Impact Factor ISSN: 1449-1907
Int J Med Sci 2020; 17(11):1508-1514. doi:10.7150/ijms.41812 This issue Cite
Research Paper
Analyzing Functional Pathways and constructing gene-gene network for Narcolepsy based on candidate genes
1. Department of Neuromedicine, Peking University People's Hospital, Beijing, China.
2. Department of Epidemiology and Biostatistics, School of Public Health, Peking University Health Science Center, Beijing, China.
Received 2019-11-5; Accepted 2020-5-15; Published 2020-6-15
Abstract
Aims: To investigate the interactions among narcolepsy-associated genes and reveal the pathways these genes involved through bioinformatics analyses.
Methods: The study was performed with the following steps: 1) Selected the previously discovered narcolepsy risk genes through literature review, 2) pathway enrichment analysis, and construction of gene-gene and protein-protein interaction (PPI) networks for narcolepsy.
Results: 1) GO analysis revealed the positive regulation of interferon-gamma production as the most enriched terms in biological process, and C-C chemokine receptor activity as the most enriched term in molecular function, 2) KEGG pathway enrichment analysis revealed selective enrichment of genes in cytokine-cytokine receptor interaction signaling pathways, and 3) five hub genes were identified (IFNAR1, IL10RB, DNMT1, TNFSF4 and NFATC2).
Conclusion: The bioinformatics results provide new insights into the molecular pathogenesis of narcolepsy and the identification of potential therapeutic targets for narcolepsy treatment.
Keywords: single-nucleotide polymorphisms, candidate gene, hypothalamic neurons, neurodegeneration, neurology
Introduction
There are two challenges in the management of narcolepsy: 1) the mechanisms of the onset and development of narcolepsy are unclear and 2) the current therapies for narcolepsy are limited, being mostly symptom-based.
Recently, the genetic risk of narcolepsy was evaluated based on the presence of HLA-DQB1*06:02, which represents an important but imperfect predictor of narcolepsy [1, 2], In addition to HLA-DQB1*06:02, many other genes have been identified as narcolepsy risk genes in international SNP-based GWASs. However, the effects of single SNPs identified by GWAS are usually small and of limited clinical significance, for both evaluating the narcolepsy risk and revealing the relevant mechanisms. The interactions among genes have not been investigated yet. Several studies investigated the benefits of conflating genomic risk estimates obtained from SNP genotyping into a genetic risk score (GRS) to predict the risk of diseases [3-7]. In addition, gene networks have been studied widely for illustrating the interactions among genes [8] and the inference of biological mechanisms [9].
The pathogenesis of narcolepsy is an urgent and needs to be revealed. However, since narcolepsy is a rare disease, related GWAS research is limited, and other data are even more scarce and difficult to obtain. Because of the incomplete data, we are not able to do deep analysis of data as that in other common diseases such as stroke, diabetes and so on. However, we still need to build prediction tools and explore the pathophysiological mechanisms of narcolepsy. In order to complement the GWASs and uncover the “missing heritability”, we selected the SNPs in recently reported narcolepsy susceptibility loci (including previously reported variants in European and Asian ancestry in case-control studies), and investigated the interactions among the candidate genes, followed by pathway enrichment analysis and construction of gene networks for narcolepsy, to reveal the potential pathways and mechanisms associated with narcolepsy.
Methods
Genetic loci selection
We selected previously discovered narcolepsy risk loci from GWAS studies published before September, 2019 [11-20]. Both the narcolepsy susceptibility loci identified in Chinese population and European population were selected. The final list of candidate genes is shown in Table 1.
