Impact Factor ISSN: 1449-1907
Int J Med Sci 2013; 10(1):34-42. doi:10.7150/ijms.5270 This issue Cite
Research Paper
GATA5 loss-of-Function Mutations Underlie Tetralogy of Fallot
1. Department of Pediatrics, Tongji Hospital, Tongji University School of Medicine, Shanghai 200065, China;
2. Department of Cardiology and Cardiovascular Research, Shanghai Chest Hospital, Shanghai Jiaotong University School of Medicine, 241 West Huaihai Road, Shanghai 200030, China.
# contributed equally to this work.
Received 2012-9-21; Accepted 2012-11-27; Published 2012-12-10
Abstract
Tetraology of Fallot (TOF) is the most common form of cyanotic congenital heart disease and is a major cause of significant morbidity and mortality. Emerging evidence demonstrates that genetic risk factors are involved in the pathogenesis of TOF. However, TOF is genetically heterogeneous and the genetic defects responsible for TOF remain largely unclear. In the present study, the whole coding region of the GATA5 gene, which encodes a zinc-finger transcription factor essential for cardiogenesis, was sequenced in 130 unrelated patients with TOF. The relatives of the index patients harboring the identified mutations and 200 unrelated control individuals were subsequently genotyped. The functional characteristics of the mutations were analyzed using a luciferase reporter assay system. As a result, 2 novel heterozygous GATA5 mutations, p.R187G and p.H207R, were identified in 2 families with autosomal dominantly inherited TOF, respectively. The variations were absent in 400 control alleles and the altered amino acids were completely conserved evolutionarily. Functional analysis showed that the GATA5 mutants were associated with significantly decreased transcriptional activation compared with their wild-type counterpart. To our knowledge, this is the first report on the association of GATA5 loss-of-function mutations with TOF, suggesting potential implications for the early prophylaxis and allele-specific therapy of human TOF.
Keywords: Congenital heart disease, Tetralogy of Fallot, Genetics, Transcription factor, GATA5.
Introduction
Congenital heart disease constitutes the most prevalent type of developmental abnormality in humans with an estimated prevalence of 1% among live births, and is the leading noninfectious cause of infant death worldwide, with approximately 30% of neonates who die of birth defects having cardiovascular malformations [1]. Congenital cardiovascular anomalies are clinically classified into at least 21 different categories with specific anatomic lesions, of which tetraology of Fallot (TOF), a tetrad consisting of pulmonary stenosis, overriding aorta, ventricular septal defect, and right ventricular hypertrophy, is the most common form of cyanotic congenital heart disease, accounting for roughly 10% of all congenital cardiovascular deformations [1,2]. TOF can occur separately or in combination with other congenital cardiac deformities, such as atrial septal defect, patent ductus arteriosis, atrioventricular septal defect, abnormal pulmonary venous return, anomalous coronary arteries, absent pulmonary valve, aorticopulmonary window, and aortic incompetence. Regardless of other complications that may be concomitant with TOF, isolated TOF may result in reduced exercise tolerance, poor quality of life, delayed fetal brain development, cardiac dysfunction or heart failure, thromboembolic stroke, pulmonary embolus, subacute bacterial endocarditis, arrhythmias, and even cardiac death [3-12]. Given no corrective surgery, 25% of patients with severe pulmonary obstruction decease within the first year, 40% decease by age 3 years, 70% by age 10 years, and 95% by age 40 years [13]. Despite the significant morbidity and mortality throughout life, the fundamental etiology responsible for TOF remains largely unclear.
Embryonic heart development is a complex and dynamic process that requires the orchestration of cardiac cell commitment, differentiation, proliferation, and migration [14]. Many factors have been involved in this process based on their temporal and spatial expression patterns and their phenotypic characteristics associated with loss or gain of function, of which transcription factors are increasingly recognized as playing critical roles in early cardiogenesis, including GATA and NK families [14-16].
The GATA zinc finger-containing transcription factors are a family of DNA binding proteins characteristic of the ability to binding preferentially to the consensus DNA sequence GATA of target genes. To date, six members of the GATA family have been identified in vertebrates (GATA1 to GATA6), of which GATA4, GATA5 and GATA6 are broadly expressed in various mesoderm and endoderm derived tissues, dominantly in embryonic heart [15,16]. Among the three cardiac GATA transcription factors, GATA4 has been most extensively explored, and a lot of GATA4 mutations have been identified in patients with various kinds of congenital heart disease, including ventricular septal defect, atrial septal defect, and TOF [17-33]. Recently, mutations in GATA6 have also been associated with a wide variety of congenital heart disease [34-37]. The GATA5 gene is another member of the GATA family, and its expression and function overlap at least partially with those of GATA4 and GATA6 during cardiovascular development [14,16], which makes it a logical candidate gene for TOF.
