immunoco-precipitation combined with mass spectrometry analysis (IP-MS).
Results: In TCGA database, the overall survival of patients with high expression of PD-L1 gene was significantly lower than that of patients with low expression of PD-L1 gene (χ2 = 12.52, P < 0.001). The samples with high expression of PD-L1 gene showed enrichment of 8 pathways including toll-like receptor signaling pathway and NOD receptor signaling pathway (P < 0.01, FDR < 0.05). Immune infiltration analysis suggested that PD-L1 were associated with monocytic lineage (r = 0.5). The proteins interacting with PD-L1 are mainly concentrated in RNA binding, ribosome, spliceosome and other biological processes or pathways.
Conclusion: PD-L1 gene may play an important role in the development of pancreatic cancer and is expected to be a prognostic indicator of pancreatic cancer.
Keywords: PD-L1, Prognostic value, Pancreatic cancer, TCGA, GSEA

 Global reach, higher impact
Global reach, higher impact