Impact Factor ISSN: 1449-1907
Int J Med Sci 2014; 11(11):1154-1160. doi:10.7150/ijms.8281 This issue Cite
Short Research Communication
Heterogeneity of Osteosarcoma Cell Lines Led to Variable Responses in Reprogramming
1. MAKNA Cancer Research Institute, Kuala Lumpur, Malaysia;
2. Faculty of Medicine and Health Sciences, University Tunku Abdul Rahman (UTAR), Selangor, Malaysia;
3. Fat Metabolism and Stem Cell Group, Laboratory of Metabolic Medicine, Singapore Bioimaging Consortium, A*STAR, 138667, Singapore;
4. Cardiovascular and Metabolic Disorders Program, Duke-NUS Graduate Medical School, Singapore;
5. Tissue Engineering Group, National Orthopaedic Centre of Excellence for Research and Learning, Department of Orthopaedic Surgery, Faculty of Medicine, University of Malaya, 50603 Kuala Lumpur, Malaysia.
* These authors contributed equally to the work.
Received 2013-12-3; Accepted 2014-7-14; Published 2014-8-22
Abstract
Four osteosarcoma cell lines, Saos-2, MG-63, G-292 and U-2 OS, were reprogrammed to pluripotent state using Yamanaka factors retroviral transduction method. Embryonic stem cell (ESC)-like clusters started to appear between 15 to 20 days post transduction. Morphology of the colonies resembled that of ESC colonies with defined border and tightly-packed cells. The reprogrammed sarcomas expressed alkaline phosphatase and pluripotency markers, OCT4, SSEA4, TRA-1-60 and TRA-1-81, as in ESC up to Passage 15. All reprogrammed sarcomas could form embryoid body-like spheres when cultured in suspension in a low attachment dish for up to 10 days. Further testing on the directed differentiation capacity of the reprogrammed sarcomas showed all four reprogrammed sarcoma lines could differentiate into adipocytes while reprogrammed Saos-2-REP, MG-63-REP and G-292-REP could differentiate into osteocytes. Among the 4 osteosarcoma cell lines, U-2 OS reported the highest transduction efficiency but recorded the lowest reprogramming stability under long term culture. Thus, there may be intrinsic differences governing the variable responses of osteosarcoma cell lines towards reprogramming and long term culture effect of the reprogrammed cells. This is a first report to associate intrinsic factors in different osteosarcoma cell lines with variable reprogramming responses and effects on the reprogrammed cells after prolonged culture.
Keywords: reprogramming, cancer cells, pluripotency, embryonic stem cells, osteosarcoma, heterogeneity.
Introduction
Cancer remains the most-feared disease of people from all walks of life. Despite intensive research efforts on studying this complex disease since the end of 19th century (1), the roles of genetic and epigenetic mechanisms involved in the tumorigenesis process are still not completely understood. However, cancer research took a new turn in 2006 when Yamanaka and colleagues discovered a novel experimental approach in direct reprogramming of somatic cells to their pluripotent state (2). The generated induced pluripotent stem cells (iPSC) hold great potentials as they are embryonic-like and are able to differentiate into all three primary lineages, and could, therefore, be custom-made to treat an individual patient in regenerative therapy (3-5).
With the introduction of this novel dedifferentiation technique, cancer cells could be reprogrammed to produce pluripotent stem cells with cancer properties. The reprogrammed cancer cells could be studied as a personalised model of the cancer from which the cells were derived, subjected to experimental manipulation in the laboratory, and utilised for testing of potential therapeutic drugs to overcome drug resistance in cancers in a personalised setting (6). In addition, it is interesting to note that cancer cells are found to be more responsive to reprogramming than normal somatic cells, suggesting differences in reprogramming potential in cancer and normal cells (7,8). Most of this cancer cell reprogramming studies determined the reprogramming efficiency at the initial stage of formation of ESC-like cells. However, the association between reprogramming efficiency and the product's capability for long term culture has not been studied. Studying the long term culture effect on the reprogrammed cells is essential as there have been publications on prolonged passage could alter cells biological properties as well as accelerating the senescence process (9-11).
