Impact Factor ISSN: 1449-1907
Int J Med Sci 2018; 15(9):859-866. doi:10.7150/ijms.24928 This issue Cite
Research Paper
Myokines related to leukocyte recruitment are down-regulated in osteosarcoma
1. Department of Orthopedics, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, Shanghai 200233, P.R. China.
2. Institution of microsurgery for limbs, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, Shanghai 200233, P.R. China.
3. Department of Oncology, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, Shanghai 200233, P.R. China.
Received 2018-1-14; Accepted 2018-4-27; Published 2018-5-26
Abstract
Myokines are cytokines that are secreted by muscle cells during exercises, muscle development and pathology. Studies have shown that expression of some individual myokines was altered in tumors. However, comprehensive analyses of myokines' expression in osteosarcoma (OS), the most common malignant tumor in musculoskeletal system, have not been performed. In this study, we analyzed the expression of 35 myokines in osteosarcoma, peritumoral skeletal muscle, and cancellous bone by qRT-PCR. Heatmap analysis based on the expression pattern of these myokines revealed that OS is more likely derived from cancellous bone than peritumoral skeletal muscle. Thus, we compared the expression of myokines between OS and cancellous bone to reveal a potential role of myokines in OS development. Our results showed that expression of 19 myokines in OS was significantly lower than that in cancellous bone. KEGG signaling pathway analysis showed that these 19 myokines are involved in several important signaling pathways, one of which was associated with leukocyte recruitment in TNF-α signaling. We verified that expression of these leukocyte recruitment-related myokines were down-regulated in OS cell line MNNG compared to those in human BMSC. Downregulation of the myokines related to leukocyte recruitment suggests that escaping from host immune system may help the occurrence of osteosarcoma.
Keywords: myokine screening, osteosarcoma, cancellous bone, peritumoral skeletal muscle, leukocyte recruitment
Introduction
Osteosarcoma (OS), the most common and frequent primary malignant solid neoplasm in musculoskeletal system, is believed to develop from primitive transformed cells of mesenchymal origins which are differentiated into osteoblastic lineages and produces malignant osteoid tissues. Although the estimated incidence rate is only 5/1,000,000, osteosarcoma (OS) has a peak incidence in teenagers and young adults [1, 2], which affects patients' life quality in their remaining years and places heavier burden on society. The major treatments of OS include tumor excision surgery, chemotherapy and radiotherapy. In the past decades, the 5-year survival rates for patients with localized OS remain approximately 60-70% [3]. Furthermore, the 5-year survival rates for patients who have remote metastatic niches is only around 20% [4, 5]. The main reasons hindering the progress of osteosarcoma treatment is the lack of knowledge of tumorigenesis and biomarkers for early diagnosis [6].
Although numerous studies have been performed to discover the biomarkers for osteosarcoma in its early stage, little successes have been achieved [7-9]. Up to now, the main diagnosis methods for osteosarcoma remain medical imaging and biopsy [10]. To our knowledge, none of the biomarkers under studies has been well accepted in clinic worldwide. It is well known tumor biomarkers are molecules that are secreted by tumors or specific responses of the body to the presence of tumors. Therefore, understanding the responses of osteosarcoma peripheral tissues (e.g. skeletal muscle and cancellous bone) or their cross-talk may help the discovery of novel early diagnosis biomarkers of osteosarcoma.
About 20 years ago, skeletal muscle has been identified as an endocrine and paracrine organ, which has the ability to produce a variety of humoral cytokines and growth factors termed as “myokine”[11, 12]. Skeletal muscle can communicate with other organs including bone, adipose tissue, brain, liver and pancreas by secreting myokines[13]. These muscle-derived peptides have been proposed to be involved in anti-inflammatory and metabolic effects, as well as mediating the preventive effects of exercise against chronic diseases [14, 15]. A few studies have focused on the functions and expression changes of myokines during exercises [16]. For example, during exercise, IL6 is the first detectable myokine which have an antiinflammatory effect[17]. Recent studies showed that a few individual myokine could mediate metabolic changes, regulate tumorigenesis and tumor invasion to other remote organs such as liver and breast [18, 19]. However, comprehensive analysis of expression for all known myokines under pathological condition, particularly in motor system diseases, is very limited [20]. Therefore, in this study, we analyzed the expression of all known myokines (n=35) in osteosarcoma tissue, cancellous bone and peritumoral skeletal muscles. Our results showed that expression of 19 myokines in OS was significantly lower than that in cancellous bone. KEGG signaling pathway analysis further showed that these 19 myokines are involved in several important signaling pathways, one of which was associated with leukocyte recruitment in TNF-α signaling. These results indicated that these differentially myokines could be potential biomarkers for the diagnosis of osteosarcoma. Downregulation of the myokines related to leukocyte recruitment in TNF-α signaling also suggests that osteosarcoma may occur by escaping from host innate immune system.