Materials and Methods
Study subjects
A cohort of 130 unrelated patients with TOF was recruited among Chinese Han population. The available relatives of the index patients carrying the identified GATA5 mutations were also enrolled. Subjects were evaluated by individual and familial histories, review of the medical records, complete physical examination, 12-lead electrocardiogram, and two-dimensional transthoracic echocardiography with color flow Doppler. All patients had an echocardiogram documented TOF. Most patients underwent cardiac surgery. The patients with known chromosomal abnormalities or syndromic cardiovascular defects were excluded from the study.
A total of 200 unrelated, ethnically matched healthy individuals randomly selected from the individuals undergoing routine physical examinations were used as control subjects to scan for likely variations in GATA5. According to a review of medical history and an analysis of echocardiographic record, the control individuals had no congenital cardiovascular deformations, except for subclinical cardiac aberrations such as bicuspid aortic valve and patent foramen ovale. The ethnic origin of a participant was ascertained by a combination of self-reported ethnicity and a personal questionnaire asking questions about the birthplace, language, religion, and ancestry.
Peripheral venous blood specimens from TOF cases and control individuals were prepared. The study protocol was reviewed and approved by the local institutional ethics committee and written informed consent was obtained from all participants or their guardians prior to study.
GATA5 genotyping
Genomic DNA from all participants was extracted from blood lymphocytes with Wizard Genomic DNA Purification Kit (Promega, Madison, WI, USA). The candidate gene GATA5 was sequenced initially in 130 unrelated patients with TOF, and genotyping GATA5 was performed subsequently for the available relatives of the index patients carrying identified mutations and the 200 unrelated, ethnically matched healthy control individuals. The referential genomic DNA sequence of GATA5 derived from GenBank (accession No. HM015595). By the aid of on-line Primer 3 software (http://frodo.wi.mit.edu), the primer pairs used to amplify the coding exons and exon/intron boundaries of GATA5 by PCR were designed as shown in Table 1. The PCR was carried out using HotStar Taq DNA Polymerase (Qiagen GmbH, Hilden, Germany) on a PE 9700 Thermal Cycler (Applied Biosystems, Foster, CA, USA), with standard conditions and concentrations of reagents. Amplified products were analyzed on 1% agarose gels stained with ethidium bromide and purified with QIAquick Gel Extraction Kit (Qiagen). Both strands of each PCR product were sequenced with a BigDye® Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems) under an ABI PRISM 3130 XL DNA Analyzer (Applied Biosystems). The sequencing primers were the same as previously designed for specific region amplification. The DNA sequences were viewed and analyzed with the DNA Sequencing Analysis Software v5.1 (Applied Biosystems). The variant was validated by re-sequencing an independent PCR-generated amplicon from the subject and met our quality control thresholds with a call rate >99%. Additionally, an identified sequence variation was searched in the single nucleotide polymorphism (SNP) database at National Center for Biotechnology Information (NCBI) to confirm its novelty.
The intronic primers to amplify the coding exons and flanking splice sites of GATA5.
| Exon | Forward primer (5′ to 3′) | Reverse primer (5′ to 3′) | Amplicon (bp) |
|---|---|---|---|
| 2 | GGC, ATA, AGC, TCG, GGC, GCT, GG | TGG, GCC, CCG, AGA, CTG, TGG, AG | 648 |
| 3 | TGA, CGA, AAG, CCG, CCA, GGC, TC | CCC, CAG, GGG, CTC, TGG, TGT, CA | 375 |
| 4 | CCG, CAA, GGC, CGA, CCT, GAG, TC | CCG, CTC, CTC, CCC, AGC, CTC, TT | 312 |
| 5 | GGG, AAT, CCA, GCT, CCA, CGG, GC | CTG, GAG, GCA, CCG, AAG, GCC, AC | 331 |
| 6 | GCC, TGC, GGT, GTG, ACC, GTG, AG | GGT, GTG, TCC, AGC, CCA, CCT, GC | 370 |
| 7 | CCC, CCA, TGC, CAT, TCC, AGG, GC | GGG, GCC, TGC, TGG, TCT, CTG, CT | 402 |
Alignment of multiple GATA5 protein sequences
Multiple GATA5 protein sequences across various species were aligned using the online program of MUSCLE, version 3.6 (http://www.ncbi.nlm.nih.gov ).
