Impact Factor ISSN: 1449-1907
Int J Med Sci 2021; 18(5):1297-1311. doi:10.7150/ijms.53531 This issue Cite
Research Paper
Identification novel prognostic signatures for Head and Neck Squamous Cell Carcinoma based on ceRNA network construction and immune infiltration analysis
1. Department of Oncology, Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan, Hubei, 430030, P.R. China.
2. Department of Orthopedics, Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan, Hubei, 430030, P.R. China.
Abstract
Background: Head and neck squamous cell carcinoma (HNSCC) is a common malignancy with high mortality and morbidity worldwide, but the underlying biological mechanisms of molecules and tumor infiltrating-immune cells (TIICs) are still unknown.
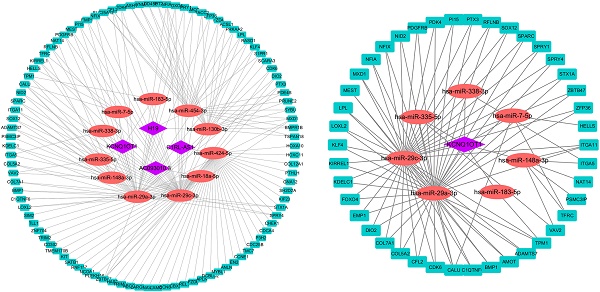
Methods and Results: We obtained mRNAs, lncRNAs, and miRNAs expression profiles of 546 HNSCC from The Cancer Genome Atlas (TCGA) database to develop a ceRNA network. CIBERSORT was employed to estimate the fraction of 22 types of TIICs in HNSCC. Univariate and multivariate Cox regression and lasso regression analyses were used to develop prognostic signatures. Then, two novel risk signatures were constructed respectively based on six ceRNAs (ANLN, KIT, PRKAA2, NFIA, PTX3 and has-miR-148a-3p) and three immune cells (naïve B cells, regulatory T cells and Neutrophils). Kaplan-Meier (K-M) analysis and Cox regression analysis further proved that these two signatures were significant prognostic factors independent of multiple clinicopathological characteristics. Two nomograms were built based on ceRNAs-riskScore and TIICs-riskScore that could be used to predict the prognosis of HNSCC. Co-expression analysis showed significant correlations between miR-148a-3p and naive B cells, naive B cells and plasmas cells.
Conclusion: Through construction of the ceRNA network and estimation of TIICs, we established two risk signatures and their nomograms with excellent utility, which indicated the potential molecular and cellular mechanisms, and predicted the prognosis of HNSCC.
Keywords: head and neck squamous carcinoma, ceRNA network, immune infiltration, prognostic signature, nomogram

 Global reach, higher impact
Global reach, higher impact