Impact Factor ISSN: 1449-1907
Int J Med Sci 2018; 15(6):610-616. doi:10.7150/ijms.24061 This issue Cite
Research Paper
Potential miRNA-target interactions for the screening of gastric carcinoma development in gastric adenoma/dysplasia
1. Department of Internal Medicine, Catholic Kwandong University, International St. Mary's Hospital, Incheon Metropolitan City, 404-834, Republic of Korea
2. Yonsei University College of Medicine, 50-Yonsei-ro, Seodaemun-gu, Seoul, Republic of Korea
3. Catholic Kwandong University, International St. Mary's Hospital, Incheon Metropolitan City, 404-834, Republic of Korea
4. Institute for Bio-Medical Convergence, College of Medicine, Catholic Kwandong University, Gangneung-si, Gangwon-do 210-701, Republic of Korea
5. Division of Gastroenterology, Department of Internal Medicine, Institute of Gastroenterology, Yonsei University College of Medicine, 50-Yonsei-ro, Seodaemun-gu, Seoul, Republic of Korea
Received 2017-11-28; Accepted 2018-3-1; Published 2018-3-14
Abstract
Although miRNA markers have been identified for the pathological development of gastric adenocarcinoma (GAC), the underlying molecule mechanism are still not fully understood. Moreover, some gastric adenoma/dysplasia may progress to GAC. In this study, the miRNA expression profiles in normal and paired low-/high-grade dysplasia were analyzed using Affymetrix Gene-Chip miRNA arrays. Of the total 2578 mature miRNA probe sets, ~1600 showed positive signals when the between normal and paired low-/high-grade dysplasia were compared. To verify the miRNA expression, qRT-PCR analysis was performed to quantify the expression of altered miRNAs between normal and paired low-/high-grade dysplasia. The analysis revealed that hsa-miR-421, hsa-miR-29b-1-5p, and hsa-miR-27b-5p were overexpressed in gastric low-/high-grade dysplasia and that based on these miRNA-target interactions, FBXO11 and CREBZF could be considered convincing markers for gastric cancer (GC) progression. Thus, we identified three miRNAs (hsa-miR-421, hsa-miR-29b-1-5p, and hsa-miR-27b-5p) with two mRNAs (FBXO11 and CREBZF) that might play an important role in the GC development from premalignant adenomas. Furthermore, these two target mRNAs and three miRNAs were predicted to be potential biomarkers for the progression of GC by miRNA-target interaction analysis.
Keywords: microRNA, gastric adenocarcinoma, hsa-miR-421, hsa-miR-29b-1-5p, hsa-miR-27b-5p
Introduction
Gastric cancer (GC) is one of the most common cancer types in the world. It is the fourth most common malignancy and the third leading cause of cancer mortality around the world [1-3]. Gastric adenomas may eventually develop into adenocarcinoma, which are the most common type of GCs; gastric adenocarcinoma (GAC) accounts for over 90% cases of all GC [2,3]. Previous studies have strongly suggested that high-grade dysplasia is highly predictive of invasive carcinoma, while the clinical significance of low-grade dysplasia is still unknown [4-7]. Several groups have studied adenomas as the intermediate step of gastric carcinoma development [2,8]. Moreover, numerous studies have identified oncogenes and tumor suppressor genes involved in the pathological development of GAC. However, the mechanism of genes is still not fully understood.
Small RNA molecules (19-23 nucleotide long) are also involved in tumor progression. MicroRNAs (miRNAs) are endogenous, non-coding single-stranded RNAs that play critical roles in the regulation of diverse biological processes, and miRNAs function as posttranscriptional gene regulators by binding to their target mRNAs [9,10]. Altered miRNA expression has been observed in various kinds of cancer, including breast cancer, hepatocellular carcinoma, colorectal cancer, and lung cancer [11-14]. Aberrant expression of miRNAs is associated with GAC development, and analyses of multiple parallel gene expression alterations are providing deeper insights into oncogenic transformation [15,16]. Therefore, different miRNAs might be involved in different pathological processes, and these miRNAs act as prognostic markers for cancer progression. miRNAs are considered as potential risk factors and are associated with an increased risk of cancer. They also pay a role in the formation and progression of GC but less is known about their role in premalignant adenomas.