The selection of SNPs
| Chr | Gene | SNP | Tag SNP | Allele | Risk Allele |
|---|---|---|---|---|---|
| 1 | TNFSF4 | rs7553711 | rs4090391 | C/T | T |
| 1 | MIR-552 | rs10915020 | rs12026171 | A/G | G |
| 3 | CCR1 | rs3181077 | - | C/T | C |
| 3 | CCR3 | rs3181077 | - | C/T | C |
| 4 | BTBD9 | rs3923809 | - | A/G | G |
| 7 | TRB | rs2854536 | - | T/C | T |
| 8 | UBXN2B | rs2859998 | rs6993992 | T/C | T |
| 10 | ZNF365 | rs10995245 | - | G/A | A |
| 10 | A1CF | rs4290173 | rs16911668 | A/G | A |
| 12 | TEAD4 | rs12425451 | rs12322530 | G/A | G |
| 14 | TRA | rs1154155 | - | T/G | T |
| 14 | rs12587781 | rs1154153 | T/C | T | |
| 14 | rs1263646 | rs1263647 | A/G | A | |
| 15 | CTSH | rs12148472 | - | T/C | C |
| 15 | rs3825932 | - | T/C | T | |
| 19 | DNMT1 | rs2290684 | rs2114724 | C/T | T |
| 19 | rs6511570 | rs7253062 | G/A | G | |
| 20 | NFATC2 | rs8119787 | - | A/G | G |
| 21 | IL10RB | rs2834118 | rs2834113 | T/G | G |
| 21 | IFNAR1 | rs2252931 | rs2834188 | A/G | A |
| 22 | CPT1B | rs5770917 | rs5770911 | C/T | C |
| 22 | CHKB | rs5770917 | rs5770911 | C/T | C |
Statistical Analyses
Go term and KEGG pathway analysis
The online software, Database for Annotation, Visualization and Integrated Discovery (DAVID, https://david.ncifcrf.gov/) 6.8 knowledgebase [22] and Retrieval Interacting genes (STRING) 10.0 platform (http://string-db.org/) [23], were used to perform Gene Ontology (GO) analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis, with a threshold of p value less than 0.05. The top 10 enrichment GO terms and KEGG pathway annotations in our study were listed. The histograms were plotted using MS Excel 2007.
Construction of gene and PPI networks, and hub gene analyses
The Search Tool for the Retrieval of Interacting genes (STRING) 10.0 platform (http://string-db.org/), an online tool for the structural and functional analysis of protein interactions [23], was used to obtain the interactive relationships among the candidate genes, which were further constructed using Cytoscape software 3.7.1 (http://www.cytoscape.org/) [24]. The plugin cytoHubba [25] was used to select the hub genes from the PPI network. As the number of the candidate genes was limited, the cut-off criteria included a combined score < 0.2 and a node degree of > 7 for the screening of hub genes from the candidate genes. The Molecular Complex Detection (MCODE; version 1.31) app in Cytoscape was used to analyze and identify PPI network modules. Among the selected genes, the gene-gene interaction network comprising the narcolepsy-genes as nodes was constructed.
Data availability
All experimental data within the article and its supplementary information are available from the corresponding author upon reasonable request.
Results
The narcolepsy-associated genes reported in previous researches
Eighteen previously reported narcolepsy-associated genes were selected, both the narcolepsy susceptibility loci identified in Chinese population and European population were selected. The candidate genes were shown in Table 1.
GO biological process analysis and KEGG pathway enrichment of the candidate genes
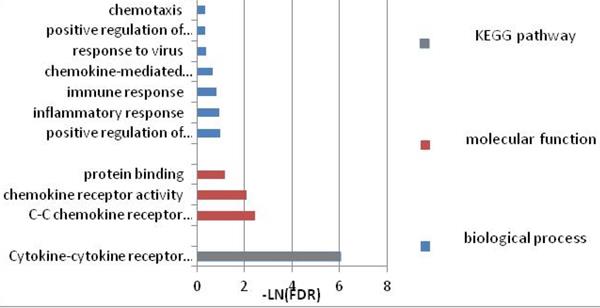
To further investigate the function of these candidate genes, we uploaded these 16 genes into DAVID and STRING for GO and KEGG pathway analyses. The results indicated that the genes were mainly enriched in the pathways associated with immune response. GO biological process analysis found that in terms of molecular function (MF), the genes were associated mainly with C-C chemokine receptor activity, C-C chemokine receptor activity, chemokine receptor activity, and protein binding. In terms of biological process (BP), the genes were associated mainly with positive regulation of interferon-gamma production, inflammatory response, and immune response. KEGG pathway analysis indicated that the relevant genes were enriched in Cytokine-cytokine receptor interaction signaling pathways. Within each of the functional groups, the enrichment terms for the candidate genes with p-value < 0.05 are listed in Table 2. The KEGG pathways and gene-gene interaction network are depicted in Figure 1 and Figure 2.