Prediction of the causative potential of a GATA5 sequence variation
The disease-causing potential of a GATA5 sequence variation was predicted by MutationTaster (an online program at http://www.mutationtaster.org), which automatically gave a probability for the variation to be either a pathogenic mutation or a benign polymorphism. Notably, the P value used here is the probability of the correct prediction rather than the probability of error as used in t test statistics (i.e., a value close to 1 indicates a high 'security' of the prediction).
Construction of recombinant pcDNA3.1-hGATA5 expression plasmid
Human fetal cardiac tissue samples were previously collected and preserved in RNAlater RNA stabilization reagent (Qiagen). Total RNA was extracted using an RNeasy Protect Mini Kit (Qiagen). Reverse transcription was performed with Oligo(dT)20 primer using SuperScript III reverse transcriptase (Invitrogen, Carlsbad, CA, USA). The full-length wild-type cDNA of the human GATA5 gene, including partial 5'- and 3'-untranslated regions, was PCR amplified using pfuUltra high-fidelity DNA polymerase (Stratagene, La Jolla, CA, USA). The primer pairs used for the specific amplification of the GATA5 transcript were: for the forward, 5'-GTA, GCT, AGC, CAC, CGC, CGT, GCC, CTG, CCG-3'; for the reverse, 5'-GAT, GCG, GCC, GCT, GTT, CCC, CTG, ACA, TGG, GC-3'. The PCR fragment with a length of 1296 base pairs was doubly digested by endonuclease NheI and NotI (TaKaRa, Dalian, Liaoning, China). The digested product was fractionated by 1.5% agarose gel electrophoresis, purified with the QIAquick Gel Extraction Kit (Qiagen), and then subcloned into pcDNA3.1 (Promega) to form a recombinant eukaryotic expression vector, pcDNA3.1-hGATA5.
Site-directed mutagenesis
The identified mutation was introduced into the wild-type GATA5 using a QuickChange II XL Site-Directed Mutagenesis Kit (Stratagene) with a complementary pair of primers. The mutant was sequenced to confirm the desired mutation and to exclude any other sequence variations.
Reporter gene assay
The atrial natriuretic factor (ANF)-luciferase reporter gene, which contains the 2600-bp 5'-flanking region of the ANF gene, namely, ANF(-2600)-Luc, was kindly provided by Dr. Ichiro Shiojima from Chiba University School of Medicine (Chiba-shi, Chiba, Japan). HEK-293 cells were cultured in Dulbecco's modified Eagle's medium supplemented with 10% fetal calf serum. The ANF(-2600)-Luc reporter construct and an internal control reporter plasmid pGL4.75 (hRluc/CMV, Promega) were used in transient transfection assay to explore the transcriptional activation function of the GATA5 mutants. HEK-293 cells were transfected with 0.4 μg of wild-type or mutant pcDNA3.1-hGATA5 expression vector, 0.4 μg of ANF(-2600)-Luc reporter construct, and 0.04 μg of pGL4.75 control reporter vector using PolyFect Transfection Reagent (Qiagen). For co-transfection experiments, 0.2 μg of wild-type pcDNA3.1-hGATA5 together with 0.2 μg of mutant pcDNA3.1-hGATA5 or 0.2 μg of empty pcDNA3.1 vector were used in the presence of 0.4 μg of ANF(-2600)-Luc and 0.04 μg of pGL4.75. Firefly luciferase and Renilla luciferase activities were measured with the Dual-Glo luciferase assay system (Promega) 48h after transfection. The activity of the ANF promoter was presented as fold activation of Firefly luciferase relative to Renilla luciferase. Three independent experiments were performed at minimum for wild-type and mutant GATA5.
Statistical analysis
Data are expressed as means ± standard deviations. Continuous variables were tested for normality of distribution, and Student's unpaired t test was used for comparison of numeric variables between two groups. Comparison of the categorical variables between two groups was performed using Pearson's χ2 test or Fisher's exact test when appropriate. A two-tailed P value less than 0.05 was considered to be statistically significant.
