Impact Factor ISSN: 1449-1907
Int J Med Sci 2014; 11(5):461-465. doi:10.7150/ijms.7967 This issue Cite
Research Paper
Commonality of Drug-associated Adverse Events Detected by 4 Commonly Used Data Mining Algorithms
1. Center for Integrative Education in Pharmacy and Pharmaceutical Sciences, Graduate School of Pharmaceutical Sciences, Kyoto University, Kyoto 606-8501, Japan;
2. Department of Systems Biosciences for Drug Discovery, Graduate School of Pharmaceutical Sciences, Kyoto University, Kyoto 606-8501, Japan.
Received 2013-10-24; Accepted 2014-2-20; Published 2014-3-15
Abstract
Objectives: Data mining algorithms have been developed for the quantitative detection of drug-associated adverse events (signals) from a large database on spontaneously reported adverse events. In the present study, the commonality of signals detected by 4 commonly used data mining algorithms was examined.
Methods: A total of 2,231,029 reports were retrieved from the public release of the US Food and Drug Administration Adverse Event Reporting System database between 2004 and 2009. The deletion of duplicated submissions and revision of arbitrary drug names resulted in a reduction in the number of reports to 1,644,220. Associations with adverse events were analyzed for 16 unrelated drugs, using the proportional reporting ratio (PRR), reporting odds ratio (ROR), information component (IC), and empirical Bayes geometric mean (EBGM).
Results: All EBGM-based signals were included in the PRR-based signals as well as IC- or ROR-based ones, and PRR- and IC-based signals were included in ROR-based ones. The PRR scores of PRR-based signals were significantly larger for 15 of 16 drugs when adverse events were also detected as signals by the EBGM method, as were the IC scores of IC-based signals for all drugs; however, no such effect was observed in the ROR scores of ROR-based signals.
Conclusions: The EBGM method was the most conservative among the 4 methods examined, which suggested its better suitability for pharmacoepidemiological studies. Further examinations should be performed on the reproducibility of clinical observations, especially for EBGM-based signals.
Keywords: adverse event, Adverse Event Reporting System, FAERS, database, data mining, signal, signal detection, proportional reporting ratio, reporting odds ratio, information component, empirical Bayes geometric mean.
Introduction
The US Food and Drug Administration (FDA) Adverse Event Reporting System (FAERS, formerly AERS) is a database that contains information on adverse event and medication error reports submitted to the FDA [1-3]. Besides those from manufacturers, reports can be submitted from health care professionals and the general public. The FAERS structure adheres to the International Safety Reporting Guidance issued by the International Conference on Harmonisation, ICH E2B, and adverse events are coded to terms in the Medical Dictionary for Regulatory Activities (MedDRA) terminology [4]. The original system was initiated in 1969; however, reporting markedly increased following the last major revision in 1997 [5, 6]. To date, the FAERS contains more than 4 million reports and is the largest repository of spontaneously reported adverse events in the world [5, 6]. The FDA releases data to the general public, and this has allowed us to conduct pharmacoepidemiological studies and/or pharmacovigilance analyses.
Data mining algorithms have been developed for the quantitative detection of signals [7-11]. A signal indicates an association between a drug and an adverse event or drug-associated adverse event, including the proportional reporting ratio (PRR) [12], reporting odds ratio (ROR) [13], information component (IC) given by a Bayesian confidence propagation neural network [14], and empirical Bayes geometric mean (EBGM) [15]. Associations with adverse events of interests were previously analyzed for 16 drugs using reports in the FAERS database between 2004 and 2009 [16-22]. Whether an adverse event is detected as a signal has been shown to depend on the algorithms; however, of the 4 methods, the ROR method provided the highest number of signals, while the EBGM method provided the lowest [23]. In the present study, the commonality of PRR-, ROR-, IC-, and EBGM-based signals was examined.