Signaling pathways enriched by the candidate genes
| Category | Term | Gene count | Background number | p-value | FDR |
|---|---|---|---|---|---|
| KEGG pathway | |||||
| hsa04060 | Cytokine-cytokine receptor interaction | 5 | 243 | 2.70E-04 | 0.002418919 |
| GO:molecular function | |||||
| GO:0016493 | C-C chemokine receptor activity | 2 | 12 | 9.91E-03 | 0.091466094 |
| GO:0004950 | chemokine receptor activity | 2 | 17 | 0.014012135 | 0.127079826 |
| GO:0005515 | protein binding | 12 | 8785 | 0.040126286 | 0.325943103 |
| GO:biological process | |||||
| GO:0032729 | positive regulation of interferon-gamma production | 2 | 46 | 0.037690464 | 0.375903573 |
| GO:0006954 | inflammatory response | 3 | 379 | 0.038655183 | 0.383537965 |
| GO:0006955 | immune response | 3 | 421 | 0.046770989 | 0.444448439 |
GO terms and pathways the narcolepsy-associated genes involved.
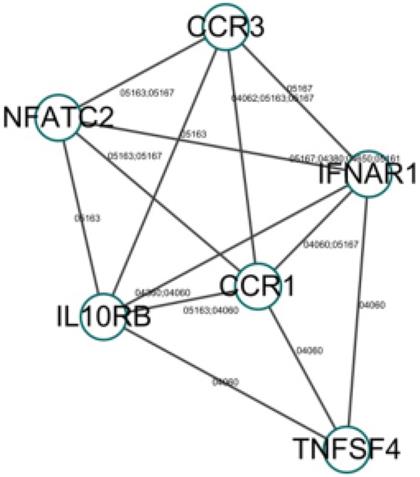
Gene-gene interaction network (the number next to the edge is the path number in the KEGG database).
PPI network integration and selection of hub genes
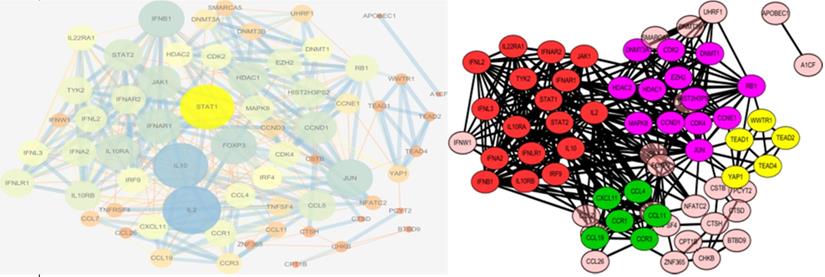
We used the STRING database and Cytoscape to investigate PPI networks. The top five hub genes were identified among the candidate genes, as well as among all the genes in the PPI network, using cytoHubba, and the genes with the highest degrees were considered as hub genes. The key genes in the PPI network among the candidate genes are shown in Table 3. These genes may play vital roles in the onset and development of narcolepsy. The PPI networks were further analyzed using the plug-in MCODE to detect potential modules, where four notable functional modules were detected using Cytoscape software (Figure 3). The top three modules were enriched mainly in cytokine-cytokine receptor interaction, protein binding, and chemokine-mediated signaling pathways, all of which were associated with immune response.
The key genes in the PPI network, among the candidate genes
| Gene | Description | Function | Degree (score) |
|---|---|---|---|
| IFNAR1 | interferon alpha and beta receptor subunit 1 | Encodes a type I membrane protein that forms one of the two chains of a receptor for interferons alpha and beta. | 23 |
| IL10RB | interleukin 10 receptor subunit beta | Encodes an accessory chain essential for the active interleukin 10 receptor complex. | 18 |
| DNMT1 | DNA methyltransferase 1 | Encodes an enzyme that transfers methyl groups to cytosine nucleotides of genomic DNA. | 14 |
| TNFSF4 | TNF superfamily member 4 | Encodes a cytokine of the tumor necrosis factor (TNF) ligand family. | 10 |
| NFATC2 | nuclear factor of activated T cell 2 | A DNA-binding protein with a REL-homology region (RHR) and an NFAT-homology region (NHR). | 7 |
The PPI network with the degree of genes is present as the size of circles, and the combined scores between proteins are presented as the size of the lines (left). The PPI network was constructed and formatted with genes separated into four notable functional modules, in different clusters with different colors(right). PPI, protein-protein interaction.
