Impact Factor ISSN: 1449-1907
Int J Med Sci 2014; 11(10):1065-1072. doi:10.7150/ijms.8473 This issue Cite
Research Paper
Bone Morphogenetic Protein-9 Induces PDLSCs Osteogenic Differentiation through the ERK and p38 Signal Pathways
1. Chongqing key Laboratory for Oral Diseases and Biomedical Sciences, Chongqing 401147, China
2. The Affiliated Hospital of Stomatology, Chongqing Medical University, Chongqing 401147, China
3. Department of Laboratory Medicine, M.O.E. Key Laboratory of Laboratory Medicine Diagnostics, Chongqing Medical University, Chongqing 400016, China.
Received 2014-1-2; Accepted 2014-7-21; Published 2014-8-6
Abstract
Periodontal ligament stem cells (PDLSCs) with bone morphogenic ability are used to treat diseases such as periodontitis. Their treatment potential is increased when used in combination with proteins that induce osteogenic differentiation. For example, bone morphogenetic protein-9 (BMP9) has been found to have potent osteogenic activity. In the present study, PDLSCs were isolated from human periodontal membrane and infected with recombinant adenoviruses expressing BMP9 (Ad-BMP9). Levels of osteogenic markers such as runt-related transcription factor 2 (Runx2), alkaline phosphatase (ALP), osteopontin (OPN), and osteocalcin (OCN) as well as mineralization ability were measured. The results showed that BMP9 promoted bone formation of PDLSCs. In other experiments, SB203580 and PD98059, which are inhibitors of p38 and ERK1/2, respectively, were used to determine if these kinases are involved in the osteogenic differentiation process. The resulting protein expression profiles and osteogenic markers of PDLSCs revealed that the mitogen-activated protein kinase (MAPK) signaling pathway might play an important role in the process of BMP9-induced osteogenic differentiation of PDLSCs.
Keywords: Periodontal ligament stem cells, BMP9, MAPK, Osteogenesis.
Introduction
Periodontitis is a very common chronic bacterial infection that mainly involves periodontal supporting tissue (i.e., periodontal ligaments, alveolar bone, and cementum). It causes progressive irreversible damage that leads to tooth loss [1]. There is a link between periodontitis and systemic diseases such as diabetes and cardiovascular disease, which seriously impact patients' quality of life [2]. The ultimate goal for the treatment of periodontitis is to reconstruct the periodontal tissue and its functions by regenerating periodontal tissue [3]. Several methods have been developed to treat periodontitis, such as guided tissue regeneration, scaling and root planning, demineralized freeze-dried bone allografts, autografts, and the use of bioactive materials [4-6]. However, these treatments have had limited success because they do not effectively regenerate periodontal tissue [7].
The periodontal ligament is a special layer of connective tissue located between the root surface cementum and alveolar bone, and it plays important roles in attaching teeth to the alveolar bone, nutrition of teeth, and repair of damaged tissue [8-10]. In the course of periodontal wound healing, progenitor cells in the periodontal ligament can migrate to the defect region and form alveolar bone tissue [10, 11]. These stem cells were first successfully discovered by Seo et al. in 2004 [12]. Periodontal ligament stem cells (PDLSCs) were isolated by single-colony selection and magnetic activated cell sorting. PDLSCs express mesenchymal stem cell markers (STRO-1, CD146, CD29, CD44, and CD106) [12, 13]. They also have self-renewal capacity and differentiation potential, and under certain culture conditions PDLSCs differentiate into osteoblasts, lipoblasts, cementum-like cells, and collagen-forming cells [13, 14]. Therefore, PDLSCs are good candidates for use in periodontal regeneration strategies [14].
Periodontal regeneration depends largely on mineralized tissue reconstruction, especially bone tissue reconstruction [7, 15]. Thus, determining how to promote osteogenic differentiation of PDLSCs has become a hot research topic in the fields of tissue engineering and regenerative medicine [3, 7, 15]. Bone morphogenetic proteins (BMPs) belong to the transforming growth factor beta superfamily, and more than 20 BMPs have been identified [16, 17]. Osteoblastic stem cells are regulated by BMPs, especially BMP2, BMP4, and BMP7 [16]. Moreover, recombinant human BMP2 and BMP7 have been used in clinical adjuvant therapy for bone regeneration [17, 18]. Cheng et al. (2003) analyzed 14 BMPs (BMP2-15) and found that BMP9 was the most potent inducer of osteogenic differentiation [19]. A distinct set of downstream target genes, which might play a role in the BMP9-induced osteogenic differentiation process, was further identified [20]. Compared with BMP2, muscle-derived stem cells infected with recombinant adenoviruses expressing BMP9 (Ad-BMP9) exhibited more effective bone regeneration in the rabbit radius defect repair model [21]. When human PDLSCs (hPDLSCs) were treated with BMP2, BMP6, and BMP7, different levels of osteogenic gene expression and mineralized nodule formation were observed [22]. However, the effect of treatment of hPDLSCs with BMP9, the BMP with the strongest ability to promote osteogenic differentiation, remains unknown. In the present study, hPDLSCs were infected with Ad-BMP9, and the early and late osteogenesis abilities of hPDLSCs were observed. Furthermore, we investigated the role of the p38 and ERK1/2 mitogen-activated protein kinase (MAPK) pathways in BMP9-induced hPDLSC osteoblast differentiation.