Methods
Data were retrieved from the public release of the FAERS database from the first quarter of 2004 through to the end of 2009. The total number of reports obtained was 2,231,029. Duplicated reports were deleted and arbitrary drug names were revised, resulting in a reduction in the number of reports from 2,231,029 to 1,644,220. Signal scores, i.e., the PRR, ROR, IC, and EBGM values, were calculated for 16 unrelated drugs to assess associations with adverse events, including 2 antimicrobials (colistin and tigecycline), 4 HMG-CoA reductase inhibitors (statins) (pravastatin, simvastatin, atorvastatin, and rosuvastatin), 2 proton pump inhibitors (PPIs) (omeprazole and esomeprazole), warfarin, 2 antiplatelets (aspirin and clopidogrel), and 5 anticancer agents (cisplatin, carboplatin, oxaliplatin, 5-fluorouracil, and capecitabine). It is noted that the associations of these drugs with adverse events have already been published [16-22]. All values reported are the mean±standard deviation (SD). The unpaired Student's t-test/Welch's test or Mann-Whitney's U test was used for two-group comparisons of the values. P values of less than 0.05 were considered significant.
Results
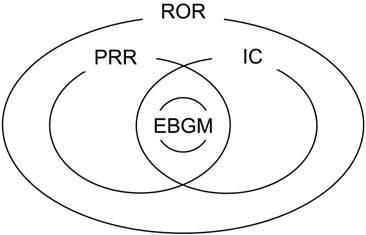
Figure 1 shows the relationship among the PRR-, ROR-, IC-, and EBGM-based signals, which was commonly observed for all 16 drugs. All EBGM-based signals were included in the PRR-based signals as well as IC- or ROR-based ones. The PRR- and IC-based signals were included in the ROR-based ones. Therefore, ROR-based signals could be stratified into 5 groups; signals detected by the ROR only, signals detected by the ROR and PRR, signals detected by the ROR and IC, signals detected by the ROR, PRR, and IC, and signals detected by the 4 methods. Table 1 lists the numbers of signals in the 5 groups. The ratio of the total number of EBGM-based signals to that of signals detected by the ROR only varied from 3.9% with omeprazole to 57.3% with oxaliplatin. The ratio of the total number of EBGM-based signals to that of ROR-based signals varied from 1.7% with omeprazole to 20.5% with oxaliplatin.
Commonality of signals detected by 4 commonly used data mining algorithms. PRR: proportional reporting ratio; ROR: reporting odds ratio; IC: information component; EBGM: empirical Bayes geometric mean. ROR-based signals were stratified into 5 groups; signals detected by the ROR only, signals detected by the ROR and PRR, signals detected by the ROR and IC, signals detected by the ROR, PRR, and IC, and signals detected by the 4 methods. The numbers of signals in the 5 groups are listed in Table 1.
Numbers of signals in the 5 groups.
| ROR only | ROR&PRR | ROR&IC | ROR&PRR&IC | ROR&PRR&IC&EBGM | |
|---|---|---|---|---|---|
| Cisplatin | 356 | 98 | 49 | 206 | 175 |
| Carboplatin | 321 | 77 | 80 | 188 | 144 |
| Oxaliplatin | 262 | 64 | 60 | 196 | 150 |
| Colistin | 166 | 18 | 1 | 30 | 23 |
| 5-Fluorouracil | 341 | 82 | 62 | 218 | 161 |
| Capecitabine | 340 | 67 | 51 | 198 | 146 |
| Pravastatin | 358 | 58 | 125 | 141 | 19 |
| Simvastatin | 284 | 61 | 268 | 101 | 30 |
| Atorvastatin | 304 | 65 | 295 | 164 | 55 |
| Rosuvastatin | 295 | 42 | 97 | 122 | 63 |
| Tigecycline | 155 | 18 | 2 | 29 | 44 |
| Omeprazole | 361 | 87 | 244 | 112 | 14 |
| Esomeprazole | 348 | 78 | 201 | 99 | 17 |
| Warfarin | 248 | 62 | 157 | 159 | 110 |
| Aspirin | 385 | 86 | 115 | 162 | 100 |
| Clopidogrel | 287 | 75 | 185 | 187 | 104 |
PRR: proportional reporting ratio; ROR: reporting odds ratio; IC: information component; EBGM: empirical Bayes geometric mean.
ROR-based signals were stratified into 5 groups; signals detected by the ROR only, signals detected by the ROR and PRR, signals detected by the ROR and IC, signals detected by the ROR, PRR, and IC, and signals detected by the 4 methods.