Material and method
Clinical tissues and cells
All the osteosarcoma tissues and peritumoral muscle tissues were obtained from the patients with osteosarcoma who received the surgery treatment in Shanghai Sixth People's Hospital. The normal cancellous bone tissues were obtained from the normal distal end of patients with osteosarcoma treated with amputation, or the patients with severe trauma who underwent the amputation in Shanghai Sixth People's Hospital. All the tissues were harvested for RNA and protein extraction. The human osteosarcoma cell line MNNG was purchased from the American Type Culture Collection (Manassas, VA, USA). The BMSC were obtained from Shanghai sixth People's Hospital (Shanghai, China)
RNA Preparation
Approximately, 100 mg tissues were put in a grind tube with steel balls and 1 ml Trizol reagent, and homogenized by a tissue grinder (Scientz-48, Ningbo, China). About 1 x 107 cells were mixed with 1 ml Trizol reagent. Total RNA from cells, osteosarcoma tissues, peritumoral muscle tissues, and cancellous bone tissues was extracted and purified using The E.Z.N.A.® Total RNA Kit IR6834-01 (OMEGA Bio-tek, Norcross, GA, U.S.A) according to the manufacturer's protocol. The concentration and quality of RNA were determined by a NanoDrop ND-1000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA).
Reverse‐transcription quantitative PCR (RT‐qPCR)
About 1 μg of total RNA was reverse transcribed using the C1000™ Thermal Cycler (Bio-Rad, CA, USA) according to the manufacturer's instruction. Thermal profile consisted of 10 minutes at 25 °C followed by 60 minutes at 55 °C and 5 minutes at 85 °C. All qPCR reactions were performed in a total volume of 10 μl mixture, including forward and reverse primers, cDNA and 1x SYBR® Green PCR Master Mix (Thermo scientific, Waltham, MA, USA). Cycling conditions were as follows: 95 °C for 10 min, followed by 40 cycles of 15 s at 95 °C and 60 s at 72 °C using a ABI 7900HT machine with software SDS version 2.3 (Thermo scientific, Waltham, MA, USA). Primers used in this study are listed in Table S1, including Eukaryotic Translation Elongation Factor 2 (eEF2) and Glyceraldehyde-3-phosphate Dehydrogenase (GAPDH) as endogenous invariant controls for data normalization.
Protein analysis
All lysates were obtained by using RIPA Lysis Buffer (Santa Cruz, Dallas, TX, USA). 30μg proteins were applied to SDS-PAGE (10% gel) and transferred to PVDF membranes (Bio-Rad Laboratories, Hercules, CA), probed with primary antibodies anti-FNDC-5 (1:4000, Abcam), anti-ANGPTL-4 (1:1000, Abcam), anti-IL-15(1:1000, Abcam), anti-MCP-1 (1:1000, Abcam) over night at 4℃. After washing, they were incubated with HRP-labelled anti-mouse (1:3000, CST) and anti-rabbit (1:3000, CST) secondary antibodies for 1h at room temperature. Anti-actin Ab was used as control of protein loading.
KEGG pathway analysis
Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways were obtained from the website: http://www.genome.jp/kegg/pathway.html to determine the differentially expressed genes. Processes and pathways that showed high enrichment of upregulated levels of myokines and certain pathway cascades were displayed.