Materials and methods
Isolation and culture of hPDLSCs
All clinical procedures were approved by the Ethics Committee of Chongqing Medical University, and informed consent from patients and their guardians was obtained. Clinically healthy and fresh premolars (n = 28) were removed from 12 patients (aged 12-14 years) during orthodontic treatment at the Maxillofacial Outpatient Surgery of The Affiliated Stomatological Hospital of Chongqing Medical University. The periodontal membrane was gently scraped from the root of each tooth and cut into small pieces (~1 mm3), and samples from individual teeth were pooled and placed in a centrifuge tube. Next, 3 mg/mL collagenase I (Sigma, St Louis, MO, USA) and 4 mg/mL dispase II (Sigma) were added to the tube, and the sample was vibrated for 1 h at 37 °C. A single-cell suspension then was obtained using a 70 µm strainer. The cell density of the single-cell suspension was adjusted to 1 × 104 per milliliter, and the sample was added to a 10 cm diameter Petri dish for single cell-derived colony screening. The sample was cultured in alpha minimal essential medium (alpha MEM, Hyclone, Logan, UT, USA) containing 15% fetal calf serum (FBS, Gibco, Grand Island, NY, USA), 2 mM L-glutamine, 100 mM L-ascorbate-2-phosphate, 100 U/mL streptomycin, and 100 U/mL penicillin at 37 °C and 5% CO2 saturated humidity. Following the method described in the literature [23], the cells were separated by cloning ring and transferred to individual vessels for further study.
Cell culture
The hPDLSCs were maintained in alpha MEM containing 10% FBS (Gibco) and 1% penicillin and streptomycin (Invitrogen, Carlsbad, CA, USA). The cells were maintained in 5% CO2 and 95% humidity. The confluent cells were digested using 0.25% trypsin containing 10 mM EDTA, re-suspended in antibiotic-free growth medium, and plated into 6-well plates at a density of 200,000 cells per well. The hPDLSCs were divided into a blank group (no adenovirus), a control group (infected with Ad-GFP, provided by Dr. Tong-chuan He, Medical Center of the University of Chicago, Chicago, IL, USA), and the experimental group (infected with Ad-BMP9).
Immunocytochemical analysis
The hPDLSCs were planted in 6-well plate at a density of 1 × 104. When cell confluence reached about 80%, cells were fixed in 4% paraformaldehyde for 30 min at room temperature. Cells were fully washed with 0.01 M PBS and incubated with 3% H2O2 at room temperature. They then were incubated with normal goat serum for 30 min at 37 °C followed by rabbit anti-CD146 and anti-Stro-1 overnight at 4 °C. The addition of goat anti-rabbit secondary antibody and visualization with diaminobenzidine (DAB) were conducted according to the manufacturer's instructions to detect the expression of Stro-1 and CD146.
Alkaline phosphatase (ALP) assays
The ALP staining assay and Great Escape SEAP Chemiluminescence assay, which were used to analyze the ALP activity, have been described previously [19, 24, 25]. Briefly, hPDLSCs were infected with Ad-BMP9 for 12 h and then treated with various concentrations of SB203580 (2, 5, and 10 µmol/L) and PD98059 (10, 25, and 50 µmol/L) for 7 d. The cells then were washed with 0.01 M PBS and incubated in the dark for 30 min with ALP staining solution containing napthol AS-MX phosphate (100 µg/mL) and Fast Blue BB salt (600 µg/mL). Finally the staining results were observed under a microscope, and representative photographs were taken. Regarding chemiluminescence assay, each condition was tested in triplicate, and the experiment was repeated three times.
Matrix mineralization assay
hPDLSCs infected with Ad-BMP9 were cultured in alpha MEM containing ascorbic acid (50 µmol/L), dexamethasone (100 nmol/L), and β-glycerophosphate (10 mmol/L). After growing for 21 d, the cells were fixed with 2.5% glutaraldehyde and stained with 2% Alizarin Red S (Sigma), which was used to detect the mineralized matrix nodules based on the presence of calcium. The staining results were observed under a microscope, and representative photographs were taken.