Table 2 lists the PRR scores of PRR-based signals. Since PRR-based signals could be divided into 2 groups based on whether adverse events were also detected as signals by the EBGM method (Figure 1), the effects of additional detection by the EBGM method on PRR scores was examined. As shown in Table 2, the scores were significantly larger for 15 of 16 drugs when adverse events were also detected as signals by the EBGM method. Tables 3 and 4 show data on the ROR and IC, respectively. The effects of additional detection by the EBGM method found for PRR scores were not observed for the ROR, whereas the IC scores of IC-based signals were the same as the PRR scores of PRR-based signals.
PRR scores of PRR-based signals (the signals detected by the PRR method).
| All | Detected by EBGM | Not detected by EBGM | p | ||||
|---|---|---|---|---|---|---|---|
| N | PRR | N | PRR | N | PRR | ||
| Cisplatin | 479 | 8.03 ± 11.29 | 175 | 12.90 ± 16.73 | 304 | 5.23 ± 4.36 | < 0.001 |
| Carboplatin | 409 | 6.80 ± 8.32 | 144 | 10.57 ± 12.25 | 265 | 4.76 ± 3.69 | < 0.001 |
| Oxaliplatin | 410 | 7.72 ± 11.47 | 150 | 11.69 ± 17.16 | 260 | 5.43 ± 4.90 | < 0.001 |
| Colistin | 71 | 29.30 ± 83.82 | 23 | 77.31 ± 136.92 | 48 | 6.29 ± 4.66 | < 0.001 |
| 5-Fluorouracil | 461 | 7.52 ± 10.03 | 161 | 11.61 ± 14.90 | 300 | 5.33 ± 4.72 | < 0.001 |
| Capecitabine | 411 | 8.09 ± 13.06 | 146 | 12.07 ± 20.26 | 265 | 5.90 ± 5.09 | < 0.001 |
| Pravastatin | 218 | 4.70 ± 4.26 | 19 | 10.48 ± 8.61 | 199 | 4.15 ± 3.11 | < 0.001 |
| Simvastatin | 192 | 4.50 ± 4.81 | 30 | 8.99 ± 10.33 | 162 | 3.66 ± 1.94 | < 0.001 |
| Atorvastatin | 284 | 3.76 ± 1.93 | 55 | 4.41 ± 1.99 | 229 | 3.61 ± 1.89 | < 0.001 |
| Rosuvastatin | 227 | 5.20 ± 5.77 | 63 | 8.50 ± 9.37 | 164 | 3.94 ± 2.65 | < 0.001 |
| Tigecycline | 91 | 37.88 ± 114.30 | 44 | 72.09 ± 158.16 | 47 | 5.85 ± 3.57 | < 0.001 |
| Omeprazole | 213 | 4.69 ± 5.05 | 14 | 12.29 ± 15.28 | 199 | 4.16 ± 2.77 | 0.003 |
| Esomeprazole | 194 | 4.65 ± 3.83 | 17 | 7.19 ± 9.50 | 177 | 4.41 ± 2.68 | 0.513 |
| Warfarin | 331 | 5.28 ± 4.95 | 110 | 7.46 ± 7.38 | 221 | 4.19 ± 2.47 | < 0.001 |
| Aspirin | 348 | 5.56 ± 4.93 | 100 | 8.05 ± 7.39 | 248 | 4.56 ± 2.96 | < 0.001 |
| Clopidogrel | 366 | 4.85 ± 3.79 | 104 | 6.77 ± 5.44 | 262 | 4.08 ± 2.52 | < 0.001 |
PRR-based signals were divided into 2 groups based on whether adverse events were also detected by the EBGM method.
ROR scores of ROR-based signals (the signals detected by the ROR method).