Statistical analysis
GraphPad Prism v.6.0 (GraphPad software, CA, USA) and SPSS versions 18.0 (SPSS Inc., Chicago, IL) were used for statistical analysis. Data are presented as the mean ± SEM. Myokine gene expression data were analyzed using a 2-△△CT method and eEF2 was used as an internal control for each technical replicate and averaged for each biological replicate. Gene expression of normal cancellous bone was regarded as the baseline for the comparison. △CT values from each group were compared by the non-parametric test to determine statistically significant differences between osteosarcoma and cancellation bone. P values of < 0.05 were considered as statistically significant.
Result
Myokines are expressed universally in musculoskeletal tissues
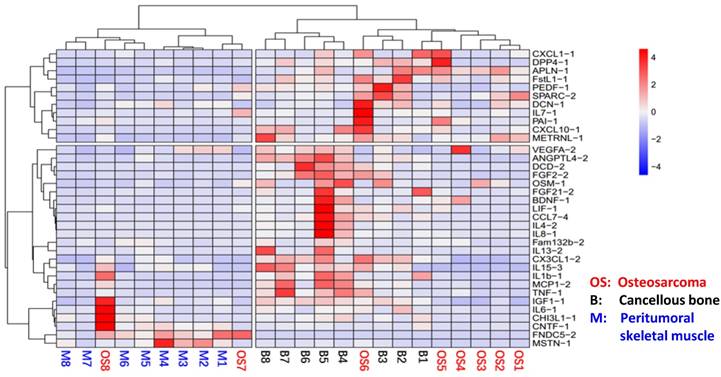
In order to analyze whether myokines are universally expressed in musculoskeletal system in OS patients, we analyzed the expression levels of all 35 myokines in OS tissue, peritumoral muscles and cancellous bones using qRT-PCR. The results showed that all 35 myokines were universally expressed in these three tissues (Table S1). Moreover, the myokine expression patterns in these three tissues can be clearly clustered into two parts in heat map pattern (Fig. 1). The left part consists of all 8 peritumoral muscle tissues and 2 OS tissues; and the right part is formed from other 6 OS tissues and all 8 cancellous bone tissues. In the right part, bone tissues B4-8 were clustered, while OS tissues OS1-5 were also clustered. These data indicated that the myokine expression is universal in musculoskeletal tissues, and myokine expression pattern of OS to cancellous bone is more similar than that to peritumoral tissues.
The relationships of osteosarcoma, cancellous bone and peritumoral skeletal muscle in myokine expression angle. The myokine expression levels of cancellous bone, peritumoral muscle and osteosarcoma tissues were clearly clustered into two parts. Myokine expression between peritumoral muscle tissues and cancellous bone tissues differed obviously, whereas the myokine expression in osteosarcoma tissues were remained between them.
Most of the myokine expression was down-regulated in OS compared with those in cancellous bone
As myokine expression pattern in cancellous bone and osteosarcoma are close and osteosarcoma is believed to be derived from transformed bone marrow mesenchymal cells, we compared the expression levels of all 35 myokines in these two tissues in order to identify myokines that are differentially expressed between osteosarcoma and cancellous bone tissues. qPCR results showed that 19 genes were significantly down-regulated in osteosarcoma compared with those in cancellous bone tissues (Fig. 2). These down-regulated genes could be divided into 5 groups according to their biological functions (Fig. 2). We also found that 3 myokines (FNDC5, IL-7 and PAI-1) were upregulated in the osteosarcoma compared to those in cancellous bone (Fold changes > 2.0). Although none of them showed significant difference between these two tissues, the expression level of myokine FNDC5 was over 20 folds higher than that in cancellous bone tissues (Fig. 2D).
The myokine expression levels in osteosarcoma tissues compared with cancellous bone tissues. (A) Expression levels of myokine related with immunological regulation and inflammatory, CCL7, CHI3L1, CX3CL1, DDP4, IL1b, IL4, IL8, IL15, LIF and MCP1 showed significant differences. (B) Expression levels of myokines related with cell survival and differentiation, DCD, FGF2 and OSM showed significant differences. (C) Expression levels of myokines related with neural growth, BDNF showed significant differences. (D) Expression levels of myokines related with metabolism, ANGPTL4, Fam132b and FGF21 showed significant differences. (E) Myokines with other functions, TNF-α showed significant differences. (n = 8, **P < 0.01, *P < 0.05) (F) The protein level of ANGPTL4, IL15 and MCP1 in three randomly chosen OS tissues and cancellous bone tissues.