Discussion
Presently, the practices used for prevention, diagnosis, and treatment of narcolepsy are far from satisfactory. Exploring the genetic and molecular level dysfunction in narcolepsy patients can help develop effective treatment strategies and provide novel predictive and diagnostic clues for narcolepsy.
The results of pathway enrichment, gene-gene network, and the PPI network analyses indicated that immune responses and inflammatory responses are the main components of narcolepsy pathogenesis. These finding highlight the feasibility of developing polygenetic methods for drug development, pathogenesis exploration, and prognosis.
In our study, the GO term and KEGG pathway analyses indicated that the narcolepsy-associated genes were mainly enriched in pathways associated with immune response. GO biological process analysis found that in terms of molecular function (MF), the genes were associated mainly with C-C chemokine receptor activity, chemokine receptor activity, and protein binding. CCR (Chemokine receptors) are cytokine receptors found on the surface of certain cells that interact with a type of cytokine known as chemokine. Nineteen distinct chemokine receptors have been described in mammals, each of which has a 7-transmembrane (7TM) structure and couples to a G-protein for signal transduction within a cell, making them members of a large protein family of G protein-coupled receptors. Interaction with their specific chemokine ligands, chemokine receptors, triggers cell responses, including the onset of chemotaxis, which traffics the cell to a desired location within the organism. Some specific chemokine receptors are associated with viral affection or contribute to inflammatory diseases.
In terms of BP, the genes were mainly associated with positive regulation of interferon-gamma production, inflammatory response, and immune response, congruent to the pathways identified in the molecular function term. Interferon-gamma is a dimerized soluble cytokine that is the only member of the type II class of interferon, also known as macrophage-activating factor. It is a central regulator of the immune response and signals via the Janus Activated Kinase (JAK)-Signal Transducer and Activator of Transcription (STAT) pathway, and has broader roles in the activation of innate and adaptive immune responses to viruses and tumors. In addition, it is associated with interferon-gamma-mediated expression activation of major histocompatibility complex (MHC) class II transplantation antigen. Our KEGG pathway analysis revealed that a high level of enrichment in the cytokine-cytokine receptor interaction signaling pathways. Cytokines are soluble extracellular proteins or glycoproteins that are crucial intercellular regulators and mobilizers of cells engaged in innate and adaptive inflammatory host defenses, cell growth, differentiation, cell death, angiogenesis, and development and repair processes aimed at the restoration of homeostasis. Cytokines are released by various cells in the body, usually in response to an activating stimulus, and they induce responses through binding to specific receptors on the cell surface of target cells.
The results of the functional enrichment analyses demonstrated that the narcolepsy-associated genes were enriched in immune-related pathways. This is consistent with previously proposed hypothesis that the immune system plays a key role in narcolepsy [36, 37], based mainly on the indirect evidence of the close association between narcolepsy risk with H1N1 and HLA gene. However, the exact underlying mechanisms of this association remain unknown [37, 38]. The enrichment of narcolepsy-associated genes in C-C chemokine receptor activity, chemokine receptor activity, interferon-gamma production, and cytokine-cytokine receptor interaction pathways indicated that the onset of narcolepsy is associated not only with the cross-reaction autoimmune processes, but also with the dysfunction of immune-regulation, especially the dysfunction of cellular immunity. These results can help provide clues to reveal the exact mechanism of immune system dysfunction in narcolepsy patients, and possibly help identify new targets for immune therapy of narcolepsy, such as immune-modulation to prevent or delay the progression of narcolepsy.