RNA isolation and quantitative real-time PCR
Total RNA was extracted from cells using Trizol reagent (Invitrogen), and it was used to generate cDNA by MMLV reverse transcriptase (Takara, Tokyo, Japan). SYBR Green (Takara) was used in qPCR analysis to amplify the osteogenesis-related genes (runt-related transcription factor 2, Runx2; osteopontin, OPN; osteocalcin, OCN) using the RG-6000 Real-Time DNA Detection System (Corbett Research, NSW, Australia). The cycling program was as follows: 94 °C for 30 s for 1 cycle and 35 cycles at 95 °C for 5 s, 58 °C for 20 s, and 72 °C for 20 s. All samples were run in triplicate and normalized based on the endogenous expression level of GAPDH. The primers for each gene were as follows: Runx2, F: 5′-ACTTCCTGTGCTCGGTGCT-3′, R: 5′-GACGGTTATGGTCAAGGTGAA-3′; OPN, F: 5′-TTGCTTTTGCCTCCTAGCA-3′, R: 5′-GTGAAAACTTCGGTTGCTGG-3′; OCN, F: 5′-GGCAGCGAGGTAGTGAAGAG-3′, R: 5′-CTGGAGAGGAGCAGAACTGG-3′; GAPDH, F: 5′-CAGCGACACCCACTCCTC-3′, R: 5′-TGAGGTCCACCACCCTGT-3′.
Western blot analysis
After hPDLSCs were infected with Ad-BMP9 for 36 h, the cells were lysed using RIPA buffer. The total lysate was centrifuged and the proteins in the supernatant were quantified using the bicinchoninic acid (BCA) assay. The proteins were denatured by boiling for 5 min, resolved on a 10% gradient SDS-PAGE, and then transferred to a PVDF membrane. The blots were blocked with 5% BSA for 2h and incubated with primary antibodies against anti-p38, anti-phospho-p38, anti-ERK1/2, anti-phospho-ERK1/2, and anti-β-actin (Santa Cruz, CA, USA) overnight at 4 °C, followed by secondary antibody incubation at 37 °C for 1 h. Blotted proteins were visualized using an ECL Chemiluminescent Substrate kit.
Statistical analysis
Each experiment in our study was repeated three times. All values in the text and figures are means ± SD. Differences between two groups were analyzed by t-test using SPSS (SPSS Inc., Chicago, IL, USA). p < 0.05 was taken as statistically significant.
Results
Isolation and identification of PDLSCs
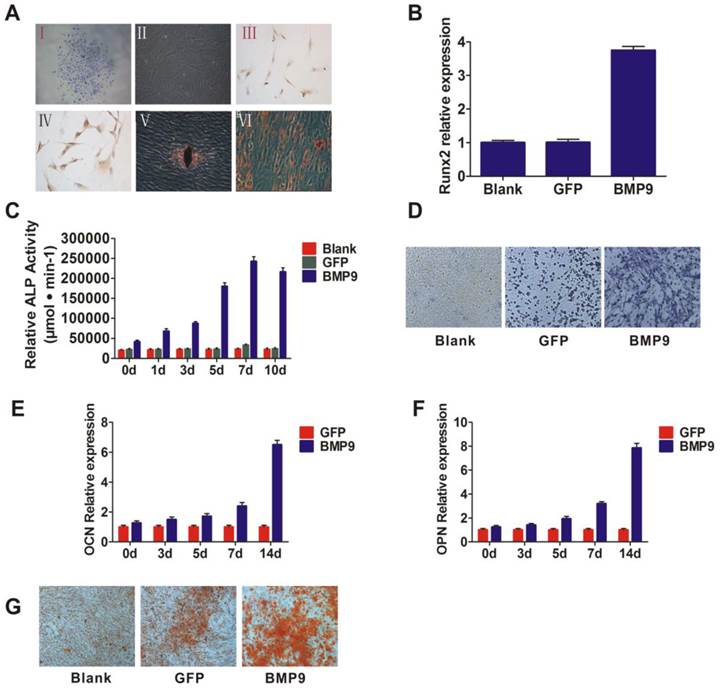
The putative stem cells were isolated and inoculated onto a Petri dish, and then adherent clonogenic cell clusters were observed 7 to 14 days later. The cells that exhibited a typical fibroblast-like spindle appearance were selected by purifying single-colony selection and multiple collecting (Fig. 1A. (I and II)). The immunohistochemical results showed that these cells expressed CD146 and Stro-1, which are early mesenchymal progenitor markers (Fig. 1A. (III and IV)). These data suggested that the isolated cells were PDLSCs. As stem cells exhibit multi-directional differentiation, we then assessed the differentiation potential of the isolated cells. The cells were cultured in osteogenic inductive medium or adipogenic inductive medium. After 4 weeks, mineralized deposits or lipid-laden vacuoles were observed (Fig. 1A. (V and VI)), which indicates that these cells could differentiate and thus likely were PDLSCs.