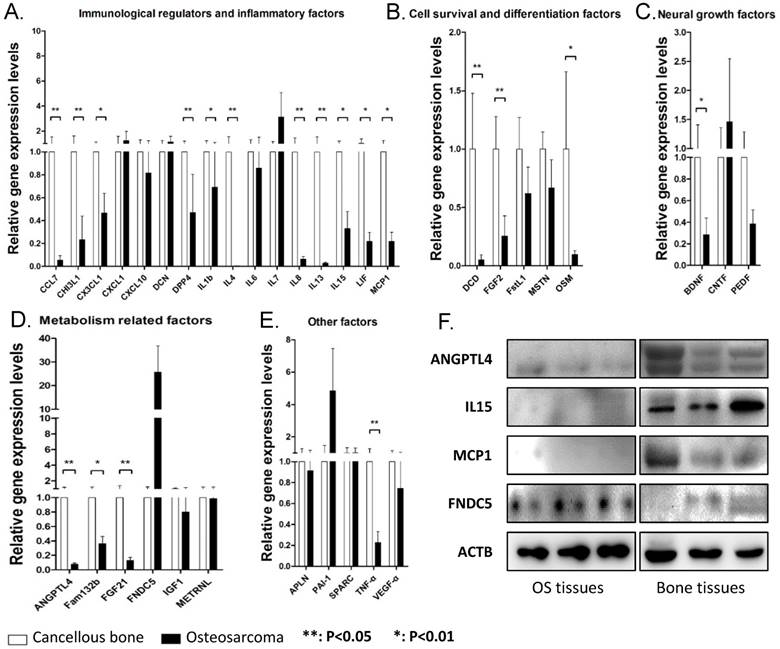
In order to verify the expressions of these myokines in their protein levels, representative OS and cancellous bone tissue lysis were analyzed by antibodies against selected myokines. The results showed that ANGPTL4, IL15 and MCP1 proteins were indeed down-regulated in OS tissues, whereas FNDC5 was up-regulated (Fig. 2F).
KEGG analysis revealed pathways that are associated with OS
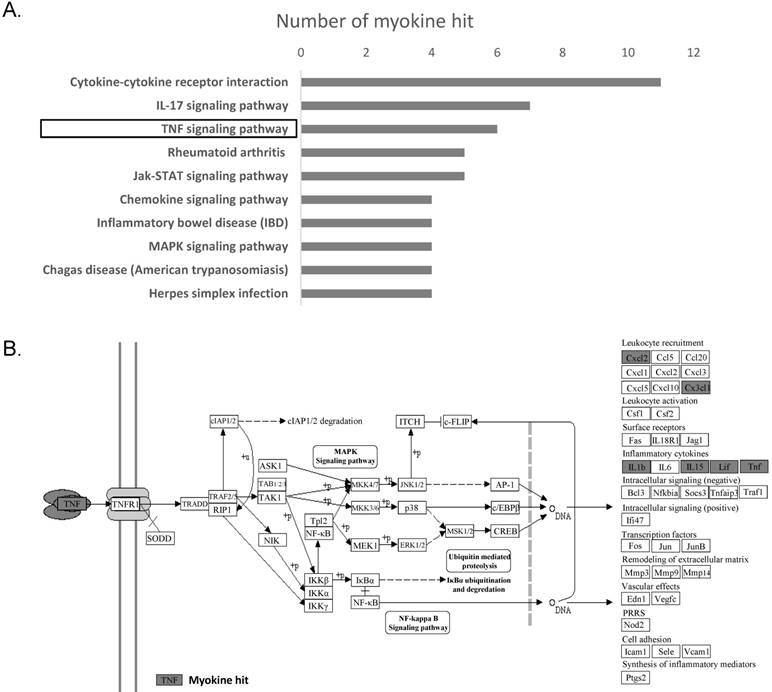
In order to identify the molecular mechanism underlying the changes in myokine expression between OS tissue and normal cancellous bone, we performed a KEGG pathway analyses on 19 genes whose expression significantly differed in these two tissues. The top 10 signaling pathway are displayed in Fig. 3a. and the top 3 are 'Cytokine-cytokine receptor interaction'(ko04060), 'IL-17 signaling pathway' (ko04657) and 'TNF signaling pathway' (ko04668). Cytokine-cytokine receptor interaction has a very broad definition, IL-17 signaling is mainly reported in the immune system and TNF signaling is associated with leukocyte recruitment and Inflammatory.