Osteosarcoma is a diverse and heterogeneous bone cancer occurring mainly in children and adolescent and is highly variable histologically (12,13). The aim of the present study was to reprogramme osteosarcoma cells into pluripotency by the four transcription factors, OCT4, SOX2, KLF4 and c-MYC, and investigate whether heterogeneity of osteosarcoma causes variable reprogramming responsiveness and the long term culture of reprogrammed osteosarcomas.
Materials and methods
Cell culture. Osteosarcoma cell lines Saos-2, MG-63, U-2 OS and G-292 were purchased from ATCC and maintained in the following conditions; Saos-2: Dulbecco's Modified Eagle Medium, low glucose (DMEM-LG), 10% fetal bovine serum (FBS), 1% Pen-Strep (Gibco, Carlsbad, CA, USA); MG-63: DMEM-LG, 10% FBS, 1% Pen-Strep; U-2 OS: McCoy's 5a Medium, 10% FBS, 1% Pen-Strep; G-292: McCoy's 5a Medium, 10% FBS, 1% Pen-Strep. Reprogrammed cells were maintained in human embryonic stem cells (hESC) medium constituted of DMEM/F12 (Gibco) supplemented with 20% knockout serum replacement (KOSR) (Gibco), 0.1 mM β-mercaptoethanol (Merck, New Jersey, USA), 10 ng/ml basic fibroblast growth factor (bFGF) (Miltenyi Biotech, Germany), 1x non-essential amino-acid (NEAA) (Gibco), 1 mM L-glutamine (Gibco) and 0.1x Pen-Strep (Gibco). Irradiated mouse embryonic fibroblasts (iMEF) (Global Stem, Rockville, MD, USA), used as feeder cells, were plated at a density of 1.5 - 2 x 105 cells to each well of six-well plates and cultured in DMEM with 10% FBS. iPC colonies were manually picked and plated on iMEF plates. All cells were cultured at 37 ˚C in a 5% CO2 incubator.
Retroviral and iPC generation. Human iPC cells were produced as previously described, with slight modification (14). Retroviral vectors for each of the transcription factors genes (pMX-c-MYC, pMX-KLF4, pMX-OCT4, or pMX-SOX2) were transfected with envelope genes (gag/pol and VSV-G) into 293FT cells using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA). 3 days after transfection, the supernatant containing viruses was harvested and filtered through a 0.45-μm filter. OS cells were seeded at a density of 40-60% confluency one day before transduction. Fresh retrovirus supernatants were supplemented with 8 mg/ml Polybrene (Millipore, Merck) prior to transduction. Equal amounts of supernatants containing each of the four retroviruses carrying the OCT4, SOX2, c-MYC and KLF4 genes, were mixed and transferred to the cells. The plates were centrifuged at 800g for 50 min and incubated overnight at 37 ˚C, 5% CO2. Fresh medium was changed on the next day and subsequently every day. Transduced cells were transferred to iMEF on day 3 post transduction. Cells were monitored every day for the formation of colonies. Colonies were manually picked on day 15 - day 20 and transferred to new iMEF.
Immunofluorescence. Cells were fixed with 4% paraformaldehyde (Sigma-Aldrich, St. Louis, MO, USA) at room temperature for 15 min and washed twice in PBS. For intracellular staining, cells were permeabilised with 0.2% Triton-X 100 (Pharmacia Biotech; Uppsala, Sweden) for 15 min. Non-specific binding was blocked by using 10% rabbit serum. The cells were then incubated overnight at 4 ˚C with specific antibodies for either OCT4 (dilution 1:100), SSEA4 (dilution 1:100), TRA-1-60 (dilution 1:100) or TRA-1-81 (dilution 1:100) (Stem Cell Technologies, Canada). After washing, cells were incubated with a secondary antibody containing fluorescein-conjugated rabbit anti-mouse IgG (Chemicon, Millipore, USA) for 1 h at room temperature. After several washes in PBS, cells were viewed for fluorescence using an inverted fluorescence microscope (Carl Zeiss, Germany).