A: Separation and identification of PDLSCs. I and II: PDLSCs formed colonies (I) and exhibited potent proliferative activity (II). Magnification, ×100; III and IV: Immunohistochemistry and phenotyping were used to determine the source of the cells. PDLSCs were labeled with CD146 (III) and Stro-1 (IV), Magnification, ×200; V and VI: Alizarin Red staining (V) and Oil Red O staining (VI) to assess the potential for cells to differentiate. Magnification, ×200. B: Runx2, the early transcriptional regulator of osteogenic differentiation, was detected by qPCR and the transcriptional activity level was higher in the BMP9 group than in the control group; C: ALP activity was detected by quantitative assay in the cells that were infected with Ad-BMP9; D: ALP activity was detected by ALP staining assay at day 7 after infection. Magnification, ×200; E and F: qPCR analysis of OCN and OPN expression at the gene level. PDLSCs were infected with Ad-BMP9 or Ad-GFP; G: Matrix mineralization of PDLSCs infected with Ad-BMP9 or Ad-GFP. At day 14, cells were subjected to Alizarin Red S staining. Magnification, ×200.
BMP9-induced osteogenic differentiation of PDLSCs
To determine the influence of BMP9 on PDLSCs, exogenous BMP9 was introduced into the cells using a recombinant adenovirus assay. To observe the cells' capacity for early osteogenesis differentiation at different time points, we investigated ALP activity via ALP staining, as ALP activity is a well-established marker for early osteogenic differentiation. After BMP9-induction for 5, 7, and 10 days, the ALP activation levels were higher than the baseline levels of the control group (stimulated with Ad-GFP) (p < 0.01) (Fig. 1C and 1D). ALP activity peaked at day 7, as shown by the enhanced ALP staining intensity at day 7. In addition, the expression of Runx2, which is an early transcriptional regulator of osteogenic differentiation, was detected at day 3, and its transcriptional activity level was significantly increased in the BMP9 treated group compared to the control (Fig. 1B). These results suggest that BMP9 promoted early osteogenic differentiation of PDLSCs.
To further investigate the ability of BMP9 to promote PDLSC osteogenic differentiation, the expression of osteogenesis differentiation-related genes was investigated at 14 days post-BMP9 stimulation. As anticipated, the expression levels of OPN and OCN were significantly higher than those in the blank group and the control group (p < 0.05) (Fig. 1E and 1F). In addition, Alizarin Red S staining indicated that Ad-BMP9 promoted matrix mineralization, whereas the control did not. (Fig. 1G). Taken together, these observations confirmed the ability of BMP9 to promote intermediate and late bone differentiation of PDLSCs.
P38 and ERK1/2 MAPKs in BMP9-induced osteogenic differentiation of PDLSCs
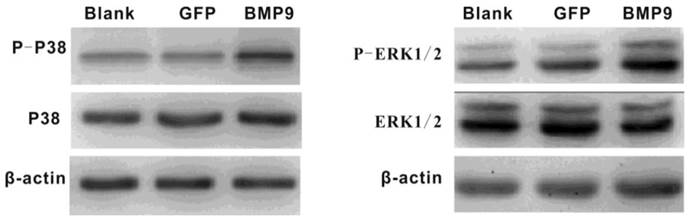
It was previously reported that p38 and ERK1/2 MAPKs play important roles in the BMP pathway. Thus, we wondered whether p38 and ERK1/2 MAPKs would be involved in BMP9-induced osteogenic differentiation of PDLSCs. Proteins were extracted after PDLSCs had been infected with Ad-BMP9 or Ad-GFP for 36 h, and the levels of each protein (p38, phosphor-p38, ERK1/2, and phosphor-ERK1/2) were measured. Western Blot analysis indicated that exposure of the cells to BMP9 effectively increased the levels of phosphor-p38 and phosphor-ERK1/2 without altering the total amounts of these proteins. This result indicates that p38 and ERK1/2 MAPKs were part of the signaling pathway by which BMP9 promoted PDLSC osteogenic differentiation (Fig. 2).
Western blot assay showed that BMP-9 activated the MAPK signaling pathway and increased the phosphorylation of p38 and ERK1/2.