| All | Detected by EBGM | Not detected by EBGM | p | ||||
|---|---|---|---|---|---|---|---|
| N | ROR | N | ROR | N | ROR | ||
| Cisplatin | 884 | 15.75 ± 34.12 | 175 | 13.92 ± 20.63 | 709 | 16.20 ± 36.69 | 0.002 |
| Carboplatin | 810 | 14.95 ± 43.93 | 144 | 11.07 ± 14.11 | 666 | 15.78 ± 47.96 | 0.001 |
| Oxaliplatin | 732 | 12.32 ± 31.94 | 150 | 12.41 ± 20.31 | 582 | 12.29 ± 34.32 | < 0.001 |
| Colistin | 238 | 57.84 ± 165.03 | 23 | 78.97 ± 141.67 | 215 | 55.58 ± 167.47 | 0.028 |
| 5-Fluorouracil | 864 | 14.89 ± 37.82 | 161 | 12.34 ± 18.38 | 703 | 15.47 ± 40.99 | 0.001 |
| Capecitabine | 802 | 17.16 ± 54.77 | 146 | 13.10 ± 25.20 | 656 | 18.06 ± 59.35 | 0.097 |
| Pravastatin | 701 | 10.00 ± 23.37 | 19 | 10.92 ± 9.30 | 682 | 9.97 ± 23.64 | 0.019 |
| Simvastatin | 744 | 5.37 ± 7.17 | 30 | 11.03 ± 16.14 | 714 | 5.13 ± 6.45 | < 0.001 |
| Atorvastatin | 883 | 5.14 ± 8.66 | 55 | 4.61 ± 2.24 | 828 | 5.18 ± 8.92 | < 0.001 |
| Rosuvastatin | 619 | 11.87 ± 27.18 | 63 | 8.93 ± 10.68 | 556 | 12.21 ± 28.44 | 0.074 |
| Tigecycline | 248 | 70.05 ± 381.27 | 44 | 74.82 ± 170.86 | 204 | 69.03 ± 413.14 | 0.008 |
| Omeprazole | 818 | 6.39 ± 11.04 | 14 | 16.92 ± 26.68 | 804 | 6.20 ± 10.51 | 0.003 |
| Esomeprazole | 743 | 6.83 ± 10.03 | 17 | 8.05 ± 11.77 | 726 | 6.80 ± 9.99 | 0.308 |
| Warfarin | 736 | 7.81 ± 13.74 | 110 | 8.36 ± 10.06 | 626 | 7.72 ± 14.30 | < 0.001 |
| Aspirin | 848 | 11.86 ± 35.85 | 100 | 8.38 ± 8.38 | 748 | 12.32 ± 38.02 | 0.033 |
| Clopidogrel | 838 | 6.20 ± 9.01 | 104 | 7.19 ± 6.26 | 734 | 6.06 ± 9.33 | < 0.001 |
ROR-based signals were divided into 2 groups based on whether adverse events were also detected by the EBGM method.
IC scores of IC-based signals (the signals detected by the IC method).
| All | Detected by EBGM | Not detected by EBGM | p | ||||
|---|---|---|---|---|---|---|---|
| N | IC | N | IC | N | IC | ||
| Cisplatin | 430 | 1.64 ± 0.67 | 175 | 2.22 ± 0.55 | 255 | 1.24 ± 0.39 | < 0.001 |
| Carboplatin | 412 | 1.51 ± 0.66 | 144 | 2.15 ± 0.53 | 268 | 1.16 ± 0.42 | < 0.001 |
| Oxaliplatin | 406 | 1.60 ± 0.69 | 150 | 2.22 ± 0.62 | 256 | 1.23 ± 0.41 | < 0.001 |
| Colistin | 54 | 1.82 ± 0.52 | 23 | 2.25 ± 0.47 | 31 | 1.51 ± 0.28 | < 0.001 |
| 5-Fluorouracil | 441 | 1.62 ± 0.70 | 161 | 2.32 ± 0.54 | 280 | 1.22 ± 0.40 | < 0.001 |
| Capecitabine | 395 | 1.66 ± 0.70 | 146 | 2.31 ± 0.63 | 249 | 1.28 ± 0.41 | < 0.001 |
| Pravastatin | 285 | 1.03 ± 0.48 | 19 | 1.98 ± 0.28 | 266 | 0.96 ± 0.41 | < 0.001 |
| Simvastatin | 399 | 0.81 ± 0.50 | 30 | 1.96 ± 0.51 | 369 | 0.72 ± 0.36 | < 0.001 |
| Atorvastatin | 514 | 0.92 ± 0.52 | 55 | 1.88 ± 0.41 | 459 | 0.80 ± 0.41 | < 0.001 |
| Rosuvastatin | 282 | 1.27 ± 0.68 | 63 | 2.18 ± 0.60 | 219 | 1.00 ± 0.42 | < 0.001 |
| Tigecycline | 75 | 2.05 ± 0.68 | 44 | 2.44 ± 0.58 | 31 | 1.50 ± 0.34 | < 0.001 |
| Omeprazole | 370 | 0.80 ± 0.50 | 14 | 1.96 ± 0.44 | 356 | 0.75 ± 0.44 | < 0.001 |
| Esomeprazole | 317 | 0.84 ± 0.48 | 17 | 1.78 ± 0.37 | 300 | 0.79 ± 0.43 | < 0.001 |
| Warfarin | 426 | 1.28 ± 0.76 | 110 | 2.19 ± 0.71 | 316 | 0.97 ± 0.47 | < 0.001 |
| Aspirin | 377 | 1.34 ± 0.68 | 100 | 2.18 ± 0.50 | 277 | 1.04 ± 0.45 | < 0.001 |
| Clopidogrel | 476 | 1.20 ± 0.66 | 104 | 2.08 ± 0.56 | 372 | 0.95 ± 0.45 | < 0.001 |
IC-based signals were divided into 2 groups based on whether adverse events were also detected by the EBGM method.