Embryoid body formation. Colonies were manually cut into small pieces and transferred to low-attachment dishes and cultured in ESC medium without FGF for 10 days. Morphology of the embryoid body was observed on day 10 using an inverted microscope.
Directed differentiation assays. To perform adipocytes differentiation, cells were cultured in an adipogenic induction medium comprising DMEM/F12 supplemented with 1.0 μM dexamethasone, 0.2 mM indomethacin, 0.01 mg/mL insulin, 0.5 mM 3-isobutyl-1-methyl-xanthine (Sigma), 10.0% FBS, 1% penicillin and streptomycin (Gibco). Medium change was performed every 3 days for 3 weeks. To carry out osteoblast differentiation; cells were cultured in an osteogenic induction medium, DMEM/F12 supplemented with 10% FBS, 50 μg/mL ascorbate-2-phosphate, 10 mM β-glycerophosphate, 100 nM dexamethasone (Sigma), 1% penicillin and streptomycin (Gibco). Medium change was performed every 3 days for 3 weeks. Both differentiation cultures were incubated in a humidified atmosphere at 37 ºC with 5% CO2. Oil Red O was used as a histological stain to visualise the presence of lipid droplets, while Alizarin Red S was used to stain matrix mineralisation associated with osteoblasts.
Results and Discussion
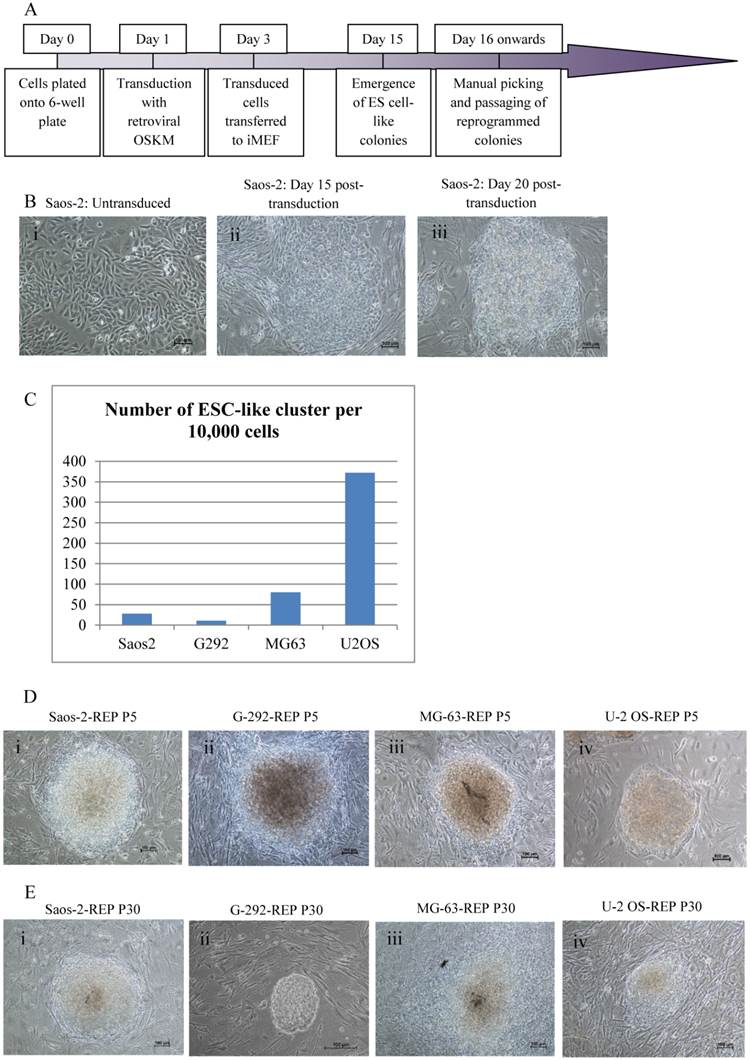
Four osteosarcoma cell lines, Saos-2, MG-63, G-292 and U-2 OS, were reprogrammed with single retroviral transduction of the transcription factors OCT4, SOX2, KLF4 and c-MYC using the experimental schedule depicted in Fig. 1A. ESC-like clusters started to appear between 15 to 20 days post-transduction (Fig. 1B) for all four cell lines. U-2 OS showed the highest reprogramming efficiency with the most ESC-like clusters, followed by MG-63, Saos-2 and G-292 (Fig. 1C). Morphology of the colonies resembled ESC colonies with defined border and cells were tightly packed with each other (Fig. 1D). Colonies