p38 inhibitor reduced BMP9-induced osteogenic differentiation of PDLSCs
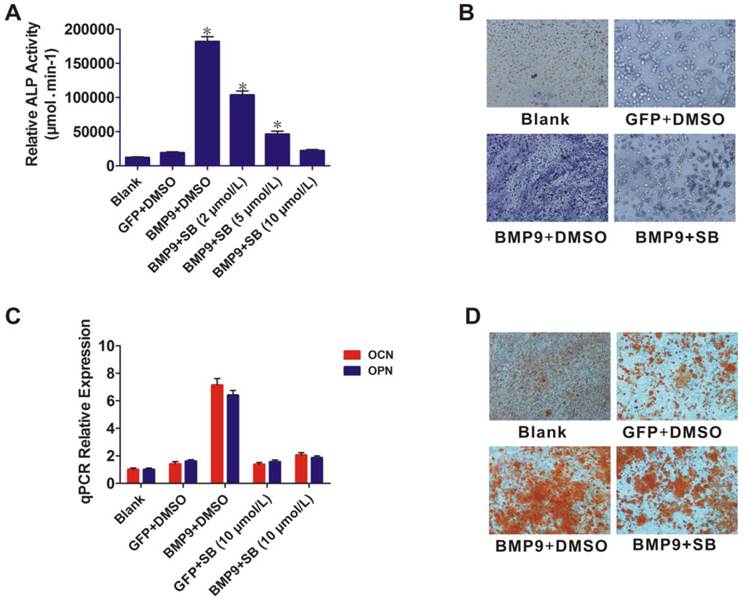
The role of p38 in BMP9-induced osteogenic differentiation of PDLSCs was further examined by using the p38 inhibitor SB203580. BMP9-induced ALP activity was significantly blocked by SB203580 (p < 0.01), and the level of inhibition was dose dependent (Fig. 3A). ALP staining showed that BMP9-induced ALP expression was significantly inhibited by 10 µmol/L SB203580 at day 7 (Fig. 3B). This inhibitory effect was confirmed in another experiment in which 10 µmol/L SB203580 was added to PDLSCs after BMP9-induction for 12 h. After culture for 10 days, total RNA was isolated from the cells and the OPN and OCN expression levels were measured using qPCR (Fig. 3C). SB203580 led to a significant decrease in BMP9-induced OPN and OCN expression (p < 0.01). We also found that SB203580 treatment decreased BMP9-induced matrix mineralization compared with that of the control (Fig. 3D). Collectively, our findings indicated that p38 might play important roles in regulating both early and late stages of BMP9-induced osteogenic differentiation of PDLSCs.
Effect of the p38 inhibitor (SB203580) on BMP9-induced osteogenic differentiation of PDLSCs. A: PDLSCs were infected with Ad-BMP9 or Ad-GFP, and 12 h later they were treated with different concentrations of SB203580 (2, 5, and 10 µmol/L). ALP activity was detected by quantitative assay after 7 days; B: ALP activity was detected by the ALP staining assay 7 days after the Ad-BMP9-induced PDLSCs were treated with SB203580 at 10µmol/L. Magnification, ×200; C: Ad-BMP9-induced PDLSCs were treated with SB203580 at 10 µmol/L, and the gene relative expression of OCN and OPN was measured after 10 days; D: Effect of SB203580 (10 µmol/L) on BMP9-induced matrix mineralization of PDLSC was determined after 21 days. Magnification, ×200.
ERK1/2 inhibitor increased BMP9-induced osteogenic differentiation of PDLSCs
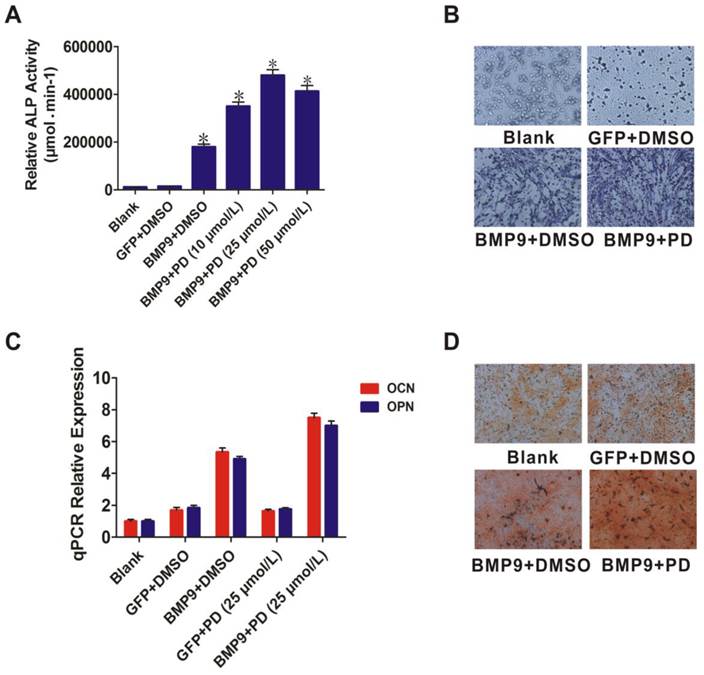
Like p38, ERK1/2 act as downstream mediators of the BMP/MAPK signaling pathway. The ERK1/2 inhibitor PD98059 was used to further evaluate the role of ERK1/2 in BMP9-induced osteogenic differentiation. PD98059 treatment increased BMP9-induced ALP activity of PDLSCs (p < 0.01) (Fig. 4A). ALP staining confirmed that 25 µmol/L PD98059 significantly promoted BMP9-induced PDLSC ALP expression at day 7 (Fig. 4B). Moreover, when PDLSCs were treated with BMP9 in the presence of PD98059 for 10 days, levels of OPN and OCN were increased relative to those of the control (p < 0.01) (Fig. 4C). In addition, Alizarin Red S staining showed that PD98059 promoted matrix mineralization compared to the control (Fig. 4D). These findings indicated that p38 and ERK1/2 have opposite effects in the process of BMP9-induced osteogenic differentiation of PDLSCs.