Results
Baseline characteristics of the study subjects
A cohort of 130 unrelated patients with TOF was requited and clinically evaluated in contrast to a total of 200 unrelated, ethnically-matched healthy individuals used as the controls. All the participants had no established environmental risk factors for congenital heart disease, such as maternal illness and drug use in the first trimester of pregnancy, parental smoking, and long-term exposure to toxicants and ionizing radiation. The baseline clinical characteristics of the 130 unrelated TOF cases are summarized in Table 2.
Clinical characteristics of the 130 unrelated patients with tetralogy of Fallot (TOF).
| Parameter | n or mean | % or range |
|---|---|---|
| Male | 71 | 55 |
| Age (years) | 3.4 | 1-32 |
| Positive family history | 56 | 43 |
| Prevalence of TOF with other cardiac defects | ||
| Isolated TOF | 98 | 75 |
| TOF and ASD | 12 | 9 |
| TOF and PDA | 10 | 8 |
| TOF and AS | 6 | 5 |
| TOF and ASD and PDA | 4 | 3 |
| Incidence of arrhythmias | ||
| Atrioventricular block | 12 | 9 |
| Atrial fibrillation | 9 | 7 |
| Treatment | ||
| Surgical repair | 127 | 98 |
| Follow-up | 3 | 2 |
ASD: atrial septal defect, PDA: patent ductus arteriosus, AS: aortic stenosis.
GATA5 sequence variation
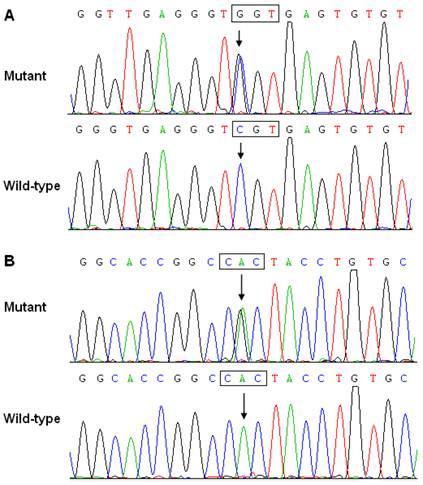
The entire coding region and the flanking splice sites of the GATA5 gene was sequenced in the 130 unrelated patients with TOF. Two heterozygous sequence variations in GATA5 were identified in 2 out of 130 patients, respectively. The total population prevalence of GATA5 variation based on the cohort patients was about 1.54%. Specifically, A transversion of cytosine into guanine at the first nucleotide of codon 187 of the GATA5 gene (c.559C>G), equivalent to the transition of arginine into glycine at amino acid position 187 (p.R187G), was identified in the index patient from family 1. A substitution of guanine for adenine in the second nucleotide of codon 207 (c.620A>G), equal to the replacement of histidine by arginine at amino acid 207 (p.H207R), was identified in the proband from family 2. The sequence electropherograms showing the identified heterozygous GATA5 variations in contrast to corresponding control sequences are illustrated in Figure 1. A schematic diagram of GATA5 showing the structural domains and the locations of the detected mutations is presented in Figure 2. The variations were neither observed in 400 control chromosomes nor found in the NCBI's SNP database, which was consulted again on September 22, 2012 (http://www.ncbi.nlm.nih.gov/SNP).
Sequence electropherograms showing the GATA5 mutations in contrast to the corresponding controls. The arrow indicates the heterozygous nucleotides of C/G in the proband from family 1 or A/G in the proband from family 2 (mutant); or the homozygous nucleotides of C/C or A/A in the corresponding control individuals (wild-type). The square denotes the nucleotides constituting a codon of GATA5.
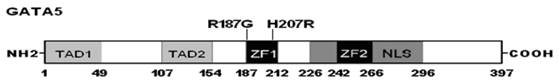
Schematic diagram of GATA5 protein structure with the tetralogy of Fallot related mutations shown. The mutations identified in patients with tetralogy of Fallot are shown above the structural domains. NH2 means amino-terminus; TAD, transcriptional activation domain; ZF, zinc finger; NLS, nuclear localization signal; and COOH, carboxyl-terminus.