were subsequently passaged onto fresh feeder cells for further expansion. Saos-2-REP and G-292-REP could maintained their morphology in culture for more than 30 passages without losing the ESC-like defined borders, whereas MG-63-REP and U-2 OS-REP colonies did not show distinct borders indicating the colonies were losing the ESC-like morphology (Fig. 1E).
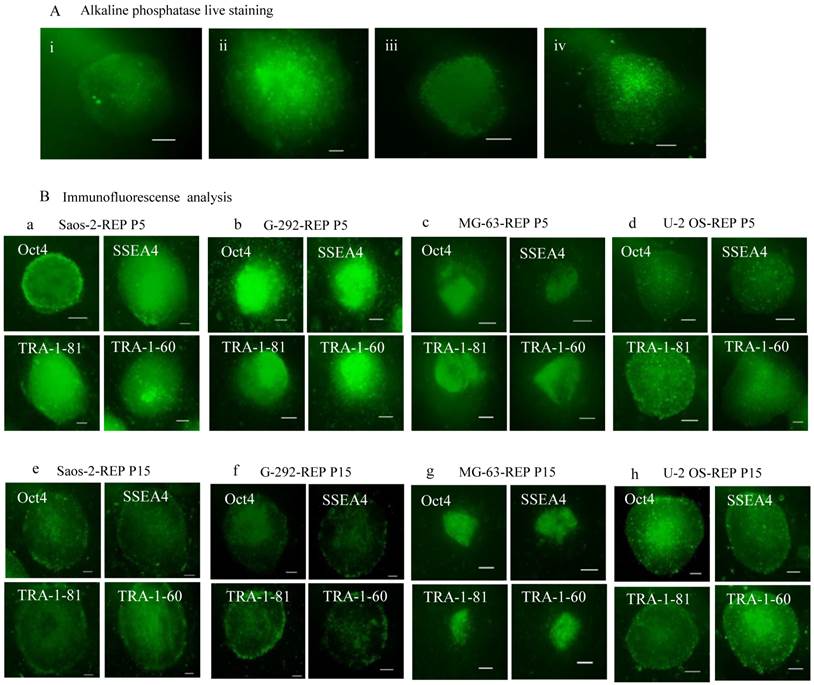
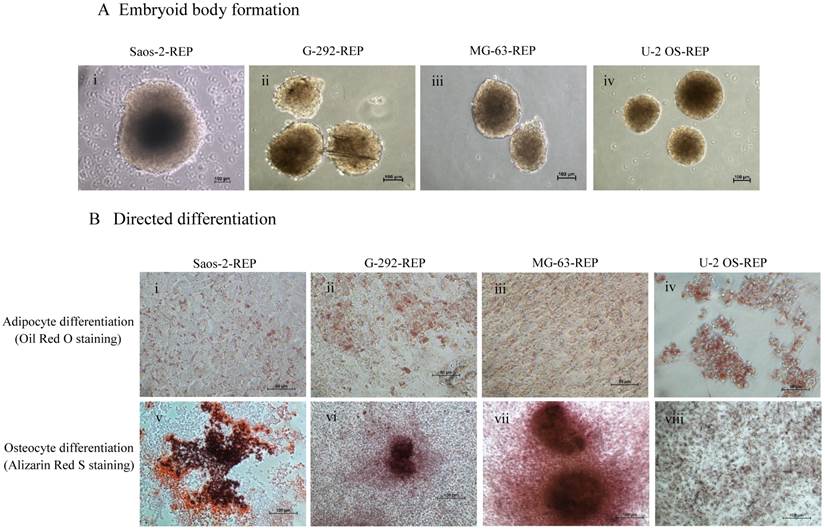
On further characterisation, all the reprogrammed osteosarcoma cells expressed alkaline phosphatase and the pluripotency markers, OCT4, SSEA4, TRA-1-60 and TRA-1-81, as in embryonic stem cells (Fig. 2). Expression of the pluripotency markers was still maintained at passage 15 albeit at a lower staining intensity (Fig. 2B e-h). Embryoid body formation is one of the hallmark characteristics of ESC. All four reprogrammed ostesarcomas formed embryoid body-like spheres for up to 10 days when cultured in suspension condition in a low attachment dish (Fig. 3A). To further test the differentiation capacity of the reprogrammed osteosarcomas, directed differentiation into adipocytes and osteocytes was performed. The results showed that all four reprogrammed osteosarcoma differentiated into adipocytes as shown with Oil Red O staining (Fig. 3B i-iv). However, only Saos-2-REP, MG-63-REP and G-292-REP differentiated into osteocytes as showed by Alizarin Red S staining (Fig. 3B v-viii).
When cancer cells are reprogrammed to pluripotency, they are known as induced pluripotent cancer (iPC) cells. iPC has been successfully achieved for a number of types of cancer (15-17). In this work, we showed for the first time that a single round of retroviral transduction of the four Yamanaka factors was enough to reprogramme four osteosarcoma cell lines into ESC-like colonies with expression of pluripotency markers. In contrast, Zhang et