TNF-α signaling pathway was found to be inhibited in osteosarcoma. (A) Top 10 signaling pathways discovered by KEGG were displayed basing on the significantly differentially expressed myokines. (B) Significant changes in myokine expression levels were observed in osteosarcoma compared to cancellous bone in the downstream of TNF signaling pathway(ko04668). Hit myokines are labeled with grey boxes.
Verification of the TNF signaling pathway in OS cell line
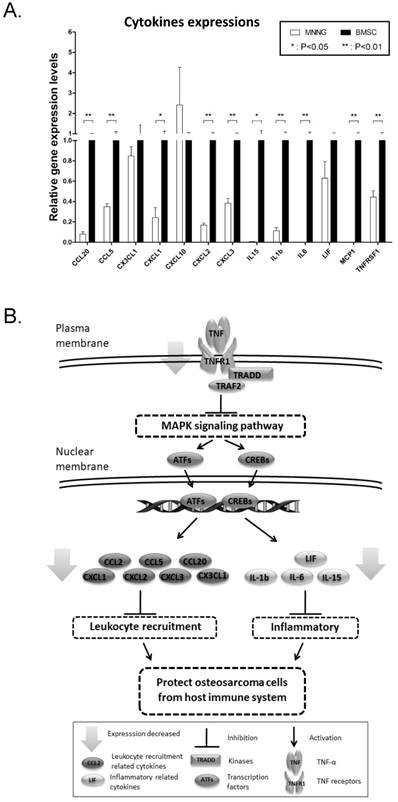
We are most interested in TNF signaling pathway as it may suggest potential inhibition of immune system in osteosarcoma tissues. To verify the TNF signaling pathway, we further determined the expression of genes that are associated with inflammatory (IL1b, IL6, IL15, LIF), leukocyte recruitment (MCP1, CCL5, CCL20, CXCL1, CXCL2, CXCL3, CXCL10, CX3CX1) and the receptor of TNF-α (TNFR1) in osteosarcoma cell line MNNG using qPCR. Human BMSC cells, which is from normal cancellous bone, are used as a normal control. Not surprisingly, the result showed that majority of these genes (with three exceptions: CX3CL1, CXCL10 and LIF) were significantly down-regulated in MNNG cell compared to that in human BMSC cells (Fig. 4A). Taken together, these results verified that TNF signaling pathway is down-regulated in OS. Down-regulation of myokines related to leukocytes recruitment in TNF signaling may protect OS from being attacked by leukocytes (Fig. 4B).
Verification of the down-regulation of TNF-α pathway. (A) Cytokines involved in inflammatory and leukocyte recruitment in the TNF signaling pathway map (ko04668) were detected by qPCR in osteosarcoma cell line MNNG and human BMSC cell. CCL20, CCL5, CXCL1, CXCL2, CXCL3, IL15, IL1b, IL6, MCP1 and TNFRSF1 were significantly decreased (**P < 0.01, *P < 0.05). (B) The diagram of how the inhibition of TNF protects osteosarcoma cells from host immune system.
Discussion
In the last decades, myokines are identified as cytokines and growth factors that exert their functions through endocrine or paracrine pattern [11, 21]. The function of myokines has been widely studied in physiological processes, such as exercise [16], fasting and temperature keeping [22], pathological processes, such as diabetes [23, 24], and obesity [25]. More importantly, many studies have revealed the role of some individual myokines in various type of cancers, such as breast cancer, prostate cancer and so on [26-28]. However, comprehensive analysis of all known myokines has not been performed under pathological condition, particularly in OS. To our knowledge, this is the first study that identified all known myokines that are differentially expressed in osteosarcoma, peritumoral skeletal muscle and cancellous bone, and explored the potential molecular mechanisms underlying these differences in osteosarcoma, and cancellous bone, which may become potential biomarkers for the diagnosis of osteosarcoma.