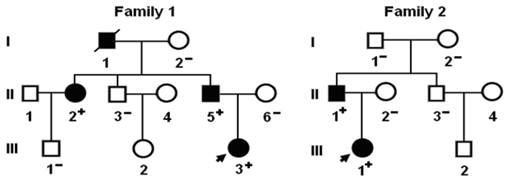
A genetic scan of the mutation carriers' family members demonstrated that in each family, the variation was present in all affected family members available, but absent in unaffected family members examined. Analysis of the pedigrees showed that the variations co-segregated with TOF with complete penetrance. The pedigree structures of the 2 families are shown in Figure 3. Additionally, in the family 1, the proband's grandfather (I-1) had also atrial septal defect and electrocardiogram documented atrial fibrillation, and her aunt (II-2) had also patent ductus arteriosus and paroxysmal atrial fibrillation. The phenotypic characteristics and results of genetic screening of the affected pedigree members are listed in Table 3.
Pedigree structures of the families with tetralogy of Fallot. Families are designated as family 1 and family 2, respectively. Family members are identified by generations and numbers. Squares indicate male family members; circles, female members; closed symbols, affected members; open symbols, unaffected members; a symbol with a slash, the deceased member; arrows, probands; ''+'', carriers of the heterozygous mutations; and ''−'', non-carriers.
Phenotypic characteristics and status of the GATA5 mutations of the affected pedigree members.
| Subject information | Phenotypes | Genotypes | |||||
|---|---|---|---|---|---|---|---|
| Identity | Gender | Age at time of Study (years) | Age at diagnosis of TOF (years) | Congenital cardiac structural defects | GATA5 mutations | ||
| Family 1 | R187G | ||||||
| I-1 | M | 55a | NA | TOF, ASD | NA | ||
| II-2 | F | 35 | 1 | TOF | ± | ||
| II-5 | M | 28 | 1 | TOF, PDA | ± | ||
| III-3 | F | 2 | 0 | TOF | ± | ||
| Family 2 | H207R | ||||||
| II-1 | M | 32 | 1 | TOF | ± | ||
| III-1 | F | 3 | 0 | TOF | ± | ||
M: male, F: female, TOF: tetralogy of Fallot, ASD: atrial septal defect, PDA: patent ductus arteriosus, NA: not available or not applicable. ±: heterozygosity. a: age at death.
Alignment of multiple GATA5 protein sequence
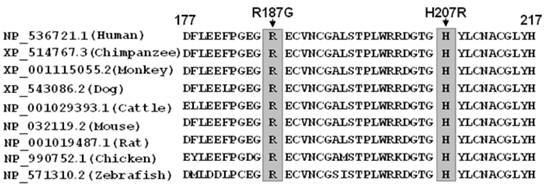
A cross-species alignment of multiple GATA5 protein sequences demonstrated that the affected amino acids were highly conserved evolutionarily (Figure 4), indicating that the amino acids are functionally important.
Alignment of multiple GATA5 protein sequences across species. The altered amino acids of p.R187 and p.H207 are completely conserved evolutionarily.

Causative potential of GATA5 sequence variations
The GATA5 sequence variations of c.559C>G and c.620A>G were both automatically predicted to be disease-causing, with P values of 0.99994 and 0.99934, respectively. No SNPs in the altered regions were found in MutationTaster database.
Transcriptional activity of the GATA5 mutants
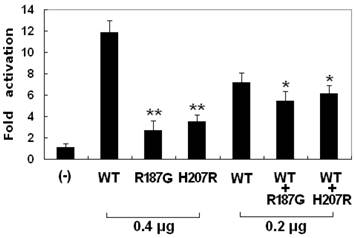
As shown in Figure 5, the wild-type GATA5, the R187G-mutant, and the H207R-mutant GATA5 activated the ANF promoter by ~12-fold, ~3-fold, and ~4-fold, respectively. When wild-type GATA5 was co-expressed with the same amount of R187G-mutant or H207R-mutant GATA5, the induced activation of the ANF promoter was ~5-fold or ~6-fold. These results indicate that both GATA5 mutants are associated with significantly reduced activation activity compared with their wild-type counterpart.
Functional defects of GATA5 caused by mutations. Activation of ANF-luciferase reporter in HEK-293 cells by wild-type GATA5 (WT), or mutant (R187G or H207R), alone or in combination, revealed significantly decreased transactivational activity by mutant proteins. Experiments were performed in triplicate and mean and standard deviations are shown. ** and * represent P < 0.001 and P < 0.005, respectively, when compared with wild-type GATA5.