al. previously reported reprogramming of sarcoma cell lines, including Saos-2 and MG-63, with lentiviral transduction of six transcription factors (7). Based on the results from our study and Zhang et al., both retrovirus and lentivirus systems can generate iPC cells with ESC-like characteristics. However, it was unclear whether the genetic and epigenetic states of iPCs generated from these two systems and with different sets of reprogramming factors are the same, or there are subtle molecular and biological differences. Further assay such as the formation of teratoma will be included in the next phase of our study to confirm the pluripotency or ESC-like state of these reprogrammed osteosarcoma cells.
Cancer cells are good candidate for reprogramming as cancer cells express some of the transcription factors crucial for generating iPSC (18-22). Despite our success with osteosarcoma cell line, our attempt on the use a single-round retroviral transduction of the four transcription factors to reprogramme an osteoblast cell line, hFOB, were unsuccessful (data not shown). Osteoblasts are more terminally differentiated somatic cells and did not seem to respond well with our reprogramming protocol. Park et al. achieved reprogramming of ~0.1% fetal fibroblasts using the Yamanaka factors but was unable to reprogramme adult fibroblasts with the same cocktail. Only upon addition of hTERT and SV40 large T-antigen to the reprogramming cocktail were Park et al. able to achieve ~0.25% of adult fibroblasts being reprogrammed (23). Hence, to force adult fibroblasts to respond to the same reprogramming treatment, additional factors are required to grow and to transform the cells to be more adaptive to reprogramming. Reprogramming of osteoblasts and other more terminally differentiated cells may, thus, require additional steps to facilitate the dedifferentiation process.