Changes in the expression pattern of miRNAs can be informative and highly significant in the gastric adenoma-carcinoma sequence progression as well. However, precancerous tissues (such as dysplasia/adenoma) have been investigated less frequently than cancerous tissues [17-19]. The molecular characteristics of adenoma, especially of biopsy specimens, have not been fully elucidated.
The aim of the present study was to perform a miRNA microarray analysis of normal, low-grade, and high-grade dysplasia using fresh frozen tissues. Changes in miRNA expression patterns, in samples obtained from the same patients, were also verified using TaqMan MicroRNA Assays.
Materials and Methods
Clinical samples and tissue harvesting
Human tissue samples were obtained from 17 patients who underwent endoscopic submucosal dissection (ESD) at the Severance Hospital of Yonsei University. The study protocol was approved by the ethics review committee of the Institutional Review Board, College of Medicine, Yonsei University. Among these, one patient was dropped from this study because no tumor was found in the ESD specimen, as well as seven patients whose RNAs samples did not meet quality control standards for miRNA analysis. The basic information of six patients (low-grade dysplasia, n=3 and high-grade dysplasia, n=3) is shown in Table 1. GAC was confirmed by histopathological examination of tumor tissues after physical resection. The histological examination was performed by experienced pathologists without authorship in this study. Tumor grade was determined by two tier WHO classification: low-grade or high- grade dysplasia. For GAC, differentiation (well, moderate, poorly differentiated or signet ring cell carcinoma), depth of invasion, and presence or absence of lymphovascular or neural invasion were recorded.
Clinicopathological features of 6 patients.
| Patient No. | Gender | Age | Histologic diagnosis | Helicobacter pylori status |
|---|---|---|---|---|
| 1 | M | 77 | Low-grade dysplasia | Positive |
| 2 | M | 74 | Low-grade dysplasia | Negative |
| 3 | F | 57 | Low-grade dysplasia | Positive |
| 4 | M | 58 | High-grade dysplasia | Negative |
| 5 | M | 61 | High-grade dysplasia | Negative |
| 6 | M | 61 | High-grade dysplasia | Positive |
Isolation and quality check of total RNA including miRNA
Total RNA was extracted from 6 matched pairs of tissue samples (3 normal and 3 low grade dysplasia samples, 3 normal and 3 high-grade dysplasia samples) using the mirVana PARIS Kit (Ambion, USA) according to manufacturer's protocol. RNA purity and integrity were evaluated by ND-1000 Spectrophotometer (NanoDrop, Wilmington, USA), and Agilent 2100 Bioanalyzer (Agilent Technologies, Palo Alto, USA).
miRNA microarray
The miRNA expression profiles were analyzed using the Affymetrix Gene-Chip miRNA 4.0 array (Homo sapiens). Total RNA (1 ug) including miRNA from tissue was biotin-labeled using the FlashTagTM Biotin HSR RNA Labeling kit (Affymetrix, Genisphere, Hatfield, PA, USA). The samples were hybridized using GeneChip® Hybridization Oven to the Affymetrix miRNA microarray according to the protocols provided by the manufacturer. The labeled RNA was heated to 99°C for 5 minutes and then to 45°C for 5 minutes. RNA-array hybridization was performed with agitation at 60 rotations per minute for 16 hours at 48°C on an Affymetrix® 450 Fluidics Station. The chips were washed and stained using a Genechip Fluidics Station 450 (Affymetrix, Santa Clara, California, United States). The chips were then scanned with an Affymetrix GCS 3000 scanner (Affymetrix, Santa Clara, California, United States). Signal values were computed using the Affymetrix® GeneChip™ Command Console software.
qRT-PCR
The cDNA was reverse transcribed using TaqMan MicroRNA Reverse Transcription Kit (Applied Biosystems, Waltham, Massachusetts, USA). To determine miRNA expression levels, qRT-PCR was performed using TaqMan MicroRNA Assays (Applied Biosystems) according to the manufacturer's instruction. U6 snRNA was used as an internal control.
In silico miRNA-mRNA target prediction
Three miRNAs that were continuously up-regulated in adenoma progression (in normal vs. low-/high-grade adenoma) by TaqMan MicroRNA Assays were selected: hsa-miR-421, hsa-miR-29b-1-5p, and hsa-miR-27b-5p. Using miRNet web database (www.mirnet.ca/) we acquired pathway enrichment from gene ontology [20]. Using the KEGG pathway databases, we examined the pathway target enrichment (p<0.05) of selected groups of miRNAs.