Discussion
In the present study, two novel heterozygous mutations of GATA5, p.R187G and p.H207R, were identified in 2 families with TOF. In each family, the mutant allele was present in all the affected family members alive but absent in unaffected relatives examined and 400 normal chromosomes from an ethnically matched control population. A cross-species alignment of GATA5 protein sequences showed that the altered amino acids were highly conserved evolutionarily. The p.R187G and p.H207R variants were both predicted to be pathogenic mutations, and the biochemical analysis revealed that the GATA5 mutant proteins were consistently associated with significantly reduced transcriptional activation. Therefore, it is very likely that functionally impaired GATA5 is responsible for TOF in these families. To our knowledge, this is the first report on the association of GATA5 loss-of-function mutations with enhanced susceptibility to TOF in humans.
The human GATA5 gene maps to chromosome 20q13.33 by fluorescence in situ hybridization, encoding a protein of 397 amino acids [38]. By alignment of GATA5 with GATA4, the functional domains of GATA5 are predicted to consist of 2 transcriptional activation domains (TAD1, amino acids 1-49; TAD2, amino acids 107-154), 2 adjacent zinc fingers (ZF1, amino acids 187-212; ZF2, amino acids 242-266), and 1 nuclear localization signal (NLS, amino acids 226-396). The two TADs are both pivotal for the transcriptional activity of GATA5. The N-terminal ZF1 is essential for DNA sequence recognition and binding to the consensus motif, while the C-terminal ZF2 is responsible for sequence specificity and stability of protein-DNA binding. The NLS is a key to the sub-cellular trafficking and distribution of GATA5. The GATA5 mutations of p.R187G and p.H207R identified in this study are both located in ZF1, thus may be expected to have impact on the transcriptional activation of GATA5 by interfering with the recognition and binding of GATA5 with target gene promoter.
It has been substantiated that GATA5 is an upstream regulator of several genes expressed during embryogenesis and cardiac morphogenesis, including the genes that encode atrial natriuretic factor (ANF), brain natriuretic peptide, α-myosin heavy chain, β myosin heavy chain, and cardiac troponin C and I (15). Hence, the functional roles of the GATA5 mutations may be assessed by using biochemical analysis of the transcriptional activity of the ANF promoter in tool cells. In this study, the functional effect of the novel p.R187G and p.H207R mutations of GATA5 identified in our familial TOF patients were evaluated by transcriptional activity assay and the results showed a significantly decreased transcriptional activity on a downstream gene. The data suggest that GATA5 loss-of-function mutations are potentially an alternative molecular mechanism involved in TOF.
That functionally compromised GATA5 predisposes to congenital cardiovascular defects has been demonstrated in animal models. In zebrafish, targeted deletion of the GATA5 gene led to embryonic lethality due to defects in endocardial and myocardial differentiation and migration, a similar phenotype to cardia bifida of GATA4-null zebrafish [39]. Although the GATA5-knockout mice were viable and without apparent cardiac defects, the mice that were compound heterozygous for both GATA5 and GATA4 or for both GATA5 and GATA6 knockout died embryonically or perinatally because of severe defects of the outflow tract development including double outlet right ventricle and ventricular septal defect [40]. These experimental findings corroborate an exquisite sensitivity of the developing cardiovascular system to the levels of GATA4, GATA5 and GATA6, and suggest that these GATA factors mar act synergistically to regulate downstream target genes.
It was interesting that atrial fibrillation was documented in 2 TOF patients harboring the p.R187G mutation of GATA5. Similar to our findings, atrial fibrillation were previously confirmed in patients with congenital cardiovascular defects who carry GATA4, GATA5, or GATA6 mutations [41-47]. These observations imply that congenital heart disease may share a common genetic origin with atrial fibrillation. It has been shown that the abnormal development of heart, especially the pulmonary vein myocardium contributes to the initiation and perpetuation of atrial fibrillation by several possible pathological mechanisms including enhanced intrinsic pacemaker activity and increased properties that facilitate reentrance [48-50]. Genetic-labeling lineage tracing analyses have revealed that NKX2-5 is highly expressed in the atria and pulmonary myocardium and plays a pivotal role in the localized formation of the sinoatrial node during embryonic development. As a repressor of the sinoatrial node lineage gene program, NKX2-5 functions to limit pacemaker activity to the sinus node and the atrioventricular node [51]. Therefore, as an important transcriptionally cooperative partner of NKX2-5 [52], GATA5, when mutated, may be involved in the formation of the atrial electrophysiological substrate in favor of atrial fibrillation.