Reprogramming of osteosarcoma cell lines. (A) The retroviral transduction reprogramming schedule used. (B) Representation images of Saos-2 colony formation post-transduction: (i) Parental Saos-2 prior to reprogramming; (ii) Day 15 post-transduction; (iii) Day 20 post-transduction. (C) The number of ESC-like clusters generated 21 days post transduction of osteosarcoma cell lines with Retroviral OSKM transcription factors. (D) Passage 5 and (E) Passage 30 clones of the reprogrammed osteosarcoma cells: (i) Saos-2 (Saos-2-REP); (ii) G-292 (G-292-REP); (iii) MG-63 (MG-63-REP); (iv) U-2 OS (U-2 OS-REP).
Expression of common pluripotency markers in the reprogrammed oesteosarcome cells. (A) Alkaline phosphatase live staining of: (i) Saos-2-REP, (ii) G-292-REP; (iii) MG-63-REP and (iv) U-2 OS-REP. (B) Immunofluorescence analysis of the reprogrammed cells at passage 5 and 15. (a) & (e) Saos-2-REP, (b) & (f) G-292-REP, (c) & (g) MG-63-REP, (d) & (h) U-2 OS-REP. The colonies expressed the common pluripotency markers OCT4, SSEA4, TRA-1-81 and TRA-1-60.
In a recent study of generation of iPC cells from gastrointestinal cancer cell lines, the researchers highlighted the importance to demonstrate universality in primary tumours and to elucidate factors that are involved in the exerting differences in responses, efficiencies and delivery systems (16). Our results showed different reprogramming efficiencies among the four cell lines used, indicating biological heterogeneity among the osteosarcoma cell lines and the effects of such heterogeneity in dedifferentiation potentials. Among the four tested cell lines, both U-2 OS and MG-63 responded well with transduction and presented the highest and second highest number of ESC-like clusters. However, the morphology of the U-2 OS-REP and MG-63-REP colonies could not be maintained for more than 30 passages, unlike G-292-REP and Saos-2-REP colonies. Our observation showed that transduction efficiency does not necessarily correlate with the success of reprogramming and maintenance of pluripotency. A previous study conducted on different osteosarcoma cell lines indeed demonstrated divergent reactions to p53 or Rb transgenes (24). Data of the said study also demonstrated variable transduction efficiencies of the osteosarcoma cell lines, with transduction of U-2 OS being most efficient compared to Saos-2 and MG-63.
As U-2 OS is the most responsive to transduction with highest number of ESC-like clusters, we further discussed on why U-2 OS behave in such a way. As in our work, U-2 OS could not maintain the effect of the transgenes. Therefore, factors intrinsic to each cell lines are probably responsible for variable transduction efficiencies and U-2 OS may have properties suppressive to the maintenance of pluripotent expression upon prolonged passage.
Formation and directed differentiation of reprogrammed cell-derived embryoid bodies. (A) Embryoid bodies formed from the reprogrammed osteosarcomas in suspension culture: (i) Saos-2-REP (ii) G-292-REP, (iii) MG-63-REP and (iv) U-2 OS-REP. (B) Directed differentiation of the reprogrammed osteosarcoma cells into adipocytes (Oil Red O staining, i-iv) and osteocytes (Alizarin Red S staining, v-viii).
Pautke et al. investigated the characteristics of osteosarcoma cell lines, Saos-2, MG-63 and U-2 OS, and showed that these cell lines exhibited very heterogeneous immunohistochemical labelling profiles and all osteosarcoma cell lines differed significantly from those of normal osteoblasts. The results revealed that Saos-2 cells resemble mature osteoblast phenotype while U-2 OS cells were negative for most of the osteoblastic markers investigated, signifying that U-2 OS cell line is not consistently classified as osteoblastic. (25). Difference in osteoblastic characterisation of osteosarcoma cell lines could impinge on reprogramming accomplishment.
Another study by Benayahu et al. confirmed variable properties and characteristics in osteosarcoma cell lines. Osteosarcoma cell lines, U-2 OS, MG-63 and Saos-2, were studied for the expression of cell cycle regulatory proteins, surface markers and several extracellular matrix (ECM) proteins. Their data demonstrated that all three cell lines have variable degree of expression on the proteins and markers conducted in the study (26). These yet undefined varied characteristics may have great influence on the reprogramming efficiency of the parental cells and long-term culture of the derived iPC cells.