Effect of the ERK1/2 inhibitor (PD98059) on BMP9-induced osteogenic differentiation of PDLSCs. A: Different doses (10, 25, and 50 µmol/L) of PD98059 were added to BMP9-induced PDLSCs and ALP activity was detected after 7 days; B: 25 µmol/L PD98059 was injected into BMP9-induced PDLSCs and ALP staining assay was used to analyze early osteogenic differentiation after 7 days. Magnification, ×200; C: OCN and OPN expression levels were detected by qPCR 10 days after BMP9-induced PDLSCs were injected with PD98059 at 25 µmol/L; D: Effect of PD98059 at 25 µmol/L on BMP9-induced matrix mineralization of PDLSCs was determined after 21 days. Magnification, ×200.
Discussion
Miller et al. (2000) isolated BMP9, also known as growth differentiation factor 2, from fetal rat liver. They reported that BMP9 plays a positive role in stem cell proliferation and regulates glucose and fat metabolism in the liver [26]. In addition, BMP9 has been shown to be a more potent inducer of osteogenic differentiation compared with other BMPs [27]. In this study, hPDLSCs were infected with Ad-BMP9. Compared with the control (Ad-GFP) and blank groups, ALP activity, gene expression of Runx2, OPN, and OCN, and calcium deposition were significantly higher in the Ad-BMP9 group. Runx2 and ALP are markers of early osteogenesis, OPN and OCN are intermediate or late period osteogenic markers, and calcium deposition is indicative of the late stage of osteogenesis [22]. Thus, our experiments confirmed that BMP9 promoted osteogenesis at the early, middle, and late stages of osteogenic differentiation of hPDLSCs.
Relatively little is known about the specific mechanisms responsible for BMP9-mediated osteogenesis, and many signaling pathways with diverse functions are thought to play roles in this process [27, 28]. The MAPK pathway, which is also called the non-canonical pathway, likely is one of these pathways [29]. It is involved in cellular responses to a diverse array of stimuli, such as proliferation, cell survival, differentiation, and environmental stress [30]. p38, ERK1/2, JNKs, and ERK5 are the members of this pathway, and different stimuli activate different MAPK members, which have different downstream targets; thus, MAPKs play distinct roles in different cellular responses [31]. Many recent studies have focused on p38 and ERK1/2. p38 has been shown to play roles in many cellular processes, such as inflammatory responses, cell cycle regulation, and cellular differentiation of neurons, adipocytes, and myelocytes; ERKs participate in cell multiplication or transformation and survival [32].
Many researchers have studied the role of MAPKs in regulating osteogenic differentiation, but results have been inconsistent. Hu et al. (2003) reported that p38 played a positive role in early and late osteogenesis of osteoblasts, bone marrow osteoprogenitor cells, and a murine osteoblast line, MC3T3-E1 [33]. Greenblatt et al. (2010) verified that p38 promoted skeleton genesis and bone homeostasis through Runx2, which is a key transcription factor associated with osteoblast differentiation [34]. They also confirmed that ERK1/2 promoted osteoblast differentiation and fetal rat skeletal development in vivo [34]. However, Higuchi et al. (2002) reported that sustained suppression of ERK1/2 could promote early osteoblast differentiation and mineralization of the extracellular matrix [35]. Viñals et al. (2002) also reported that p38 MAPK had a negative role in osteoblast differentiation [36].
In the present study, we found that the expression levels of early, middle, and late osteogenic markers were significantly reduced when the p38 signal pathway was inhibited. In contrast, levels of the osteogenic markers were elevated when ERK1/2 was inhibited. These findings revealed that both p38 and ERK1/2 were involved in the BMP9-mediated osteoinductive process but that they had opposite effects. Zhao et al. (2012) analyzed mesenchymal progenitor cell lines (C3H10T1/2, C2C12, HEK293, and HCT116) in vitro and in vivo and demonstrated that p38 promoted osteogenic differentiation that but ERK1/2 did not [30]. In addition, Li et al. (2012) reported the same conclusion in BMP9-induced osteogenic differentiation of rat dental follicle stem cells [36]. From these various observations, we propose that the p38 and ERK1/2 pathways co-determine BMP9-promoted osteogenesis.
Since Seo et al. (2004) first successfully isolated PDLSCs, these cells have been the focus of many studies due to their ability to induce periodontal tissue regeneration [12]. PDLSCs have become popular periodontal tissue engineering seed cells [3, 37, 38]. However, they lack a specific marker to tag. Thus, in the current study PDLSCs were identified based on their biological characteristics: their capacity to self-renew and to form cell colonies, the presence of stem cell markers (STRO-1, CD146+, CD44+, CD105+ and CD166+), and their capacity to differentiate into multiple types of cells (i.e., bone, fat, and nerve) [13, 39]. Furthermore, our experimental results confirmed the BMP9-induced differentiation capacity of PDLSCs and showed that the p38 and ERK1/2 MAPK pathway is involved in BMP9-induced osteogenic differentiation. By using specific inhibitors, we found that p38 and ERK1/2 had opposite effects on BMP9-induced PDLSCs differentiation. These findings provide new insight into the ability of BMP9 to promote PDLSC osteogenic differentiation and the possible interaction between the MAPK pathway and other cell pathways. Our results suggest that the application of PDLSCs combined with BMP9 could prove to be a useful approach to promoting periodontal tissue regeneration in the clinical setting.
