ISSN: 1449-1907
Int J Med Sci 2017; 14(6):578-584. doi:10.7150/ijms.15699 This issue Cite
Research Paper
Hypoxia-regulated MicroRNA-210 Overexpression is Associated with Tumor Development and Progression in Upper Tract Urothelial Carcinoma
1. Graduate Institute of Medicine, College of Medicine, Kaohsiung Medical University, Kaohsiung, Taiwan;
2. Department of Urology, School of Medicine, College of Medicine, Kaohsiung Medical University, Kaohsiung, Taiwan;
3. Department of Urology, Kaohsiung Medical University Hospital, Kaohsiung Medical University, Kaohsiung, Taiwan;
4. Department of Urology, Ministry of Health and Welfare Pingtung Hospital, Pingtung, Taiwan;
5. Department of Urology, Kaohsiung Municipal Ta-Tung Hospital, Kaohsiung, Taiwan;
6. Department of Microbiology, School of Medicine, College of Medicine, Kaohsiung Medical University, Kaohsiung, Taiwan;
7. Department of Pathology, Chi Mei Medical Center, Tainan, Taiwan;
8. National Institute of Cancer Research, National Health Research Institutes, Tainan, Taiwan;
9. Department of Biotechnology, Southern Taiwan University of Science and Technology, Tainan, Taiwan.
Received 2016-3-30; Accepted 2016-11-15; Published 2017-5-11
Abstract
Background: Hypoxia has been shown to facilitate tumor progression. Hypoxia-regulated microRNA-210 (miR-210) may play an important role in carcinogenesis and tumor progression. In this study, we evaluated the clinical significance of miR-210 expression in upper tract urothelial carcinoma (UTUC).
Methods: Eighty-three UTUC patients participated in this study. All of them provided cancer tissue samples and 50 of them provided non-cancerous urothelium samples. Clinicopathologic data were collected by reviewing medical records. The expression of miR-210 and hypoxia-inducible factor-1α (HIF-1α) was determined by quantitative real-time polymerase chain reaction. The relationship between clinicopathologic variables and the expression of miR-210 and HIF-1α was analyzed statistically.
Results: MiR-210 is overexpressed in UTUC compared to non-cancerous urothelium (p < 0.001); it is also upregulated in high-stage and high-grade tumors (p = 0.020 and 0.049, respectively). HIF-1α is overexpressed in UTUC and correlates positively with miR-210 expression (r = 0.442, p = 0.001).
Conclusion: Both miR-210 and HIF-1α are involved in promoting UTUC carcinogenesis. MiR-210 is also correlated with tumor progression. Further studies are needed to clarify the underlying mechanism.
Keywords: Upper tract urothelial carcinoma, HIF-1α, miR-210.
Introduction
Upper tract urothelial carcinoma (UTUC) includes renal pelvis cancer and ureter cancer. Radical nephroureterectomy with bladder cuff excision is the standard treatment for UTUC [1, 2]. The incidence of bladder cancer recurrence within five years after nephroureterectomy is 20-75% [3]. High bladder cancer recurrence and distant metastases are the major causes of complications after primary nephroureterectomy [1, 2]. Therefore, UTUC patients should receive periodic cystoscopy follow-ups. The pathologic stage of the primary tumor, lymph node status, presence of distant metastases, and tumor grade are important prognostic factors [2, 4]. Five-year survival rates of patients with stage Ta-T1, T2, T3, and T4 tumors were 60-90%, 43-75%, 16-33%, and 0-4% respectively [5]. Disease progression and tumor metastasis are commonly observed after surgery. Identifying novel biomarkers to facilitate clinical prognosis and therapeutic intervention is an important clinical issue.