Statistical analysis
Raw data were extracted automatically in Affymetrix data extraction protocol using the software provided by Affymetrix GeneChip® Command Console® Software (AGCC). The CEL files import, miRNA level RMA+DABG-All analysis and result export were performed using Affymetrix® Expression Console™ Software. Array data were filtered by probes annotated species. Comparative analysis was carried out between test and control samples using fold-change and independent T-test, in which the null hypothesis was that no difference exists among the 2 groups. False discovery rate (FDR) was controlled by adjusting the p value using Benjamini-Hochberg algorithm. All statistical tests and visualization of differentially expressed genes were conducted using R statistical language v. 3.1.2.
Results
Microarray analysis of miRNA expression in different patient groups
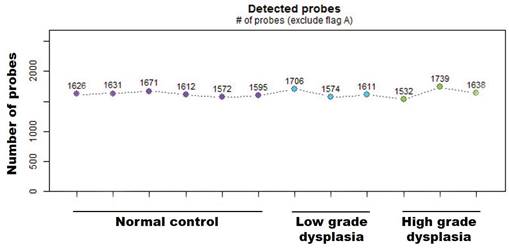
First, the absolute number of expressed miRNAs in the analyzed sample groups (normal and low-/high-grade dysplasia from matched pair tissues) was determined based on the intensity values of oligonucleotide probes for 2578 human mature miRNAs that were synthesized on the surface of GeneChip miRNA 4.0 arrays. Present values (based on hybridization) were calculated with the Expression Console Software (Affymetrix) using the statistical present/absent calls method. Of the 2578 mature miRNA probe sets, ~1600 showed positive values in all tissue samples (Fig. 1). There were no significant differences in detection probes between the diagnostic groups.
Number of miRNAs positive values from the mature miRNA probe sets in normal and low-/high-grade dysplasia on the Gene-Chip miRNA 4.0 array.
Discovery of expressed miRNA between normal and low-/high-grade dysplasia of patients
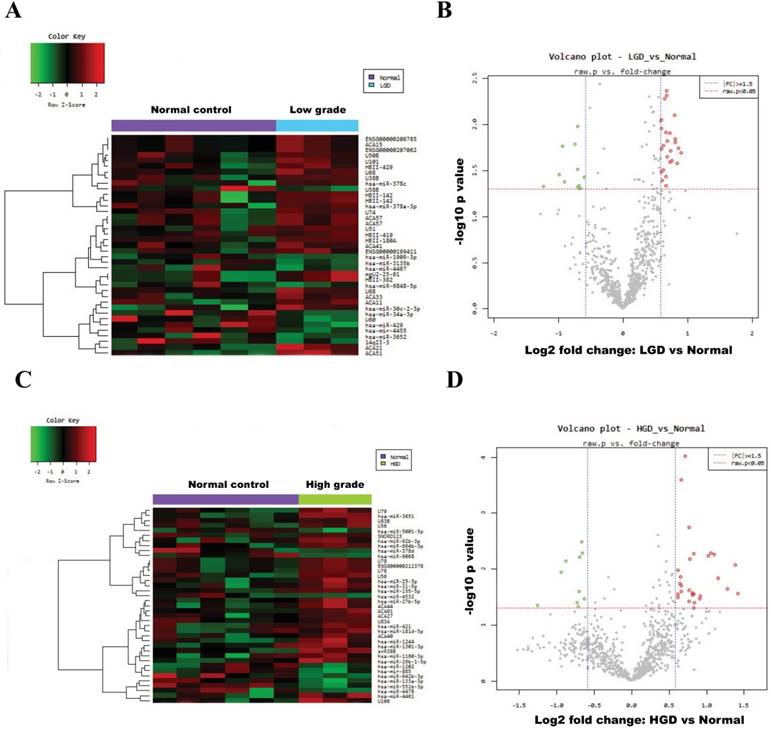
Fig. 2A and Fig. 2C show heatmaps of microarray data describing the miRNA expression profiles in normal vs. low-/high-grade dysplasia of patients. The volcano plot graph of the miRNA array shows that log 2 of the fold change in the expression of each miRNA between normal and low-/high-grade dysplasia is versus its -log 10 of P value from the t-test (Fig. 2B and 2D). The vertical blue line indicates that the threshold of fold change in miRNA expression is ≥1.5. The horizontal red line indicates that the threshold of P value of the t-test is 0.05. Up-regulated (red spots) and down-regulated (green spots) miRNAs which showed significantly different expression between control group and low-/high-grade dysplasia tissues (Fig. 3).