Besides intrinsic factors governing the reprogramming responsiveness of the cells, migration and invasion potentials of the cancer cells could play a role too. Fromigue et al. showed that Saos-2 had the best capacity in migration, while MG-63 had the least migration capacity, and U-2 OS was the most invasive (27). The invasiveness and the higher proliferation rate (28) of U-2 OS could have increased the transduction efficiency of the cells, but may also have decreased the long term effect of transgenes in the cells. Recent work by Xu et al. reported that hyperproliferation of somatic cells could be harmful for reprogramming (29).
Conclusion
We demonstrated here that osteosarcoma cell lines can be reprogrammed to induced pluripotent cancer cells with variable reprogramming efficiency. U-2 OS cells, with higher proliferation rate and more invasive character, respond better to retroviral transduction but could not maintain the ESC-like morphology in prolonged culture suggesting the inability to maintain the effects of the transgenes in the cells. This observation suggests that efficiency of transduction may not correlate with the sustainability of reprogramming.
Acknowledgements
This research is supported by HIR-MoE Grant (Reference number - UM.C/625/1/HIR/MOHE/CHAN/03, account number - A000003-50001). The experiments were mainly carried out in the research laboratories at the Sungai Long Campus of the Universiti Tunku Abdul Rahman (UTAR).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Weinstein IB, Case K. The history of Cancer Research: introducing an AACR Centennial series. Cancer Res. 2008;68:6861-62
2. Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126:663-76
3. Puri MC, Nagy A. Concise review: Embryonic stem cells versus induced pluripotent stem cells: the game is on. Stem Cells. 2012;30:10-4
4. Amabile G, Meissner A. Induced pluripotent stem cells: current progress and potential for regenerative medicine. Trends in Molecular Medicine. 2009;15:59-68
5. Lu SJ, Lee RJ, Napoli C, Oh S, Kimbrel EA, Feng Q. The promise and therapeutic potential of human ES and iPS cells. Stem Cells Int. 2011;2011:959275
6. Tafani M. Reprogramming cancer stem cells. J Cancer Sc Ther. 2012;4:xxv-xxvi
7. Zhang X, Cruz FD, Terry M, Remotti F, Matushansky I. Terminal differentiation and loss of tumorigenicity of human cancers via pluripotency-based reprogramming. Oncogene. 2013;32:2249-60
8. Spike BT, Wahl GM. p53, stem cells, and reprogramming: Tumor suppression beyond guarding the genome. Genes & cancer. 2011;2:404-19
9. Bonab MM, Alimoghaddam K, Talebian F, Ghaffari SH, Ghavamzadeh A, Nikbin B. Aging of mesenchymal stem cell in vitro. BMC Cell Biol. 2006;7:14
10. Izadpanah R, Kaushal D, Kriedt C, Tsien F, Patel B, Dufour J. et al. Long-term in vitro expansion alters the biology of adult mesenchymal stem cells. Cancer Res. 2008;68:4229-38
11. Briquet A, Dubois S, Bekaert S, Dolhet M, Beguin Y, Gothot A. Prolonged ex vivo culture of human bone marrow mesenchymal stem cells influences their supportive activity toward NOD / SCID-repopulating cells and committed progenitor cells of B lymphoid and myeloid lineages. Hematologica. 2010;95:47-56
12. Yuan J, Ossendorf C, Szatkowski JP, Bronk JT, Yaszemski M, Bolander ME. et al. Osteoblastic and osteolytic human osteosarcomas can be studied with a new xenograft mouse model producing spontaneous metastases. Cancer Invest. 2009;27:435-42