In this study, the expression of 35 myokines (to the best of our knowledge, these are almost all the myokine genes) were analyzed in the osteosarcoma tissues, normal cancellous bone tissues and peritumoral muscle tissues. It is well known that bone and skeletal muscle are both derived from somatic mesoderm during embryonic development. Muscle pouch has also been fully used in ectopic bone formation induced by BMPs [29]; hence, we thought that there might be some similarity of myokine expression between these three tissues [30, 31]. As expected, the results from qPCR screening showed a universal expression of myokines in OS, peritumoral muscle and cancellous bones. Besides, the clustered heat map (Fig. 1) showed that the expression of myokines in osteosarcoma is more similar to that in the cancellous bone compared with that in peritumoral muscle tissues. This indicates that myokines can be used as a parameter to tell the relations between musculoskeletal tissues. Based on the different expression profile between cancellous bone and osteosarcoma, the result suggests that the inhibition of the TNF-α signaling pathway probably protects osteosarcoma cells from the attacking of leukocytes according to the KEGG signaling pathway analysis.
As known to all, tumor necrosis factor alpha (TNF-α), which is mainly secreted by activated macrophages and activated T lymphocytes, has been demonstrated to play an important role in inflammatory, apoptosis, angiogenesis, and cell proliferation via the regulation of various signaling pathways, such as MAPK signaling pathway [32, 33]. On the other hand, it has the ability to cause apoptosis of tumor-associated endothelial cells by clustering death domain-containing proteins, leading to caspase activation [34], ultimately leading to the complete destruction of the tumor vasculature [35]. With regard to the functions of TNF-α in osteosarcoma, some studies showed that TNF-α and inflammatory cytokines are required for the tumorigenesis and inducing the migration and invasion of osteosarcoma [36-38], whereas some diverse conclusions were observed in other studies. A recent study reported that TNF-α were upregulated when the tumorigenesis gene HOTAIR was knocked down in osteosarcoma to inhibit the proliferation and differentiation [39]. Moreover, Pahl et. al. also showed that human macrophages which take a major responsibility for TNF-α secretion can be induced to exert direct anti-tumor activity against osteosarcoma cells [40]. Consistent with this result, our study also indicates that TNF signaling is probably attenuated to reduce the expressions of leukocyte-related cytokines in osteosarcoma cells, which protects osteosarcoma cells from the attack of leukocytes from host. This finding enriches our knowledge of how osteosarcoma protect themselves from host immune system, which could offer a novel approach to develop osteosarcoma therapies in the future.
In conclusion, we demonstrated that the expression of myokines is universal in OS, peritumoral muscle and cancellous bone and the relation between cancellous bone and OS is closer than that between peritumoral muscle and OS in terms of myokines' expression pattern. Moreover, we also indicate that the TNF-α signaling pathway is inhibited in OS tissue and cell, resulting in suppression of downstream cytokines related to inflammation and leukocyte recruitment in OS. Our results also suggest that myokines that are associated with TNF signaling pathway could be potential biomarkers for the diagnosis of OS.
Supplementary Material
Table S1.
Acknowledgements
This work was supported by the Shanghai Jiaotong University Medical and Industry cross fund (YG2017MS21) and National Natural Science Foundation of China (No.81501939).