Hierarchical clustering heat map showing differential miRNA expression in normal vs low-/high-grade dysplasia and volcano plot graph of miRNA array results. (A) and (B) for normal vs low-grade dysplasia; (C) and (D) for normal vs high-grade dysplasia. The vertical blue line indicates that the threshold for fold change in miRNA expression is ≥1.5. The horizontal red line indicates that the threshold p value of the t-test is 0.05.
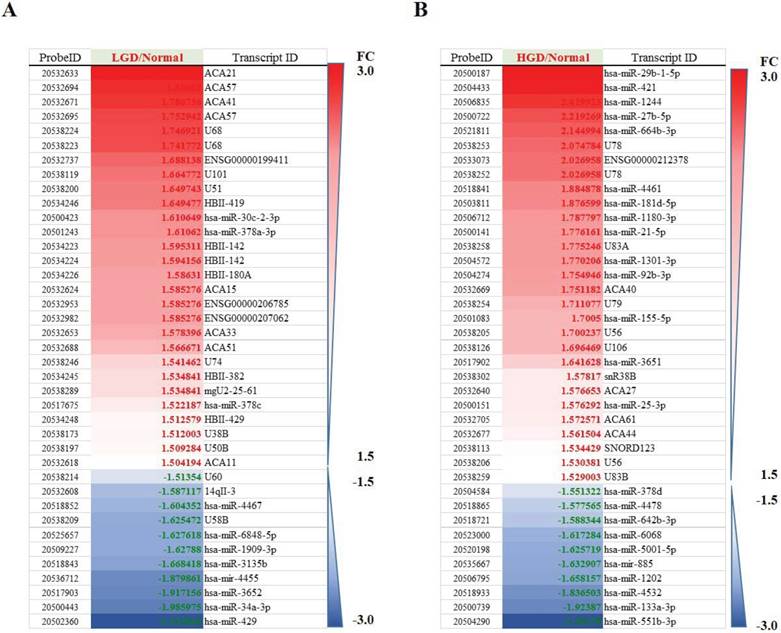
Differentially regulated miRNAs in tissues from normal group compared with low-grade dysplasia (A)/High-grade dysplasia (B).
miRNA microarray validation
In order to confirm the accuracy and reliability of the microarray data, the same tissue/RNA samples used in miRNA microarray analysis were analyzed using TaqMan MicroRNA Assays. qRT-PCR validation was done on four tissue samples pooled (with equal ng of RNA of the samples in each group) according to the analyzed diagnostic groups (normal, low-grade dysplasia, and high-grade dysplasia). miRNAs showing altered expression in normal and low-/high-grade dysplasia in the microarrays were selected, and the expression tendencies were detected by qRT-PCR (Fig. 4A). Three miRNAs (hsa-miR-421, hsa-miR-29b-1-5p, hsa-miR-27b-5p) were identified to be consistently upregulated in pathological progression from normal to low-/high-grade dysplasia in the qRT-PCR results.
Validation of miRNAs and miRNA-Target interaction. (A) Three miRNAs (miR-421, miR-29b-1-5p, miR-27b-5p) were selected as they were consistently up-regulated from normal to low-/high-grade dysplasia, (B) miRNA-target interactions and functional associations through network-based visual analysis, (C) miRNA-targets associated with pathways in cancer using KEGG pathway enrichment analysis. Significant differences between normal and low-/high-grade dysplasia were determined via ANOVA, with p values indicated as *p<0.05 and **p<0.01.