Hypoxia is the condition of inadequate oxygen in the tissue. Hypoxia is one of the most important environmental factors that induce cancer metastasis [6]. Hypoxia also plays an important role in metabolism, angiogenesis, innate immunity, and induction of stemness [7]. In response to hypoxia stress, cells alter DNA transcription in conjunction with the oxygen-monitoring machinery, which includes hypoxia-inducible factors (HIFs) that are the major components of hypoxia signaling pathways [8]. HIF-1 is a heterodimer, composed of one oxygen-sensitive subunit, HIF-1α, and one oxygen-insensitive subunit, HIF-1β. In hypoxic tumor microenvironments, the oxygen-dependent HIF-1α degradation pathway is halted, resulting in elevated levels of HIF-1α. More than 1000 target genes, including several microRNAs (miRNAs), are regulated by HIFs to mediate the effects induced by hypoxia [9].
MiRNAs are relatively small, 18-24 nucleotide long, noncoding RNAs. MiRNAs commonly regulate gene expression negatively by repressing translation or directing sequence-specific degradation of complementary mRNAs [10]. A single miRNA can bind to as many as 200 mRNA targets and function as either a tumor suppressor or an oncogene, depending on the characteristics of the target gene [11]. They regulate diverse biological processes and potentially influence almost all genetic pathways [11]. MiR-210, which is located on chromosome 11p15.5, is overexpressed in many human cancers [12-14]. MiR-210 overexpression, specifically under hypoxia, affects many processes involved in tumor development, including the promotion of angiogenesis and a reduction in DNA repair capabilities [13, 15].
Previously, we have shown that HIF-1α overexpression plays an important role in predicting prognosis in UTUC [16]. Here, we compared miR-210 expression between UTUC and paired non-cancerous urothelium to clarify the role of miR-210 in UTUC. To our knowledge, the role of miR-210 in UTUC has not been reported yet. This is the first study to evaluate systematically the expression of miR-210 in UTUC samples.
Materials and methods
Surgical specimens and clinicopathologic data
Eighty-three patients with UTUC who received nephroureterectomy in Kaohsiung Medical University Hospital from 2013 to 2015, participated in this study. All of them provided cancerous urothelium samples and 50 of them provided paired non-cancerous urothelium samples. Clinicopathologic data were collected by reviewing medical records. Both cancerous and non-cancerous urothelium samples were confirmed by a clinical pathologist. All clinical samples were obtained with informed consent from each subject and the protocol for this study was reviewed and approved by the Institutional Review Board of Kaohsiung Medical University Hospital (KMUH-IRB-20130369 and KMUH-IRB-20130089).
RNA extraction and complementary DNA (cDNA) preparation
Tissues were immediately frozen in liquid nitrogen at the time of surgery and stored at -80°C. Total RNA, including mRNA and miRNA, was extracted from frozen tissues using the Quick-RNATM MiniPrep Kit (Zymo Research Corp., Irvine, CA, USA), according to the manufacturer's protocol. The concentration and purity of the total RNA were evaluated using an ultraviolet spectrophotometer. For the individual miRNA assays, cDNA of the miRNA was synthesized using a unique primer (miR-210-specific stem-loop primers) by using 20 ng of the total RNA. For the mRNA quantitative assay, cDNA was synthesized from 1 μg of total RNA with random hexamer primers by using reverse transcriptase (Applied Biosystems, Waltham, MA, USA). All the cDNA samples were stored at -20°C until analysis.
MicroRNA profile analyses
MicroRNA profile analyses of the BFTC-909 cell line (obtained from Bioresource Collection and Research Center, Hsinchu, Taiwan) and one pair of UTUC samples (comprising cancerous and non-cancerous urothelium from the same patient) were performed. RNA was reverse transcribed using TaqMan Megaplex Primer Pools A and B mix and TaqMan Reverse Transcription Kit (Applied Biosystems, Waltham, MA, USA). The miRNA expression profiles were assayed on the TaqMan Array Human MicroRNA A + B Cards Set (Applied Biosystems, Waltham, MA, USA). qRT- PCR was performed on 7500HT Fast Real Time PCR System (Applied Biosystems, Waltham, MA, USA). According to this miRNA expression profile, we found miR-210 to be the most promising candidate and consequently our further studies focused only on miR-210.