We also constructed a PPI network and identified the top five hub genes, i.e., IFNAR1, IL10RB, DNMT1, TNFSF4, and NFATC2. Our results indicated that IFNAR1 was at the core of the PPI network. It belongs to the type II cytokine receptor family and encodes a type I membrane protein that forms one of the two chains of the type I interferon receptor. Binding and activation of the receptor leads to the phosphorylation of several proteins, including STAT1 and STAT2. The encoded protein also functions as an antiviral factor; it transforms the cell to an antiviral state in coordination with other cytokines [39, 40]. Various studies have reported that the pathways associated with IFNAR1 are associated with several diseases caused by viral affection [41-44]. It has been reported that the onset of narcolepsy is associated with H1N1 infection [45, 46]. It provides indirect evidence that the onset and progression of narcolepsy may be associated with abnormal immune response following a virus infection, in which the IFNAR1 dysfunction likely plays an important role, finally leading to the destruction of hypocretin neurons due to immune attack. The results indicated that the immune dysfunction of narcolepsy is associated not only with abnormal antigen recognition and cross-reaction, as indicated by the association of the HLA-DQB1*06:02 and TRA with narcolepsy risk [13, 20, 47], but also with the dysfunction of IFNAR1 immune-modulation and anti-viral processes. Thus, IFNAR1 may serve as a novel target for immune therapy of narcolepsy in the future.
Based on this analysis, the PPI network was identified and divided into four groups according to the protein-protein interactions. The top module included genes such as IFNAR1, IL2, and IL10RB, which are enriched mainly in the cytokine-cytokine receptor pathway. This result is in accordance with the results of the pathway enrichment analysis, and supported the role of IFNAR1 as the hub gene in the PPI network.
Nevertheless, the present study has some limitations. First, non-genetic factors may contribute to the onset of narcolepsy, and the effect of gene-environment interactions on narcolepsy incidence should also be considered. Second, the biological verification was not involved in the study. So further experimental studies are needed to confirm the identified hub genes and pathways, including RT-qPCR validation of these hub genes in clinical samples.
Conclusion
Through a comprehensive bioinformatic and heredity statistics re-analysis of GWAS data, we identified some crucial genes and pathways that were closely correlated with narcolepsy. Therefore, an overview regarding the molecular pathogenesis of narcolepsy and the potential for identification new drug targets for narcolepsy was provided. The results reported in the present study can eventually offer novel clues for screening and clinical diagnosis of narcolepsy, as well as understanding the molecular mechanisms underlying the pathogenesis of narcolepsy. However, further molecular biological experiments are required to verify these findings.
Acknowledgements
We thank Prof. Yiqun Wu from Peking University, Healthscience Center for advice, Shiying Wang from Peking University, College of Life Sciences for help with statistical analyses, and Prof. Fang Han from Peking University, People's Hospital for advice.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Mignot E, Lin L, Rogers W, Honda Y, Qiu X, Lin X. et al. Complex HLA-DR and -DQ interactions confer risk of narcolepsy-cataplexy in three ethnic groups. American journal of human genetics. 2001;68:686-99
2. Han F, Lin L, Schormair B, Pizza F, Plazzi G, Ollila HM. et al. HLA DQB1*06:02 negative narcolepsy with hypocretin/orexin deficiency. Sleep. 2014;37:1601-8
3. Allman R, Dite GS, Hopper JL, Gordon O, Starlard-Davenport A, Chlebowski R. et al. SNPs and breast cancer risk prediction for African American and Hispanic women. Breast cancer research and treatment. 2015;154:583-9