Discussion
Several studies previously compared data mining algorithms [13, 24-29]; however, as Bate and Evans recently concluded [7], different algorithms have slightly different properties such that one may consequently be preferable in a particular application. If used for pharmacovigilance, data mining algorithms should be assessed from the standpoint of early and timely signal detection [30-33]. Although few studies have published comparative data, Chen et al. recently compared the timing of early signal detection with PRR, ROR, IC, and EBGM using the FAERS database, and concluded that the ROR performed better [30]. We previously reported that the ROR method provided the highest number of signals, while the EBGM method provided the lowest [23]. The difference in the number of signals can be attributed to a higher rate of false positives or lower ability to detect signals. In the present study, the commonality of signals was clarified, as shown in Figure 1. The EBGM method was shown to be the most conservative among the 4 methods, which suggested that it was suitable for pharmacoepidemiological studies. In contrast, the ROR method was shown to be the most comprehensive, indicating its usefulness for pharmacovigilance. These results were consistent with the findings of Chen et al [30]. These 4 data mining algorithms were used in our previous studies [16-22], and adverse events were listed as drug-associated, when at least 1 of the 4 indices met the criteria. However, the results shown in Figure 1 demonstrated that lists of adverse events were only identical when the ROR method was applied, which suggested that care should be taken in interpreting data when signals are not detected by the EBGM method.
Based on the number of signals, 16 drugs could be classified into 4 groups. Group 1 included 2 antimicrobials, which were characterized by the lower number of signals. The total number of co-occurrences with colistin was only 1,491, and 1,906 for tigecycline. These were markedly less than those of the other 14 drugs; from 33,197 with oxaliplatin to 220,194 with atorvastatin. The lower number of signals can be explained by comparatively infrequent use, and, therefore, a smaller number of reports in the database. This is not related to the reliability of the signals.
Group 2 included 4 statins and 2 PPIs characterized by a lower number of EBGM-based signals, and group 3 included warfarin and 2 antiplatelets by a higher number of EBGM-based signals. Group 4 included 5 anticancer agents characterized by a much higher number of EBGM-based signals. The total number of ROR-based signals was similar among drugs in groups 2-4; from 619 with rosuvastatin to 884 with cisplatin. The ROR method is feasible for detecting more signals, including false positives, than the EBGM method. The difference observed in the ratio of EBGM-based to ROR-based signals may reflect whether adverse events are generally found.
A pilot study performed by Hochberg et al. in 2009 concerning drug-versus-drug comparisons revealed that the rank-order of adverse event rates in the FAERS database was consistent with the results of published studies [34], which encouraged the use of the database for comparisons. In other investigations, the number of reports with or without normalization by usage or sales during the corresponding period was used to compare drugs [35]; however, adverse events are underreported, which may lead to incorrect conclusions [36-38]. Signal scores have also been considered inappropriate for determining the rank-order of drugs in terms of risk; however, few studies have been published to date. In the present study, the EBGM method was shown to be the most conservative among the 4 methods; therefore, it is important to confirm whether this method can provide important information similar to that in well-organized clinical studies.