Quantitative real-time PCR (qRT-PCR) for miR-210 and HIF-1α expression
For miR-210 analysis, the TaqMan miRNA qRT-PCR assay (Applied Biosystems, Waltham, MA, USA) was used to quantify the levels of miR-210. The qRT-PCR was performed at 95°C for 20 s, followed by 40 cycles of 95°C for 5 s and 60°C for 35 s. The relative expression level of the miRNA was normalized to that of the internal control, RNU6B, by using the equation log (2-ΔCt), where ΔCt = (CtmiR-210 - Ct RNU6B) [17].
For HIF-1α analysis, cDNA was synthesized using reverse transcriptase (Applied Biosystems, Waltham, MA, USA). HIF-1α expression levels were quantified using qRT-PCR, which was performed using Fast SYBR Green Master Mix (Applied Biosystems, Waltham, MA, USA) with specific oligonucleotide primers (HIF-1α primers: Forward TGTGACCATGAGGAAATGAGAGA; Reverse TTTTGTTCTTTACCCTTTTTCACAAG; GAPDH was used as the internal control, Forward: CCTGCACCACCAACTGCTTA; Reverse: GGGCCATCCACAGTCTTCTG). Expression was analyzed using 7500HT Real-Time PCR System (Applied Biosystems, Waltham, MA, USA). The fold change in HIF-1α expression in cancerous vs. non-cancerous urothelium was calculated by the comparative threshold count method, after normalization to the internal control, GAPDH. All experiments were performed in triplicate.
Statistical analysis
All statistical analyses were performed using the SPSS 20.0 statistical package (IBM Corp., NY, USA) and SigmaPlot (Systat Software, San Jose, CA, USA). MiR-210 expression has been presented as the median and interquartile range due to its failure to pass the normality test of distribution. Wilcoxon signed-rank test was used to compare the differences in miR-210 expression between UTUC and paired non-cancerous urothelium, and Mann-Whitney U tests were used for comparisons between different age, gender, stage, and grade groups. Student's t test was used to compare HIF-1α expression levels in UTUC and paired non-cancerous urothelium. We used Receiver Operating Characteristic (ROC) curve analysis to test the accuracy of distinguishing UTUC from normal urothelium based on miR-210 and HIF-1α expression levels. The ROC curves are produced by calculating the sensitivity and specificity at different thresholds. The ROC curve is a graphical representation of test characteristics, with sensitivity on the y-axis and 1-specificity on the x-axis, over all possible cut-off points for defining a positive and a negative test result. In this study, the result was considered positive (UTUC) when the probability lies above a cut-off point. P values less than 0.05 were considered statistically significant.
Results
MiR-210 is the candidate miRNA in microRNA array profile
In order to search the candidate miRNAs of interest in the Taiwanese population with UTUC, we used the ABI TaqMan microRNA array to compare the miRNA profiles of BFTC-909 cell line, UTUC tissue and paired non-cancerous urothelium from the same patient. MiR-210 was found to be highly expressed in BFTC-909 cell line, and UTUC tissue, compared to that in paired non-cancerous urothelium (data not shown); this suggests that miR-210 may be involved in UTUC carcinogenesis.