Acknowledgements
This work was supported by the Medical Scientific Research Program of the Health Bureau of Chongqing (2012-1-056), the Chongqing Science & Technology Commission of Yubei district (2013-31), the Chongqing Science & Technology Commission of Yuzhong district (x11073), and the Natural Science Foundation of Chongqing (cstc2014jcyjA0240).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Lim JC, Mitchell CH. Inflammation, pain, and pressure--purinergic signaling in oral tissues. Journal of Dental Research. 2012;91:1103-9
2. Jeftha A, Holmes H. Periodontitis and cardiovascular disease. SADJ: Journal of the South African Dental Association = tydskrif van die Suid-Afrikaanse Tandheelkundige Vereniging. 2013;68:60-63
3. Hughes FJ, Ghuman M, Talal A. Periodontal regeneration: a challenge for the tissue engineer? Proceedings of the Institution of Mechanical Engineers Part H, Journal of Engineering in Medicine. 2010;224:1345-58
4. Bono A, Brunotto M. Amoxicillin/metronidazole or scaling and root planing in the treatment of chronic periodontitis. Acta Odontologica Latinoamericana: AOL. 2010;23:196-203
5. Kodama T, Minabe M, Sugiyama T. et al. Guided tissue regeneration using a collagen barrier and bone swaging technique in noncontained infrabony defects. The International Journal of Periodontics & Restorative Dentistry. 2013;33:805-812
6. Wood RA, Mealey BL. Histologic comparison of healing after tooth extraction with ridge preservation using mineralized versus demineralized freeze-dried bone allograft. Journal of Periodontology. 2012;83:329-336
7. Rodriguez-Lozano FJ, Insausti CL, Iniesta F. et al. Mesenchymal dental stem cells in regenerative dentistry. Medicina Oral, Patologia Oral y Cirugia Bucal. 2012;17:e1062-1067
8. Chen SC, Marino V, Gronthos S. et al. Location of putative stem cells in human periodontal ligament. Journal of Periodontal Research. 2006;41:547-553
9. Nanci A, Bosshardt DD. Structure of periodontal tissues in health and disease. Periodontology 2000. 2006;40:11-28
10. Shimono M, Ishikawa T, Ishikawa H. et al. Regulatory mechanisms of periodontal regeneration. Microscopy Research and Technique. 2003;60:491-502
11. Ohta S, Yamada S, Matuzaka K. et al. The behavior of stem cells and progenitor cells in the periodontal ligament during wound healing as observed using immunohistochemical methods. Journal of Periodontal Research. 2008;43:595-603
12. Seo BM, Miura M, Gronthos S. et al. Investigation of multipotent postnatal stem cells from human periodontal ligament. Lancet. 2004;364:149-155
13. Mrozik K, Gronthos S, Shi S. et al. A method to isolate, purify, and characterize human periodontal ligament stem cells. Methods in Molecular Biology. 2010;666:269-284
14. Zhang C, Li J, Zhang L. et al. Effects of mechanical vibration on proliferation and osteogenic differentiation of human periodontal ligament stem cells. Archives of Oral Biology. 2012;57:1395-1407
15. Tang N, Zhao Z, Zhang L. et al. Up-regulated osteogenic transcription factors during early response of human periodontal ligament stem cells to cyclic tensile strain. Archives of Medical Science: AMS. 2012;8:422-430
16. Jain AP, Pundir S, Sharma A. Bone morphogenetic proteins: The anomalous molecules. Journal of Indian Society of Periodontology. 2013;17:583-586
17. Yoshida M, Okubo N, Chosa N. et al. TGF-beta-operated growth inhibition and translineage commitment into smooth muscle cells of periodontal ligament-derived endothelial progenitor cells through Smad- and p38 MAPK-dependent signals. International Journal of Biological Sciences. 2012;8:1062-1074
18. Leach J, Bittar RG. BMP-7 (OP-1) safety in anterior cervical fusion surgery. Journal of Clinical Neuroscience: official journal of the Neurosurgical Society of Australasia. 2009;16:1417-1420
19. Cheng H, Jiang W, Phillips FM. et al. Osteogenic activity of the fourteen types of human bone morphogenetic proteins (BMPs). The Journal of Bone and Joint Surgery American volume. 2003;85A:1544-1552
20. Kang Q, Sun MH, Cheng H. et al. Characterization of the distinct orthotopic bone-forming activity of 14 BMPs using recombinant adenovirus-mediated gene delivery. Gene Therapy. 2004;11:1312-1320