Ethics Committee Approval
This study was approved by the Ethics Committee of Shanghai Jiao Tong University Affiliated Sixth People's Hospital and performed according to the Principles of the Declaration of Helsinki.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Botter SM, Neri D, Fuchs B. Recent advances in osteosarcoma. Current opinion in pharmacology. 2014;16:15-23
2. Ottaviani G, Jaffe N. The epidemiology of osteosarcoma. Cancer treatment and research. 2009;152:3-13
3. Longhi A, Fabbri N, Donati D, Capanna R, Briccoli A, Biagini R. et al. Neoadjuvant chemotherapy for patients with synchronous multifocal osteosarcoma: results in eleven cases. Journal of chemotherapy. 2001;13(3):324-30
4. Bacci G, Briccoli A, Rocca M, Ferrari S, Donati D, Longhi A. et al. Neoadjuvant chemotherapy for osteosarcoma of the extremities with metastases at presentation: recent experience at the Rizzoli Institute in 57 patients treated with cisplatin, doxorubicin, and a high dose of methotrexate and ifosfamide. Annals of oncology: official journal of the European Society for Medical Oncology. 2003;14(7):1126-34
5. Chen L, Wang Q, Wang GD, Wang HS, Huang Y, Liu XM. et al. miR-16 inhibits cell proliferation by targeting IGF1R and the Raf1-MEK1/2-ERK1/2 pathway in osteosarcoma. FEBS letters. 2013;587(9):1366-72
6. Marina N, Gebhardt M, Teot L, Gorlick R. Biology and therapeutic advances for pediatric osteosarcoma. The oncologist. 2004;9(4):422-41
7. Fei D, Li Y, Zhao D, Zhao K, Dai L, Gao Z. Serum miR-9 as a prognostic biomarker in patients with osteosarcoma. The Journal of international medical research. 2014;42(4):932-7
8. Zhou G, Lu M, Chen J, Li C, Zhang J, Chen J. et al. Identification of miR-199a-5p in serum as noninvasive biomarkers for detecting and monitoring osteosarcoma. Tumour biology: the journal of the International Society for Oncodevelopmental Biology and Medicine. 2015;36(11):8845-52
9. Kubo T, Shimose S, Matsuo T, Fujimori J, Arihiro K, Ochi M. Interferon-alpha/beta receptor as a prognostic marker in osteosarcoma. The Journal of bone and joint surgery American volume. 2011;93(6):519-26
10. Moore DD, Luu HH. Osteosarcoma. Cancer treatment and research. 2014;162:65-92
11. Karstoft K, Pedersen BK. Skeletal muscle as a gene regulatory endocrine organ. Current opinion in clinical nutrition and metabolic care. 2016;19(4):270-5
12. Pedersen BK, Akerstrom TC, Nielsen AR, Fischer CP. Role of myokines in exercise and metabolism. Journal of applied physiology. 2007;103(3):1093-8
13. Petersen AM, Pedersen BK. The anti-inflammatory effect of exercise. Journal of applied physiology. 2005;98(4):1154-62
14. Benatti FB, Pedersen BK. Exercise as an anti-inflammatory therapy for rheumatic diseases-myokine regulation. Nature reviews Rheumatology. 2015;11(2):86-97
15. Pedersen BK, Saltin B. Exercise as medicine - evidence for prescribing exercise as therapy in 26 different chronic diseases. Scandinavian journal of medicine & science in sports. 2015;25(Suppl 3):1-72
16. Huh JY, Siopi A, Mougios V, Park KH, Mantzoros CS. Irisin in response to exercise in humans with and without metabolic syndrome. The Journal of clinical endocrinology and metabolism. 2015;100(3):E453-7
17. Steensberg A, Fischer CP, Keller C, Moller K, Pedersen BK. IL-6 enhances plasma IL-1ra, IL-10, and cortisol in humans. American journal of physiology Endocrinology and metabolism. 2003;285(2):E433-7
18. Pedersen BK, Febbraio MA. Muscle as an endocrine organ: focus on muscle-derived interleukin-6. Physiological reviews. 2008;88(4):1379-406
19. Kjaer M, Pollack SF, Mohr T, Weiss H, Gleim GW, Bach FW. et al. Regulation of glucose turnover and hormonal responses during electrical cycling in tetraplegic humans. The American journal of physiology. 1996;271(1 Pt 2):R191-9
20. Paulson TA, Bishop NC, Smith BM, Goosey-Tolfrey VL. Inflammation-mediating cytokine response to acute handcycling exercise with/without functional electrical stimulation-evoked lower-limb cycling. Journal of rehabilitation research and development. 2014;51(4):645-54