MiR-210 and HIF-1α expression in human UTUC tissues
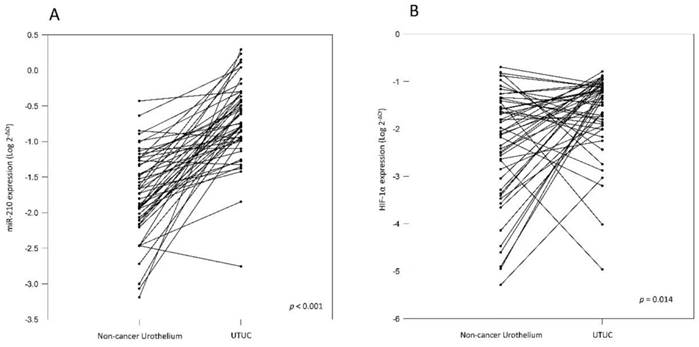
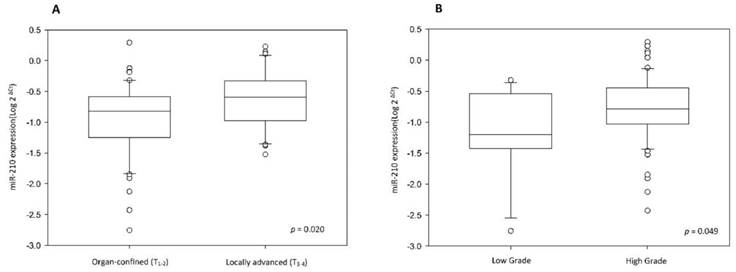
To validate miR-210 expression, we analyzed cancer tissue samples from 83 UTUC patients, 50 of whom also provided paired non-cancerous urothelium. The correlation between clinicopathologic features and miR-210 expression was analyzed and it has been summarized in Table 1. The results show that miR-210 expression is significantly upregulated in UTUC tissues than in non-cancerous urothelium (Table 1 and Fig. 1A, p < 0.001). There are no significant associations between miR-210 expression and patients' gender and age groups (p = 0.181 and 0.639, respectively). Interestingly, we also found that miR-210 showed significant overexpression in high-stage and high-grade tumors (Table 1 and Fig. 2, p = 0.020 and 0.049, respectively). These data support the hypothesis that miR-210 may act as an important oncogenic factor in UTUC development. Since miR-210 is a hypoxia-related miRNA, we also examined HIF-1α expression in UTUC and non-cancerous urothelium. HIF-1α was found to be significantly overexpressed in UTUC than in paired non-cancerous urothelium (Table 1 and Fig. 1B, p = 0.014). There are no significant associations between HIF-1α expression and patients' gender and age groups (p = 0.078 and 0.885, respectively). We also found that HIF-1α showed significant overexpression in high-stage tumors but not in high-grade tumors (p = 0.015 and 0.265, respectively). These results suggest that HIF-1α is involved in UTUC development.
Relationship between miR-210 expression and clinicopathologic features in UTUC patients.
| Clinicopathologic feature | n | MiR-210 relevant expression(2-ΔCt) | HIF-1α relevant expression(2-ΔCt) | |||
|---|---|---|---|---|---|---|
| Median (25%~75%) | P value$ | Mean ± standard deviation | P value* | |||
| Tissue | UTUC | 50 | 0.176 (0.104~0.442) | <0.001 | 0.053 ± 0.043 | 0.014 |
| Non-cancerous urothelium | 50 | 0.020 (0.008~0.059) | 0.029 ± 0.046 | |||
| Gender | Male | 35 | 0.138 (0.043~0.387) | 0.181 | 0.057 ± 0.053 | 0.078 |
| Female | 48 | 0.181 (0.094~0.326) | 0.040 ± 0.035 | |||
| Age | < 70 | 42 | 0.153 (0.070~0.396) | 0.639 | 0.048 ± 0.042 | 0.885 |
| ≥ 70 | 41 | 0.163 (0.064~0.286) | 0.046 ± 0.046 | |||
| Stage | Organ-confined (T1-2) | 50 | 0.151 (0.056~0.259) | 0.020 | 0.037 ± 0.041 | 0.015 |
| Locally advanced (T3-4) | 33 | 0.257 (0.106~0.467) | 0.061 ± 0.045 | |||
| Grade | Low | 11 | 0.063 (0.038~0.290) | 0.049 | 0.033 ± 0.036 | 0.265 |
| High | 72 | 0.163 (0.093~0.356) | 0.049 ± 0.045 | |||
$Wilcoxon signed-rank test *Paired-t test
(A) Comparison of miR-210 levels in 50 pairs of UTUC and the corresponding non-cancerous urothelium. The expression level of miR-210 is significantly higher in UTUC than in non-cancerous urothelium (B) Comparison of HIF-1α levels in 50 pairs of UTUC and the corresponding non-cancerous urothelium. The expression level of HIF-1α is significantly higher in UTUC than in non-cancerous urothelium.