Acknowledgements
This study was supported by the Funding Program for Next Generation World-Leading Researchers, Japan, and a Grant-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Rodriguez EM, Staffa JA, Graham DJ. The role of databases in drug postmarketing surveillance. Pharmacoepidemiol Drug Saf. 2001;10:407-410
2. Wysowski DK, Swartz L. Adverse drug event surveillance and drug withdrawals in the United States, 1969-2002: the importance of reporting suspected reactions. Arch Intern Med. 2005;165:1363-1369
3. US Food and Drug Administration (FDA). http://www.fda.gov/Drugs/GuidanceComplianceRegulatoryInformation/Surveillance/AdverseDrugEffects/default.htm
4. MedDRA MSSO. http://www.meddramsso.com/index.asp
5. Moore TJ, Cohen MR, Furberg CD. Serious adverse drug events reported to the Food and Drug Administration, 1998-2005. Arch Intern Med. 2007;167:1752-1759
6. Weiss-Smith S, Deshpande G, Chung S. et al. The FDA drug safety surveillance program: adverse event reporting trends. Arch Intern Med. 2011;171:591-593
7. Bate A, Evans SJ. Quantitative signal detection using spontaneous ADR reporting. Pharmacoepidemiol Drug Saf. 2009;18:427-436
8. Hauben M, Reich L. Drug-induced pancreatitis: lessons in data mining. Br J Clin Pharmacol. 2004;58:560-562
9. Almenoff J, Tonning JM, Gould AL. et al. Perspectives on the use of data mining in pharmaco-vigilance. Drug Saf. 2005;28:981-1007
10. Almenoff JS, Pattishall EN, Gibbs TG. et al. Novel statistical tools for monitoring the safety of marketed drugs. Clin Pharmacol Ther. 2007;82:157-166
11. Hauben M, Bate A. Decision support methods for the detection of adverse events in post-marketing data. Drug Discov Today. 2009;14:343-357
12. Evans SJ, Waller PC, Davis S. Use of proportional reporting ratios (PRRs) for signal generation from spontaneous adverse drug reaction reports. Pharmacoepidemiol Drug Saf. 2001;10:483-486
13. van Puijenbroek EP, Bate A, Leufkens HG. et al. A comparison of measures of disproportionality for signal detection in spontaneous reporting systems for adverse drug reactions. Pharmacoepidemiol Drug Saf. 2002;11:3-10
14. Bate A, Lindquist M, Edwards IR. et al. A Bayesian neural network method for adverse drug reaction signal generation. Eur J Clin Pharmacol. 1998;54:315-321
15. Szarfman A, Machado SG, O'Neill RT. Use of screening algorithms and computer systems to efficiently signal higher-than-expected combinations of drugs and events in the US FDA's spontaneous reports database. Drug Saf. 2002;25:381-392
16. Sakaeda T, Kadoyama K, Okuno Y. Adverse event profiles of platinum agents: Data mining of the public version of the FDA Adverse Event Reporting System, AERS, and reproducibility of clinical observations. Int J Med Sci. 2011;8:487-491
17. Sakaeda T, Kadoyama K, Tamon A. et al. Data mining of the public version of the FDA Adverse Event Reporting System, AERS: Colistin-associated adverse events. Jpn J Chemother. 2011;59:610-613
18. Kadoyama K, Miki I, Tamura T. et al. Adverse event profiles of 5-fluorouracil and capecitabine: Data mining of the public version of the FDA Adverse Event Reporting System, AERS, and reproducibility of clinical observations. Int J Med Sci. 2012;9:33-39
19. Sakaeda T, Kadoyama K, Okuno Y. Statin-associated muscular and renal adverse events: Data mining of the public version of the FDA Adverse Event Reporting System. PLoS ONE. 2011;6:e28124
20. Kadoyama K, Sakaeda T, Tamon A. et al. Adverse event profile of tigecycline: Data mining of the public version of the US Food and Drug Administrtation Adverse Event Reporting System. Biol Pharm Bull. 2012;35:967-970
21. Tamura T, Sakaeda T, Kadoyama K. et al. Omeprazole- and esomeprazole-associated hypomagnesaemia: Data mining of the public version of the FDA Adverse Event Reporting System. Int J Med Sci. 2012;9:322-326