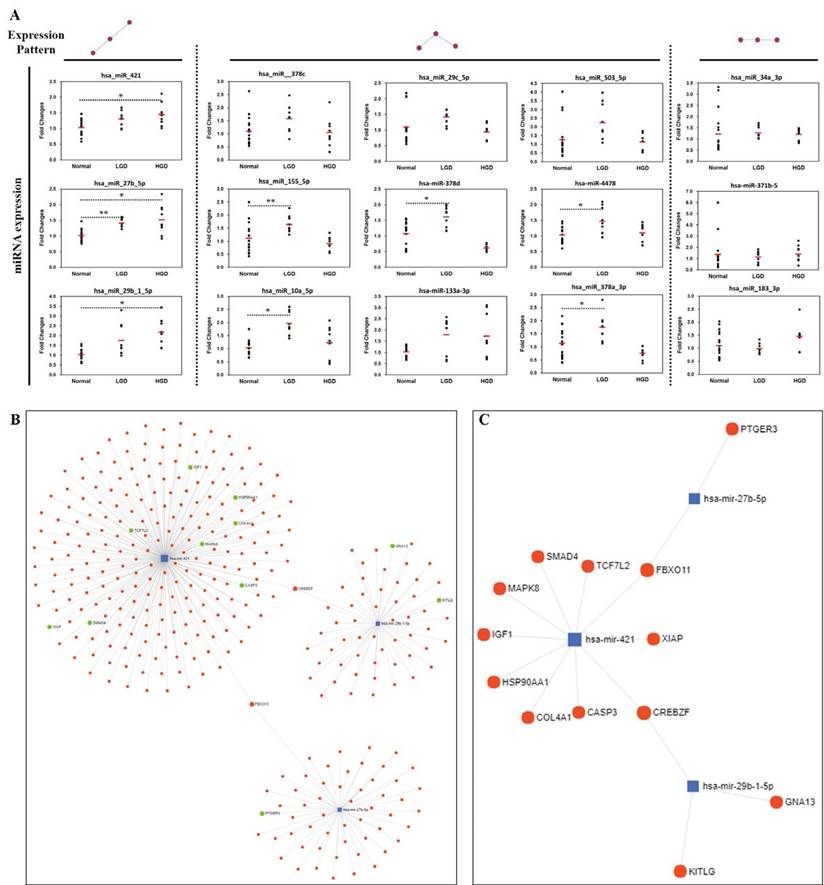
miRNA-target interaction and biological function/pathway analysis
Considering that miRNA can post-transcriptionally regulate the expression of target genes, we analyzed the miRNA-target interacttions and functional association through network-based visual analysis (miRNet) (Fig. 4B and 4C). Herein, we identified that two mRNAs (FBXO11 and CREBZF) along with their corresponding miRNAs, which were significantly associated with GC progression. Therefore, three miRNAs (hsa-miR-421, hsa-miR-29b-1-5p, and hsa-miR-27b-5p) with two mRNAs (FBXO11 and CREBZF) might play an important role in the development of GC from premalignant adenomas. According to the results of the miRNet network analysis, two targets and three miRNAs were predicted as potential biomarkers for the progression of GC. Moreover, KEGG pathway enrichment analysis revealed that the miRNA-targets were significantly associated with pathways in cancer, colorectal cancer, and focal adhesion (Table 2).
Functional association of KEGG databases of the selected miRNAs-target interaction genes.
| Name | Number of hit genes | P value |
|---|---|---|
| Pathways in cancer | 11 | 5.14e-12 |
| Colorectal cancer | 4 | 0.000256 |
| Focal adhesion | 4 | 0.0177 |
| Progesterone-mediated oocyte maturation | 3 | 0.0177 |
| Prostate cancer | 3 | 0.0177 |
| Toxoplasmosis | 3 | 0.0338 |
| FoxO signaling pathway | 3 | 0.0419 |
| Wnt signaling pathway | 3 | 0.0419 |
| Hepatitis B | 3 | 0.0419 |
| PI3K-Akt signaling pathway | 4 | 0.0423 |
Discussion
In this study, we have carried out a high throughput screening of microRNA expression alterations in normal and paired low-/high-grade dysplasia tissues using microarray analysis. We investigated the expression levels of 2578 mature miRNA in normal and low- and high-grade dysplasia tissues using GeneChip® miRNA 4.0 Arrays. Besides the previously described miRNAs between normal and gastric carcinoma lesions, we also identified new, lesser known miRNAs (hsa-miR-421, hsa-miR-29b-1-5p, and hsa-miR-27b-5p), with altered expression in differently graded dysplasia compared with normal pair-tissues.