(A) MiR-210 expression level is higher in locally advanced UTUC than in organ confined UTUC. (B) MiR-210 expression level is higher in high-grade UTUC than in low-grade UTUC.
(A) The ROC curve analysis for miR-210 expression to calculate the best cut-off point that discriminates between UTUC and non-cancerous urothelium. The area under curve (AUC) is 0.904 (95% CI = 0.843-0.865, p < 0.001). (B) The ROC curve analysis for HIF-1α expression. The AUC is 0.701 (95% CI = 0.597-0.805, p = 0.001).
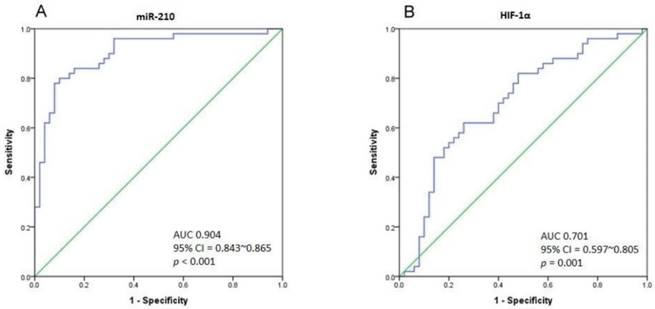
ROC curve analysis for distinguishing UTUC from non-cancerous urothelium
As there is a significant difference in miR-210 expression between UTUC and non-cancerous urothelium, we performed Receiver Operating Characteristic (ROC) curve analysis to determine the best cut-off point of miR-210 expression with the highest sensitivity and specificity. The ROC curve is shown in Fig. 3A. The area under curve (AUC) is 0.904 (95% CI = 0.843-0.865, p < 0.001). Taking 0.1001 to be the cut-off point of miR-210 expression, we could gain a sensitivity of 80% and specificity of 90% in distinguishing UTUC from non-cancerous urothelium. The AUC of the ROC curve in HIF-1α analysis is 0.701 (95% CI = 0.597-0.805, p = 0.001), and the sensitivity and specificity are 62% and 74% respectively, with the cut-off value of 0.0299 (Fig. 3B). Importantly, miR-210 and HIF-1α expression are found to correlate positively (Pearson Correlation, r = 0.442, p = 0.001).
Discussion
From our TaqMan microRNA array data, we identified miR-210 to be an oncogenic factor in UTUC. MiR-210 was validated in 50 pairs of UTUC and adjacent non-cancerous urothelium and was found to be significantly upregulated in UTUC compared to adjacent non-cancerous urothelium. We also found miR-210 overexpression in high-stage and high-grade UTUC. In gastric cancer, the level of its expression is closely related to tumor size, depth of invasion, lymph node metastasis, distant metastasis, and TNM stage [13]. MiR-210 is strongly upregulated in non-small cell lung cancer compared to normal tissue, and is associated with tumor stage [18]. However, Zaravinos et al. showed equal expression of miR-210 in bladder cancer and normal urothelium, but they also found miR-210 to be an independent prognostic factor for overall survival [19]. Takikawa et al. reported that the inhibition of miR-210 expression decreased migration and the expression of vimentin and snai1, and increased the membrane-associated expression of β-catenin in Panc-1 cells (pancreatic cancer cell line) co-cultured with pancreatic stellate cells [20]. Our results are in agreement with the above-mentioned published results and suggest that miR-210 may play an important role in promoting UTUC carcinogenesis.