In conclusion, this work provides the first genetic evidence for the association of functionally compromised GATA5 with increased vulnerability to TOF, implying the potential implications for the early prophylaxis and personalized treatment of TOF.
Acknowledgements
We are really thankful to the subjects for their devotion to the study. This work was supported in part by grants from the National Natural Science Fund of China (81070153, 81270161 and 30570768), the Personnel Development Foundation of Shanghai, China (2010019), the Natural Science Fund of Shanghai, China (10ZR1428000 and 10ZR1433100), and the Key Program of Basic Research of Shanghai, China (10JC1414002).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Roger VL, Go AS, Lloyd-Jones DM, Benjamin EJ, Berry JD, Borden WB, Bravata DM, Dai S, Ford ES, Fox CS, Fullerton HJ, Gillespie C, Hailpern SM, Heit JA, Howard VJ, Kissela BM, Kittner SJ, Lackland DT, Lichtman JH, Lisabeth LD, Makuc DM, Marcus GM, Marelli A, Matchar DB, Moy CS, Mozaffarian D, Mussolino ME, Nichol G, Paynter NP, Soliman EZ, Sorlie PD, Sotoodehnia N, Turan TN, Virani SS, Wong ND, Woo D, Turner MB. American Heart Association Statistics Committee and Stroke Statistics Subcommittee. Heart disease and stroke statistics--2012 update: a report from the American Heart Association. Circulation. 2012;125:e2-220
2. van der Linde D, Konings EE, Slager MA, Witsenburg M, Helbing WA, Takkenberg JJ, Roos-Hesselink JW. Birth prevalence of congenital heart disease worldwide: a systematic review and meta-analysis. J Am Coll Cardiol. 2011;58:2241-7
3. Buys R, Cornelissen V, Van De Bruaene A, Stevens A, Coeckelberghs E, Onkelinx S, Thomaes T, Delecluse C, Budts W, Vanhees L. Measures of exercise capacity in adults with congenital heart disease. Int J Cardiol. 2011;153:26-30
4. Müller J, Hess J, Hager A. Exercise performance and quality of life is more impaired in Eisenmenger syndrome than in complex cyanotic congenital heart disease with pulmonary stenosis. Int J Cardiol. 2011;150:177-81
5. Teixeira FM, Coelho RM, Proença C, Silva AM, Vieira D, Vaz C, Moura C, Viana V, Areias JC, Areias ME. Quality of life experienced by adolescents and young adults with congenital heart disease. Pediatr Cardiol. 2011;32:1132-8
6. Müller J, Hess J, Hager A. Minor symptoms of depression in patients with congenital heart disease have a larger impact on quality of life than limited exercise capacity. Int J Cardiol. 2012;154:265-9
7. Shedeed SA, Elfaytouri E. Brain maturity and brain injury in newborns with cyanotic congenital heart disease. Pediatr Cardiol. 2011;32:47-54
8. Perry JC. Sudden cardiac death and malignant arrhythmias: the scope of the problem in adult congenital heart patients. Pediatr Cardiol. 2012;33:484-90
9. Silka MJ, Bar-Cohen Y. A contemporary assessment of the risk for sudden cardiac death in patients with congenital heart disease. Pediatr Cardiol. 2012;33:452-60
10. Knirsch W, Nadal D. Infective endocarditis in congenital heart disease. Eur J Pediatr. 2011;170:1111-27
11. Murakami T, Niwa K, Yoshinaga M, Nakazawa M. Factors associated with surgery for active endocarditis in congenital heart disease. Int J Cardiol. 2012;157:59-62
12. Cheng HH, Almodovar MC, Laussen PC, Wypij D, Polito A, Brown DW, Emani SM, Pigula FA, Allan CK, Costello JM. Outcomes and risk factors for mortality in premature neonates with critical congenital heart disease. Pediatr Cardiol. 2011;32:1139-46