13. Gibbs CP, Levings PP, Ghivizzani SC. Evidence for the osteosarcoma stem cell. Curr Orthop Pract. 2011;22:322-6
14. Sugii S, Kida Y, Berggren WT, Evans RM. Feeder-dependent and feeder-independent iPS cell derivation from human and mouse adipose stem cells. Nature Protocols. 2011;6:346-58
15. Carette JE, Pruszak J, Varadarajan M, Blomen VA, Gokhale S, Camargo FD. et al. Generation of iPSCs from cultured human malignant cells. Blood. 2010;115:4039-42
16. Miyoshi N, Ishii H, Nagai K, Hoshino H, Mimori K, Tanaka F. et al. Defined factors induce reprogramming of gastrointestinal cancer cells. PNAS. 2010;107:40-5
17. Lin SL, Chang DC, Chang-Lin S, Lin CH, Wu DTS, Chen DT. et al. Mir-302 reprograms human skin cancer cells into a pluripotent ES-cell-like state. RNA. 2008;14:2115-24
18. Utikal J, Maherali N, Kulalert W, Hochedlinger K. Sox2 is dispensable for the reprogramming of melanocytes and melanoma cells into induced pluripotent stem cells. J Cell Sci. 2009;122:3502-10
19. Liao DJ, Dickson RB. c-Myc in breast cancer. Endocrine-related Cancer. 2000;7:143-64
20. Lin CY, Lovén J, Rahl PB, Paranal RM, Burge CB, Bradner JE. et al. Transcriptional amplification in tumor cells with elevated c-Myc. Cell. 2012;151:56-67
21. Lin ZS, Chu HC, Yen YC, Lewis BC, Chen YW. Krüppel-like factor 4, a tumor suppressor in hepatocellular carcinoma cells reverts epithelial mesenchymal transition by suppressing slug expression. PloS One. 2012;7:e43593
22. Le Magnen C, Bubendorf L, Ruiz C, Zlobec I, Bachmann A, Heberer M. et al. Klf4 transcription factor is expressed in the cytoplasm of prostate cancer cells. Euro J Cancer. 2013;49:955-63
23. Park IH, Zhao R, West JA, Yabuuchi A, Huo H, Ince TA. et al. Reprogramming of human somatic cells to pluripotency with defined factors. Nature. 2008;451:141-6
24. Hellwinkel OJC, Müller J, Pollmann A, Kabisch H. Osteosarcoma cell lines display variable individual reactions on wildtype p53 and Rb tumour-suppressor transgenes. J Gene Med. 2005;7:407-19
25. Pautke C, Schieker M, Tischer T, Kolk A, Neth P, Mutschler W. et al. Characterization of osteosarcoma cell lines MG-63, Saos-2 and U-2 OS in comparison to human osteoblasts. Anticancer Res. 2004;24:3743-8
26. Benayahu D, Shur I, Marom R, Meller I, Issakov J. Cellular and molecular properties associated with osteosarcoma. Cells. 2002;114:108-14
27. Fromigué O, Hamidouche Z, Marie PJ. Blockade of the RhoA-JNK-c-Jun-MMP2 cascade by atorvastatin reduces osteosarcoma cell invasion. J Biol Chem. 2008;283:30549-56
28. Lucero CM, Vega OA, Osorio MM, Tapia JC, Antonelli M, Stein GS. et al. The cancer-related transcription factor Runx2 modulates cell proliferation in human osteosarcoma cell lines. J Cell Physiol. 2013;228:714-23
29. Xu Y, Wei X, Wang M, Zhang R, Fu Y, Xing M. et al. Proliferation rate of somatic cells affects reprogramming efficiency. J Biol Chem. 2013;288:9767-78
Author contact
Corresponding author: Professor Emeritus Dr. Cheong Soon Keng, Faculty of Medicine & Health Sciences, Universiti Tunku Abdul Rahman (UTAR), Jalan Sungai Long, Bandar Sungai Long, 43000 Kajang, Phone number: 03-9019 4722 Email: cheongskedu.my.

 Global reach, higher impact
Global reach, higher impact