21. Xiang L, Liang C, Zhen-Yong K. et al. BMP9-induced osteogenetic differentiation and bone formation of muscle-derived stem cells. Journal of Biomedicine & Biotechnology. 2012;2012:610952
22. Xu WP, Shiba H, Mizuno N. et al. Effect of bone morphogenetic proteins-4, -5 and -6 on DNA synthesis and expression of bone-related proteins in cultured human periodontal ligament cells. Cell Biology International. 2004;28:675-682
23. Ge S, Zhao N, Wang L. et al. Bone repair by periodontal ligament stem cellseeded nanohydroxyapatite-chitosan scaffold. International Journal of Nanomedicine. 2012;7:5405-5414
24. Peng Y, Kang Q, Cheng H. et al. Transcriptional characterization of bone morphogenetic proteins (BMPs)-mediated osteogenic signaling. Journal of Cellular Biochemistry. 2003;90:1149-1165
25. Luo J, Tang M, Huang J. et al. TGFbeta/BMP type I receptors ALK1 and ALK2 are essential for BMP9-induced osteogenic signaling in mesenchymal stem cells. The Journal of Biological Chemistry. 2010;285:29588-29598
26. Miller AF, Harvey SA, Thies RS. et al. Bone morphogenetic protein-9. An autocrine/paracrine cytokine in the liver. The Journal of Biological Chemistry. 2000;275:17937-17945
27. Lamplot JD, Qin J, Nan G. et al. BMP9 signaling in stem cell differentiation and osteogenesis. American Journal of Stem Cells. 2013;2:1-21
28. Cao X, Chen D. The BMP signaling and in vivo bone formation. Gene. 2005;357:1-8
29. Xu DJ, Zhao YZ, Wang J. et al. Smads, p38 and ERK1/2 are involved in BMP9-induced osteogenic differentiation of C3H10T1/2 mesenchymal stem cells. BMB Reports. 2012;45:247-252
30. Zhao Y, Song T, Wang W. et al. P38 and ERK1/2 MAPKs act in opposition to regulate BMP9-induced osteogenic differentiation of mesenchymal progenitor cells. PloS One. 2012;7:e43383
31. Pearson G, Robinson F, Beers Gibson T. et al. Mitogen-activated protein (MAP) kinase pathways: regulation and physiological functions. Endocrine Reviews. 2001;22:153-183
32. Guicheux J, Lemonnier J, Ghayor C. et al. Activation of p38 mitogen-activated protein kinase and c-Jun-NH2-terminal kinase by BMP-2 and their implication in the stimulation of osteoblastic cell differentiation. Journal of Bone and Mineral Research: the official journal of the American Society for Bone and Mineral Research. 2003;18:2060-2068
33. Hu Y, Chan E, Wang SX. et al. Activation of p38 mitogen-activated protein kinase is required for osteoblast differentiation. Endocrinology. 2003;144:2068-2074
34. Greenblatt MB, Shim JH, Zou W. et al. The p38 MAPK pathway is essential for skeletogenesis and bone homeostasis in mice. The Journal of Clinical Investigation. 2010;120:2457-2473
35. Higuchi C, Myoui A, Hashimoto N. et al. Continuous inhibition of MAPK signaling promotes the early osteoblastic differentiation and mineralization of the extracellular matrix. Journal of Bone and Mineral Research: the official journal of the American Society for Bone and Mineral Research. 2002;17:1785-1794
36. Li C, Yang X, He Y. et al. Bone morphogenetic protein-9 induces osteogenic differentiation of rat dental follicle stem cells in P38 and ERK1/2 MAPK dependent manner. International Journal of Medical Sciences. 2012;9:862-871
37. Liu Y, Zheng Y, Ding G. et al. Periodontal ligament stem cell-mediated treatment for periodontitis in miniature swine. Stem Cells. 2008;26:1065-1073
38. Ding G, Liu Y, Wang W. et al. Allogeneic periodontal ligament stem cell therapy for periodontitis in swine. Stem Cells. 2010;28:1829-1838
39. Chen D, Zhao M, Mundy GR. Bone morphogenetic proteins. Growth Factors. 2004;22:233-241
Author contact
Corresponding author: Yaguang Weng, Department of Laboratory Medicine, M.O.E. Key Laboratory of Laboratory Medicine Diagnostics, Chongqing Medical University, 1#, Yixueyuan Road, Chongqing 400016, China. Tel: +86 023 68485239; Fax: +86 023 68485006; Email: yaguangwengcom Qiong Shi, Department of Laboratory Medicine, M.O.E. Key Laboratory of Laboratory Medicine Diagnostics, Chongqing Medical University, 1#, Yixueyuan Road, Chongqing 400016, China. Tel: +86 023 68485239; Fax: +86 023 68485006; Email: anniesq8718com.

 Global reach, higher impact
Global reach, higher impact