The AUC of the ROC curve suggests that the miR-210 levels can be used to distinguish UTUC from normal urothelium, with high accuracy. We found that by taking 0.1001 to be the cut-off point of miR-210 expression, we could gain a sensitivity of 80% and specificity of 90% to discriminate UTUC from adjacent non-cancerous urothelium. This may help pathologists to identify precancerous lesions before histological characteristics are manifested. Besides, serum miR-210 level has been found to be upregulated in bladder cancer patients thus implying that it can potentially be used to predict bladder cancer recurrence [21]. However, even after undergoing radical surgery, UTUC patients with a high pT stage show poor clinical outcomes owing to high local tumor recurrence and distant metastases. It is of utmost importance to identify novel targets for UTUC that may not only serve as prognostic adjuncts but also aid therapeutic intervention. In this study, we have not measured serum miR-210 levels. Further studies on serum miR-210 levels in UTUC patients should be conducted to develop a reliable biomarker for pre-operative diagnosis.
Interestingly, the role of miR-210 in the regulation of hypoxia-induced pathogenesis is well-documented; it is known to be mediated by HIF-1α through the hypoxia-responsive element located on the proximal promoter of the miR-210-coding DNA [22]. Hypoxia, a hallmark of all solid tumors, is strongly associated with malignant progression and poor clinical outcome [23]. Previously, we found that tumor HIF-1α expression is significantly correlated with tumor T stage, N stage, and grade. HIF-1α overexpression is a significant predictor for cancer-specific survival and tumor recurrence [16]. Recently, hypoxia-regulated miRNAs have been researched in many cancer types. Among these miRNAs, miR-210 is the only one that is consistently upregulated in both normal and hypoxic tumor cells; it is generally recognized as a robust HIF target [24]. Here, we show that miR-210 and HIF-1α expression are positively correlated (Pearson Correlation, r = 0.442, p = 0.001) in UTUC.
MiR-210 downregulation significantly suppresses clonogenicity, migration, and invasion, as well as induces cell apoptosis, increases the percentage of cells in the G1 phase, and decreases the percentage of cells in the S phase in vitro. These results indicate that miR-210 downregulation significantly suppresses tumor growth in nude mouse models [25]. In the immune response, miR-210 decreases the susceptibility of tumor cells to lysis by cytotoxic T lymphocytes (CTLs). Moreover, the coordinate silencing of PTPN1, HOXA1, and TP53 dramatically decreases tumor cell susceptibility to CTL-mediated lysis, thus supporting their primary role in mediating the immunosuppressive effects of miR-210 [26]. However, the effect of miR-210 may depend on the cancer type; although its high-level expression is an independent prognostic factor in breast cancer [27], it is frequently deleted in epithelial ovarian cancer [28]. Thus, its role needs to be investigated in individual types of cancer. The oncogenic character of miR-210 is due to its targets in several carcinogenesis pathways. MiR-210 targets MNT (Max's Next Tango) and E2F3 (E2F transcription factor 3), which are crucial factors in cell cycle progression. MiR-210 also binds to 3'UTR of RAD52 mRNA and interferes in DNA repair via the HDR (homology-dependent repair) pathway. In HIF-VEGF pathway, miR-210, induced by HIF, targets on EFNA3 (ephrin A3) and PTP1B (phospho-tyrosine phosphatase-1B), which are negative regulators of VEGF [8]. Further functional analysis of miR-210 and identification of its target molecules in UTUC carcinogenesis are essential to identify novel prognostic markers or targets for treatment.
Finally, this study has several limitations. First, due to the inadequate follow-up period, only few cases of bladder cancer recurrence or cancer-specific death were observed. Therefore, we could not conclude a significant effect of miR-210 on survival. Second, as we did not collect patients' blood samples, we could neither test the miR-210 serum levels nor isolate circulating tumor cells. Hence, further evaluation after a long-term follow-up is needed. Third, we used only one cell line and one pair of UTUC and adjacent non-cancerous urothelium for miRNA screening. The significant miRNAs identified in this discovery stage might not have adequate power, but we later used 50 pairs of UTUC and adjacent non-cancerous urothelium for validation to minimize the possibility of false positivity.