A previous study using deep sequencing analysis, found that hsa-miR-421 is differentially expressed and plays important roles in carcinogenesis in human GC [21]. Moreover, hsa-miR-421 was detected in serum, and this miRNA was suggested to be a novel class of biomarker for diagnosis of cancer and other diseases [22]. However, new findings from this study demonstrated that hsa-miR-29b-1-5p and hsa-miR-27b-5p might be overexpressed in GC progression from adenoma/dysplasia. Based on the prediction in miRNet network analysis, two targets, FBXO11 and CREBZF, were identified as potential biomarkers for the progression of GC. Functionally, FBXO11 encodes a member of the F-box protein family and PKD1 phosphorylation-dependent degradation of SNAIL in epithelial-mesenchymal transition (EMP) and metastasis [23]. Previous results have demonstrated that FBXO11 complex also mediates ubiquitination and degradation of DTL, an important step for the regulation of TGF-beta signaling, cell migration and the timing of the cell-cycle progression and exit [24,25]. Another predicted target CREBZF is a transcription factor that target was suggested as a novel positive regulator of p53 [26]. Previous research identified that CREBZF activates transcription when bound to HCFC1 and suppresses the expression of HSV proteins in cells infected with the virus in an HCFC1-dependent manner [27]. Furthermore, CREBZF suppresses the HCF1-dependent transcriptional activation by CREB3 and reduces the amount of CREB3 in the cell [28].
Collectively, by integrating the results of miRNA expression alteration by GeneChip® miRNA 4.0 Arrays with TaqMan RT-PCR from adenoma/dysplasia, our data reveals that FBXO11 and CREBZF are closely related molecules for cancer development. Furthermore, correlation between miRNA expression and predicted target mRNA could be explored by functional analysis of GAC development related alterations, and this might serve as a valuable early diagnostic marker.
Abbreviations
EGC: early gastric cancer; GC: gastric cancer; GAC: gastric adenocarcinoma; HGD: high-grade dysplasia; LGD: low-grade dysplasia; miRNAs: microRNAs.
Acknowledgements
This study was supported by National Research Foundation of Korea (NRF-2014R1A1A3050247).
Authors' Contributions
Yu Jin Kim participated in the study design and drafted the manuscript. Ki-Chul Hwang conceived the study and drafted the manuscript. Sang Woo Kim and Yong Chan Lee are corresponding authors. All authors read and approved the final manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Lu Y, Tang L, Zhang Q, Zhang Z, Wei W. MicroRNA-613 inhibits the progression of gastric cancer by targeting CDK9. Artificial Cells, Nanomedicine, and Biotechnology. 2017:1-5
2. Taniyama D, Taniyama K, Kuraoka K, Zaitsu J, Saito A, Nakatsuka H. et al. Long-term follow-up study of gastric adenoma; tumor-associated macrophages are associated to carcinoma development in gastric adenoma. Gastric Cancer. 2017;20:929-939
3. Siegel R, Ma J, Zou Z, Jemal A. Cancer statistics, 2014. CA: Cancer Journal for Clinicians. 2014;64:9-29
4. Rugge M, Farinati F, Di Mario F, Baffa R, Valiante F, Cardin F. Gastric epithelial dysplasia: a prospective multicenter follow-up study from the Interdisciplinary Group on Gastric Epithelial Dysplasia. Human Pathology. 1991;22:1002-1008
5. Rugge M, Farinati F, Baffa R, Sonego F, Di Mario F, Leandro G. Gastric epithelial dysplasia in the natural history of gastric cancer: a multicenter prospective follow-up study. Interdisciplinary Group on Gastric Epithelial Dysplasia. Gastroenterology. 1994;107:1288-1296
6. Sipponen P. Gastric dysplasia. Current Topics in Pathology. 1990;81:61-76
7. Yamada H, Ikegami M, Shimoda T, Takagi N, Maruyama M. Long-term follow-up study of gastric adenoma/dysplasia. Endoscopy. 2004;36:390-396
8. Abozeid M, Rosato A, Sommaggio R. Immunotherapeutic Strategies for Gastric Carcinoma: A Review of Preclinical and Clinical Recent Development. BioMed Research International. 2017;2017:5791262
9. Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136:215-233