21. Lightfoot AP, Cooper RG. The role of myokines in muscle health and disease. Current opinion in rheumatology. 2016;28(6):661-6
22. Li X, Fang W, Hu Y, Wang Y, Li J. Characterization of fibronectin type III domain-containing protein 5 (FNDC5) gene in chickens: Cloning, tissue expression, and regulation of its expression in the muscle by fasting and cold exposure. Gene. 2015;570(2):221-9
23. Akerstrom TC, Krogh-Madsen R, Petersen AM, Pedersen BK. Glucose ingestion during endurance training in men attenuates expression of myokine receptor. Experimental physiology. 2009;94(11):1124-31
24. Gray SR, Kamolrat T. The effect of exercise induced cytokines on insulin stimulated glucose transport in C2C12 cells. Cytokine. 2011;55(2):221-8
25. Peterson JM, Mart R, Bond CE. Effect of obesity and exercise on the expression of the novel myokines, Myonectin and Fibronectin type III domain containing 5. PeerJ. 2014;2:e605
26. Hojman P, Dethlefsen C, Brandt C, Hansen J, Pedersen L, Pedersen BK. Exercise-induced muscle-derived cytokines inhibit mammary cancer cell growth. American journal of physiology Endocrinology and metabolism. 2011;301(3):E504-10
27. Gannon NP, Vaughan RA, Garcia-Smith R, Bisoffi M, Trujillo KA. Effects of the exercise-inducible myokine irisin on malignant and non-malignant breast epithelial cell behavior in vitro. International journal of cancer. 2015;136(4):E197-202
28. Hayes BD, Brady L, Pollak M, Finn SP. Exercise and Prostate Cancer: Evidence and Proposed Mechanisms for Disease Modification. Cancer epidemiology, biomarkers & prevention: a publication of the American Association for Cancer Research, cosponsored by the American Society of Preventive Oncology. 2016;25(9):1281-8
29. Scott MA, Levi B, Askarinam A, Nguyen A, Rackohn T, Ting K. et al. Brief review of models of ectopic bone formation. Stem cells and development. 2012;21(5):655-67
30. Braun T, Gautel M. Transcriptional mechanisms regulating skeletal muscle differentiation, growth and homeostasis. Nature reviews Molecular cell biology. 2011;12(6):349-61
31. DiGirolamo DJ, Kiel DP, Esser KA. Bone and skeletal muscle: neighbors with close ties. Journal of bone and mineral research: the official journal of the American Society for Bone and Mineral Research. 2013;28(7):1509-18
32. Wajant H. The role of TNF in cancer. Results and problems in cell differentiation. 2009;49:1-15
33. Wang X, Lin Y. Tumor necrosis factor and cancer, buddies or foes? Acta pharmacologica Sinica. 2008;29(11):1275-88
34. Lin Y, Devin A, Rodriguez Y, Liu ZG. Cleavage of the death domain kinase RIP by caspase-8 prompts TNF-induced apoptosis. Genes & development. 1999;13(19):2514-26
35. Lejeune FJ, Lienard D, Matter M, Ruegg C. Efficiency of recombinant human TNF in human cancer therapy. Cancer immunity. 2006;6:6
36. Mori T, Sato Y, Miyamoto K, Kobayashi T, Shimizu T, Kanagawa H. et al. TNFalpha promotes osteosarcoma progression by maintaining tumor cells in an undifferentiated state. Oncogene. 2014;33(33):4236-41
37. Rutkowski P, Kaminska J, Kowalska M, Ruka W, Steffen J. Cytokine and cytokine receptor serum levels in adult bone sarcoma patients: correlations with local tumor extent and prognosis. Journal of surgical oncology. 2003;84(3):151-9
38. Liu C, Zhao P, Yang Y, Xu X, Wang L, Li B. Ampelopsin suppresses TNF-alpha-induced migration and invasion of U2OS osteosarcoma cells. Molecular medicine reports. 2016;13(6):4729-36
39. Zheng H, Min J. Role of Long Noncoding RNA HOTAIR in the Growth and Apoptosis of Osteosarcoma Cell MG-63. BioMed research international. 2016;2016:5757641
40. Pahl JH, Kwappenberg KM, Varypataki EM, Santos SJ, Kuijjer ML, Mohamed S. et al. Macrophages inhibit human osteosarcoma cell growth after activation with the bacterial cell wall derivative liposomal muramyl tripeptide in combination with interferon-gamma. Journal of experimental & clinical cancer research: CR. 2014;33:27
Author contact
Corresponding authors: Shumin Zhou: zhoushumin_zwcom, No.600, Yishan Road, Shanghai 200233, P.R. China or Qingcheng Yang: tjyqccom, No.600, Yishan Road, Shanghai 200233, P.R. China

 Global reach, higher impact
Global reach, higher impact