Conclusion
Our study demonstrates that miR-210 is overexpressed in UTUC compared to non-cancerous urothelium. Moreover, miR-210 is upregulated in high-stage and high-grade tumors. This suggests that miR-210 may be involved in carcinogenesis and tumor progression pathways in UTUC. MiR-210 and its target mRNAs are potential candidates for UTUC treatment. Further research is necessary to clarify the underlying mechanisms.
Abbreviations
UC: urothelial carcinoma; UTUC: upper tract urothelial carcinoma; HIF-1α: hypoxia-inducible factor-1α; miRNA: microRNA; miR: microRNA.
Acknowledgements
This study was supported by grants from Kaohsiung Medical University “Aim for the Top Universities” (KMU-TP104E31, KMU-TP105G00, KMU-TP105G01, KMU-TP105G02), the health and welfare surcharge of tobacco products, Ministry of Health and Welfare (MOHW105-TDU-B-212-134007), Ministry of Science and Technology (MOST103-2314-B-037-059-MY3), and Kaohsiung Medical University Hospital (KMUH102-2M41, KMUH102-2R40, KMUH101-1R46).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Roupret M, Babjuk M, Comperat E, Zigeuner R, Sylvester R, Burger M. et al. European guidelines on upper tract urothelial carcinomas: 2013 update. Eur Urol. 2013;63:1059-71
2. Roupret M, Zigeuner R, Palou J, Boehle A, Kaasinen E, Sylvester R. et al. European guidelines for the diagnosis and management of upper urinary tract urothelial cell carcinomas: 2011 update. Eur Urol. 2011;59:584-94
3. Chou YH, Huang CH. Unusual clinical presentation of upper urothelial carcinoma in Taiwan. Cancer. 1999;85:1342-4
4. Li WM, Shen JT, Li CC, Ke HL, Wei YC, Wu WJ. et al. Oncologic outcomes following three different approaches to the distal ureter and bladder cuff in nephroureterectomy for primary upper urinary tract urothelial carcinoma. Eur Urol. 2010;57:963-9
5. Smith A, Matin S, Jarrett T. Urothelial Tumors of the Upper Urinary Tract and Ureter. In: (ed.) Wein A, Kavoussi L, Partin A, Peters C. Campbell-Walsh Urology; 11 ed. Elsevier. 2016:1374
6. Tsai YP, Wu KJ. Hypoxia-regulated target genes implicated in tumor metastasis. J Biomed Sci. 2012;19:102
7. Majmundar AJ, Wong WJ, Simon MC. Hypoxia-inducible factors and the response to hypoxic stress. Mol Cell. 2010;40:294-309
8. Dang K, Myers KA. The role of hypoxia-induced miR-210 in cancer progression. Int J Mol Sci. 2015;16:6353-72
9. Semenza GL. Hypoxia-inducible factors: mediators of cancer progression and targets for cancer therapy. Trends Pharmacol Sci. 2012;33:207-14
10. Wightman B, Ha I, Ruvkun G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell. 1993;75:855-62
11. Esquela-Kerscher A, Slack FJ. Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer. 2006;6:259-69
12. Valera VA, Walter BA, Linehan WM, Merino MJ. Regulatory Effects of microRNA-92 (miR-92) on VHL Gene Expression and the Hypoxic Activation of miR-210 in Clear Cell Renal Cell Carcinoma. J Cancer. 2011;2:515-26
13. Yu P, Fan S, Huang L, Yang L, Du Y. MIR210 as a potential molecular target to block invasion and metastasis of gastric cancer. Med Hypotheses. 2015;84:209-12
14. Ho AS, Huang X, Cao H, Christman-Skieller C, Bennewith K, Le QT. et al. Circulating miR-210 as a Novel Hypoxia Marker in Pancreatic Cancer. Transl Oncol. 2010;3:109-13