10. Shukla GC, Singh J, Barik S. MicroRNAs: Processing, Maturation, Target Recognition and Regulatory Functions. Molecular and Cellular Pharmacology. 2011;3:83-92
11. Zhu J, Zheng Z, Wang J, Sun J, Wang P, Cheng X. et al. Different miRNA expression profiles between human breast cancer tumors and serum. Frontiers in Genetics. 2014;5:149
12. Wang Y, Tian Y. miRNA for diagnosis and clinical implications of human hepatocellular carcinoma. Hepatology Research. 2016;46:89-99
13. Noguchi T, Toiyama Y, Kitajima T, Imaoka H, Hiro J, Saigusa S. et al. miRNA-503 Promotes Tumor Progression and Is Associated with Early Recurrence and Poor Prognosis in Human Colorectal Cancer. Oncology. 2016;90:221-231
14. Zhang S, Gao L, Thakur A, Shi P, Liu F, Feng J. et al. miRNA-204 suppresses human non-small cell lung cancer by targeting ATF2. Tumour biology. 2016;37:11177-11186
15. Cui L, Zhang X, Ye G, Zheng T, Song H, Deng H. et al. Gastric juice MicroRNAs as potential biomarkers for the screening of gastric cancer. Cancer. 2013;119:1618-1626
16. Konishi H, Ichikawa D, Komatsu S, Shiozaki A, Tsujiura M, Takeshita H. et al. Detection of gastric cancer-associated microRNAs on microRNA microarray comparing pre- and post-operative plasma. British Journal of Cancer. 2012;106:740-747
17. Zhang X, Peng Y, Jin Z, Huang W, Cheng Y, Liu Y. et al. Integrated miRNA profiling and bioinformatics analyses reveal potential causative miRNAs in gastric adenocarcinoma. Oncotarget. 2015;6:32878-32889
18. Hao NB, He YF, Li XQ, Wang K, Wang RL. The role of miRNA and lncRNA in gastric cancer. Oncotarget. 2017;8:81572-81582
19. Anauate AC, Leal MF, Wisnieski F, Santos LC, Gigek CO, Chen ES. et al. Identification of suitable reference genes for miRNA expression normalization in gastric cancer. Gene. 2017;621:59-68
20. Fan Y, Siklenka K, Arora SK, Ribeiro P, Kimmins S, Xia J. miRNet - dissecting miRNA-target interactions and functional associations through network-based visual analysis. Nucleic Acids Research. 2016;44:W135-W141
21. Guo J, Miao Y, Xiao B, Huan R, Jiang Z, Meng D, Wang Y. Differential expression of microRNA species in human gastric cancer versus non-tumorous tissues. Journal of Gastroenterology and Hepatology. 2009;24:652-657
22. Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K. et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Research. 2008;18:997-1006
23. Zheng H, Shen M, Zha YL, Li W, Wei Y, Blanco MA. et al. PKD1 phosphorylation-dependent degradation of SNAIL by SCF-FBXO11 regulates epithelial-mesenchymal transition and metastasis. Cancer Cell. 2014;26:358-373
24. Abbas T, Mueller AC, Shibata E, Keaton M, Rossi M, Dutta A. CRL1-FBXO11 promotes Cdt2 ubiquitylation and degradation and regulates Pr-Set7/Set8-mediated cellular migration. Molecular Cell. 2013;49:1147-1158
25. Abbas T, Keaton M, Dutta A. Regulation of TGF-beta signaling, exit from the cell cycle, and cellular migration through cullin cross-regulation: SCF-FBXO11 turns off CRL4-Cdt2. Cell Cycle. 2013;12:2175-2182
26. Lopez-Mateo I, Villaronga MA, Llanos S, Belandia B. The transcription factor CREBZF is a novel positive regulator of p53. Cell Cycle. 2012;11:3887-3895
27. Lu R, Misra V. Zhangfei: a second cellular protein interacts with herpes simplex virus accessory factor HCF in a manner similar to Luman and VP16. Nucleic Acids Research. 2000;28:2446-2454
28. Misra V, Rapin N, Akhova O, Bainbridge M, Korchinski P. Zhangfei is a potent and specific inhibitor of the host cell factor-binding transcription factor Luman. Journal of Biological Chemistry. 2005;280:15257-15266
Author contact
Corresponding authors: Sang Woo Kim, Catholic Kwandong University, International St. Mary's Hospital, Incheon Metropolitan City, 404-834, Republic of Korea. Tel: +82-32-290-2612, Fax: +82-32-290-2774, E-mail: ksw74ac.kr (S.W. Kim) and Yong Chan Lee, Department of Internal Medicine, Yonsei University College of Medicine 50-1 Yonsei-ro, Seodaemun-gu, Seoul 03722, Korea. Tel: +82-2-2228-1960, Fax: +82-2-393-6884, E-mail: leeycac (Y.C. Lee)

 Global reach, higher impact
Global reach, higher impact