22. Tamura T, Sakaeda T, Kadoyama K. et al. Aspirin- and clopidogrel-associated bleeding complications: Data mining of the public version of the FDA Adverse Event Reporting System. Int J Med Sci. 2012;9:441-446
23. Sakaeda T, Tamon A, Kadoyama K, Okuno Y. Data mining of the public version of the FDA Adverse Event Reporting System. Int J Med Sci. 2013;10:796-803
24. Gould AL. Practical pharmacovigilance analysis strategies. Pharmacoepidemiol Drug Saf. 2003;12:559-574
25. Rothman KJ, Lanes S, Sacks ST. The reporting odds ratio and its advantages over the proportional reporting ratio. Pharmacoepidemiol Drug Saf. 2004;13:519-523
26. Waller P, van Puijenbroek E, Egberts A. et al. The reporting odds ratio versus the proportional reporting ratio: 'deuce'. Pharmacoepidemiol Drug Saf. 2004;13:525-526
27. Matsushita Y, Kuroda Y, Niwa S. et al. Criteria revision and performance comparison of three methods of signal detection applied to the spontaneous reporting database of a pharmaceutical manufacturer. Drug Saf. 2007;30:715-726
28. Hochberg AM, Hauben M, Pearson RK. et al. An evaluation of three signal-detection algorithms using a highly inclusive reference event database. Drug Saf. 2009;32:509-525
29. Hochberg AM, Hauben M. Time-to-signal comparison for drug safety data-mining algorithms vs. traditional signaling criteria. Clin Pharmacol Ther. 2009;85:600-606
30. Chen Y, Guo JJ, Steinbuch M. et al. Comparison of sensitivity and timing of early signal detection of four frequently used signal detection methods: An empirical study based on the US FDA Adverse Event Reporting System database. Pharm Med. 2008;22:359-365
31. Motola D, Piccinni C, Biagi C. et al. Cardiovascular, ocular and bone adverse reactions associated with thiazolidinediones: a disproportionality analysis of the US FDA adverse event reporting system database. Drug Saf. 2012;35:315-323
32. Sommet A, Grolleau S, Bagheri H. et al. Was the thrombotic risk of rofecoxib predictable from the French Pharmacovigilance Database before 30 September 2004? Eur J Clin Pharmacol. 2008;64:829-834
33. Lindquist M, Ståhl M, Bate A. et al. A retrospective evaluation of a data mining approach to aid finding new adverse drug reaction signals in the WHO international database. Drug Saf. 2000;23:533-542
34. Hochberg AM, Pearson RK, O'Hara DJ. et al. Drug-versus-drug adverse event rate comparisons: a pilot study based on data from the US FDA Adverse Event Reporting System. Drug Saf. 2009;32:137-146
35. Bailie GR. Comparison of rates of reported adverse events associated with i.v. iron products in the United States. Am J Health Syst Pharm. 2012;69:310-320
36. Chertow GM, Mason PD, Vaage-Nilsen O. et al. Update on adverse drug events associated with parenteral iron. Nephrol Dial Transplant. 2006;21:378-382
37. Wysowski DK, Swartz L, Borders-Hemphill BV. et al. Use of parenteral iron products and serious anaphylactic-type reactions. Am J Hematol. 2010;85:650-654
38. Auerbach M, Kane RC. Caution in making inferences from FDA's Adverse Event Reporting System. Am J Health Syst Pharm. 2012;69:922-923
Author contact
Corresponding author: Toshiyuki Sakaeda, Ph.D., Center for Integrative Education in Pharmacy and Pharmaceutical Sciences, Graduate School of Pharmaceutical Sciences, Kyoto University, Kyoto 606-8501, Japan, Tel: +81-75-753-9560, Fax: +81-75-753-9253, e-mail: sakaedatkyoto-u.ac.jp; Yasushi Okuno, Ph.D., Department of Systems Biosciences for Drug Discovery, Graduate School of Pharmaceutical Sciences, Kyoto University, Kyoto 606-8501, Japan, Tel&Fax: +81-75-753-4559, e-mail: okunokyoto-u.ac.jp.

 Global reach, higher impact
Global reach, higher impact