4. Dauriz M, Porneala BC, Guo X, Bielak LF, Peyser PA, Durant NH. et al. Association of a 62 Variants Type 2 Diabetes Genetic Risk Score With Markers of Subclinical Atherosclerosis: A Transethnic, Multicenter Study. Circulation Cardiovascular genetics. 2015;8:507-15
5. French L, Gray C, Leonard G, Perron M, Pike GB, Richer L. et al. Early Cannabis Use, Polygenic Risk Score for Schizophrenia and Brain Maturation in Adolescence. JAMA psychiatry. 2015;72:1002-11
6. Gui L, Wu F, Han X, Dai X, Qiu G, Li J. et al. A multilocus genetic risk score predicts coronary heart disease risk in a Chinese Han population. Atherosclerosis. 2014;237:480-5
7. Abelson S, Collord G, Ng SWK, Weissbrod O, Mendelson Cohen N, Niemeyer E. et al. Prediction of acute myeloid leukaemia risk in healthy individuals. Nature. 2018;559:400-4
8. Zhang KK, Xiang M, Zhou L, Liu J, Curry N, Heine Suner D. et al. Gene network and familial analyses uncover a gene network involving Tbx5/Osr1/Pcsk6 interaction in the second heart field for atrial septation. Human molecular genetics. 2016;25:1140-51
9. Yip DK, Chan LL, Pang IK, Jiang W, Tang NLS, Yu W. et al. A network approach to exploring the functional basis of gene-gene epistatic interactions in disease susceptibility. Bioinformatics (Oxford, England). 2018;34:1741-9
10. Medicine AAoS. International Classification of Sleep Disorders.3rd ed. Darien,IL: American Academy of Sleep Medicine. 2014
11. Han F, Faraco J, Dong XS, Ollila HM, Lin L, Li J. et al. Genome wide analysis of narcolepsy in China implicates novel immune loci and reveals changes in association prior to versus after the 2009 H1N1 influenza pandemic. PLoS genetics. 2013;9:e1003880
12. Toyoda H, Miyagawa T, Koike A, Kanbayashi T, Imanishi A, Sagawa Y. et al. A polymorphism in CCR1/CCR3 is associated with narcolepsy. Brain, behavior, and immunity. 2015;49:148-55
13. Han F, Lin L, Li J, Aran A, Dong SX, An P. et al. TCRA, P2RY11, and CPT1B/CHKB associations in Chinese narcolepsy. Sleep medicine. 2012;13:269-72
14. Miyagawa T, Kawashima M, Nishida N, Ohashi J, Kimura R, Fujimoto A. et al. Variant between CPT1B and CHKB associated with susceptibility to narcolepsy. Nature genetics. 2008;40:1324-8
15. Holm A, Lin L, Faraco J, Mostafavi S, Battle A, Zhu X. et al. EIF3G is associated with narcolepsy across ethnicities. European journal of human genetics: EJHG. 2015;23:1573-80
16. Faraco J, Lin L, Kornum BR, Kenny EE, Trynka G, Einen M. et al. ImmunoChip study implicates antigen presentation to T cells in narcolepsy. PLoS genetics. 2013;9:e1003270
17. Hor H, Kutalik Z, Dauvilliers Y, Valsesia A, Lammers GJ, Donjacour CE. et al. Genome-wide association study identifies new HLA class II haplotypes strongly protective against narcolepsy. Nature genetics. 2010;42:786-9
18. Kornum BR, Kawashima M, Faraco J, Lin L, Rico TJ, Hesselson S. et al. Common variants in P2RY11 are associated with narcolepsy. Nature genetics. 2011;43:66-71
19. Shimada M, Miyagawa T, Kawashima M, Tanaka S, Honda Y, Honda M. et al. An approach based on a genome-wide association study reveals candidate loci for narcolepsy. Human genetics. 2010;128:433-41
20. Tafti M, Hor H, Dauvilliers Y, Lammers GJ, Overeem S, Mayer G. et al. DQB1 locus alone explains most of the risk and protection in narcolepsy with cataplexy in Europe. Sleep. 2014;37:19-25
21. Johnson AD, Handsaker RE, Pulit SL, Nizzari MM, O'Donnell CJ, de Bakker PI. SNAP: a web-based tool for identification and annotation of proxy SNPs using HapMap. Bioinformatics (Oxford, England). 2008;24:2938-9
22. Huang da W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature protocols. 2009;4:44-57
23. Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M. et al. The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. 2017; 45: D362-d8.
24. Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome research. 2003;13:2498-504
25. Chin CH, Chen SH, Wu HH, Ho CW, Ko MT, Lin CY. cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC systems biology. 2014;8(Suppl 4):11
26. Kraft P, Hunter DJ. Genetic risk prediction-are we there yet? The New England journal of medicine. 2009;360:1701-3
27. Qi L, Parast L, Cai T, Powers C, Gervino EV, Hauser TH. et al. Genetic susceptibility to coronary heart disease in type 2 diabetes: 3 independent studies. Journal of the American College of Cardiology. 2011;58:2675-82
28. Yamasaki M, Miyagawa T, Toyoda H, Khor SS, Liu X, Kuwabara H. et al. Evaluation of polygenic risks for narcolepsy and essential hypersomnia. Journal of human genetics. 2016;61:873-8
29. Capittini C, De Silvestri A, Terzaghi M, Scotti V, Rebuffi C, Pasi A. et al. Correlation between HLA-DQB1*06:02 and narcolepsy with and without cataplexy: approving a safe and sensitive genetic test in four major ethnic groups. A systematic meta-analysis. Sleep medicine. 2018;52:150-7
30. Dauvilliers Y, Neidhart E, Lecendreux M, Billiard M, Tafti M. MAO-A and COMT polymorphisms and gene effects in narcolepsy. Molecular psychiatry. 2001;6:367-72
31. Kernohan KD, Cigana Schenkel L, Huang L, Smith A, Pare G, Ainsworth P. et al. Identification of a methylation profile for DNMT1-associated autosomal dominant cerebellar ataxia, deafness, and narcolepsy. Clinical epigenetics. 2016;8:91
32. Moghadam KK, Pizza F, Tonon C, Lodi R, Carelli V, Poli F. et al. Polysomnographic and neurometabolic features may mark preclinical autosomal dominant cerebellar ataxia, deafness, and narcolepsy due to a mutation in the DNA (cytosine-5-)-methyltransferase gene, DNMT1. Sleep medicine. 2014;15:582-5
33. Gibson G. Rare and common variants: twenty arguments. Nature reviews Genetics. 2012;13:135-45
34. Golan D, Lander ES, Rosset S. Measuring missing heritability: inferring the contribution of common variants. Proceedings of the National Academy of Sciences of the United States of America. 2014;111:E5272-81
35. Fuchsberger C, Flannick J, Teslovich TM, Mahajan A, Agarwala V, Gaulton KJ. et al. The genetic architecture of type 2 diabetes. Nature. 2016;536:41-7
36. Taheri S. The Immune Basis of Narcolepsy: What Is the Evidence? Sleep medicine clinics. 2017;12:279-87
37. Barateau L, Liblau R, Peyron C, Dauvilliers Y. Narcolepsy Type 1 as an Autoimmune Disorder: Evidence, and Implications for Pharmacological Treatment. CNS drugs. 2017;31:821-34
38. Degn M, Kornum BR. Type 1 narcolepsy: a CD8(+) T cell-mediated disease? Annals of the New York Academy of Sciences. 2015;1351:80-8
39. Schreiber G. The molecular basis for differential type I interferon signaling. The Journal of biological chemistry. 2017;292:7285-94
40. Schreiber G, Piehler J. The molecular basis for functional plasticity in type I interferon signaling. Trends in immunology. 2015;36:139-49
41. Sebina I, Haque A. Effects of type I interferons in malaria. Immunology. 2018;155:176-85
42. Scagnolari C, Antonelli G. Type I interferon and HIV: Subtle balance between antiviral activity, immunopathogenesis and the microbiome. Cytokine & growth factor reviews. 2018;40:19-31
43. Rehman SU, Rauf M, Abbas Z, Hamed MH, Qadri I. Role of Some Predominant Host Immunomodulators' Single Nucleotide Polymorphisms in Severity of Hepatitis B Virus and Hepatitis C Virus Infection. Viral immunology. 2016;29:536-45
44. Lehmann MH, Torres-Dominguez LE, Price PJ, Brandmuller C, Kirschning CJ, Sutter G. CCL2 expression is mediated by type I IFN receptor and recruits NK and T cells to the lung during MVA infection. Journal of leukocyte biology. 2016;99:1057-64
45. Szakacs A, Darin N, Hallbook T. Increased childhood incidence of narcolepsy in western Sweden after H1N1 influenza vaccination. Neurology. 2013;80:1315-21
46. Partinen M, Kornum BR, Plazzi G, Jennum P, Julkunen I, Vaarala O. Narcolepsy as an autoimmune disease: the role of H1N1 infection and vaccination. The Lancet Neurology. 2014;13:600-13
47. Pelin Z, Guilleminault C, Risch N, Grumet FC, Mignot E. HLA-DQB1*0602 homozygosity increases relative risk for narcolepsy but not disease severity in two ethnic groups. US Modafinil in Narcolepsy Multicenter Study Group. Tissue antigens. 1998;51:96-100
Author contact
Corresponding author: Jun Zhang, Department of Neuromedicine, Peking University People's Hospital, 11 Xizhimen South Street, Beijing, China; E-mail: zhangjunedu.cn; TEL: +86 13517379562; FAX: +86 10 82236809.

 Global reach, higher impact
Global reach, higher impact