15. Ivan M, Huang X. miR-210: fine-tuning the hypoxic response. Adv Exp Med Biol. 2014;772:205-27
16. Ke HL, Wei YC, Yang SF, Li CC, Wu DC, Huang CH. et al. Overexpression of hypoxia-inducible factor-1alpha predicts an unfavorable outcome in urothelial carcinoma of the upper urinary tract. Int J Urol. 2008;15:200-5
17. Yang IP, Tsai HL, Hou MF, Chen KC, Tsai PC, Huang SW. et al. MicroRNA-93 inhibits tumor growth and early relapse of human colorectal cancer by affecting genes involved in the cell cycle. Carcinogenesis. 2012;33:1522-30
18. Tafsiri E, Darbouy M, Shadmehr MB, Zagryazhskaya A, Alizadeh J, Karimipoor M. Expression of miRNAs in non-small-cell lung carcinomas and their association with clinicopathological features. Tumour Biol. 2015;36:1603-12
19. Zaravinos A, Radojicic J, Lambrou GI, Volanis D, Delakas D, Stathopoulos EN. et al. Expression of miRNAs involved in angiogenesis, tumor cell proliferation, tumor suppressor inhibition, epithelial-mesenchymal transition and activation of metastasis in bladder cancer. J Urol. 2012;188:615-23
20. Takikawa T, Masamune A, Hamada S, Nakano E, Yoshida N, Shimosegawa T. miR-210 regulates the interaction between pancreatic cancer cells and stellate cells. Biochem Biophys Res Commun. 2013;437:433-9
21. Yang Y, Qu A, Liu J, Wang R, Liu Y, Li G. et al. Serum miR-210 Contributes to Tumor Detection, Stage Prediction and Dynamic Surveillance in Patients with Bladder Cancer. PLoS One. 2015;10:e0135168
22. Chan YC, Banerjee J, Choi SY, Sen CK. miR-210: the master hypoxamir. Microcirculation. 2012;19:215-23
23. Harris AL. Hypoxia-a key regulatory factor in tumour growth. Nat Rev Cancer. 2002;2:38-47
24. Devlin C, Greco S, Martelli F, Ivan M. miR-210: More than a silent player in hypoxia. IUBMB Life. 2011;63:94-100
25. Liu C, Tang X. Downregulation of microRNA-210 inhibits osteosarcoma growth in vitro and in vivo. Mol Med Rep. 2015;12:3674-80
26. Noman MZ, Buart S, Romero P, Ketari S, Janji B, Mari B. et al. Hypoxia-inducible miR-210 regulates the susceptibility of tumor cells to lysis by cytotoxic T cells. Cancer Res. 2012;72:4629-41
27. Camps C, Buffa FM, Colella S, Moore J, Sotiriou C, Sheldon H. et al. hsa-miR-210 Is induced by hypoxia and is an independent prognostic factor in breast cancer. Clin Cancer Res. 2008;14:1340-8
28. Giannakakis A, Sandaltzopoulos R, Greshock J, Liang S, Huang J, Hasegawa K. et al. miR-210 links hypoxia with cell cycle regulation and is deleted in human epithelial ovarian cancer. Cancer Biol Ther. 2008;7:255-64
Author contact
![]() Corresponding author: Wen-Jeng Wu, Department of Urology, Kaohsiung Medical University Hospital, Kaohsiung Medical University, No. 100, TzYou 1st Road, Kaohsiung City 807, Taiwan. Tel.: +886-7-3208212, Fax: +886-7-3211033 E-mail: wejewukmu.edu.tw.
Corresponding author: Wen-Jeng Wu, Department of Urology, Kaohsiung Medical University Hospital, Kaohsiung Medical University, No. 100, TzYou 1st Road, Kaohsiung City 807, Taiwan. Tel.: +886-7-3208212, Fax: +886-7-3211033 E-mail: wejewukmu.edu.tw.

 Global reach, higher impact
Global reach, higher impact