13. Starr JP. Tetralogy of Fallot: Yesterday and today. World J Surg. 2010;34:658-68
14. Cecchetto A, Rampazzo A, Angelini A, Bianco LD, Padalino M, Stellin G, Daliento L. From molecular mechanisms of cardiac development to genetic substrate of congenital heart diseases. Future Cardiol. 2010;6:373-93
15. Pikkarainen S, Tokola H, Kerkelä R, Ruskoaho H. GATA transcription factors in the developing and adult heart. Cardiovasc Res. 2004;63:196-207
16. Peterkin T, Gibson A, Loose M, Patient R. The roles of GATA-4, -5 and -6 in vertebrate heart development. Semin Cell Dev Biol. 2005;16:83-94
17. Garg V, Kathiriya IS, Barnes R, Schluterman MK, King IN, Butler CA, Rothrock CR, Eapen RS, Hirayama-Yamada K, Joo K, Matsuoka R, Cohen JC, Srivastava D. GATA4 mutations cause human congenital heart defects and reveal an interaction with TBX5. Nature. 2003;424:443-7
18. Okubo A, Miyoshi O, Baba K, Takagi M, Tsukamoto K, Kinoshita A, Yoshiura K, Kishino T, Ohta T, Niikawa N, Matsumoto N. A novel GATA4 mutation completely segregated with atrial septal defect in a large Japanese family. J Med Genet. 2004;41:e97
19. Sarkozy A, Conti E, Neri C, D'Agostino R, Digilio MC, Esposito G, Toscano A, Marino B, Pizzuti A, Dallapiccola B. Spectrum of atrial septal defects associated with mutations of NKX2.5 and GATA4 transcription factors. J Med Genet. 2005;42:e16
20. Hirayama-Yamada K, Kamisago M, Akimoto K, Aotsuka H, Nakamura Y, Tomita H, Furutani M, Imamura S, Takao A, Nakazawa M, Matsuoka R. Phenotypes with GATA4 or NKX2.5 mutations in familial atrial septal defect. Am J Med Genet A. 2005;135:47-52
21. Reamon-Buettner SM, Borlak J. GATA4 zinc finger mutations as a molecular rationale for septation defects of the human heart. J Med Genet. 2005;42:e32
22. Nemer G, Fadlalah F, Usta J, Nemer M, Dbaibo G, Obeid M, Bitar F. A novel mutation in the GATA4 gene in patients with Tetralogy of Fallot. Hum Mutat. 2006;27:293-4
23. Tomita-Mitchell A, Maslen CL, Morris CD, Garg V, Goldmuntz E. GATA4 sequence variants in patients with congenital heart disease. J Med Genet. 2007;44:779-83
24. Rajagopal SK, Ma Q, Obler D, Shen J, Manichaikul A, Tomita-Mitchell A, Boardman K, Briggs C, Garg V, Srivastava D, Goldmuntz E, Broman KW, Benson DW, Smoot LB, Pu WT. Spectrum of heart disease associated with murine and human GATA4 mutation. J Mol Cell Cardiol. 2007;43:677-85
25. Zhang W, Li X, Shen A, Jiao W, Guan X, Li Z. GATA4 mutations in 486 Chinese patients with congenital heart disease. Eur J Med Genet. 2008;51:527-35
26. Hamanoue H, Rahayuningsih SE, Hirahara Y, Itoh J, Yokoyama U, Mizuguchi T, Saitsu H, Miyake N, Hirahara F, Matsumoto N. Genetic screening of 104 patients with congenitally malformed hearts revealed a fresh mutation of GATA4 in those with atrial septal defects. Cardiol Young. 2009;19:482-5
27. Chen MW, Pang YS, Guo Y, Pan JH, Liu BL, Shen J, Liu TW. GATA4 mutations in Chinese patients with congenital cardiac septal defects. Pediatr Cardiol. 2010;31:85-9
28. Chen Y, Mao J, Sun Y, Zhang Q, Cheng HB, Yan WH, Choy KW, Li H. A novel mutation of GATA4 in a familial atrial septal defect. Clin Chim Acta. 2010;411:1741-5
29. Butler TL, Esposito G, Blue GM, Cole AD, Costa MW, Waddell LB, Walizada G, Sholler GF, Kirk EP, Feneley M, Harvey RP, Winlaw DS. GATA4 mutations in 357 unrelated patients with congenital heart malformation. Genet Test Mol Biomarkers. 2010;14:797-802
30. Salazar M, Consoli F, Villegas V, Caicedo V, Maddaloni V, Daniele P, Caianiello G, Pachón S, Nuñez F, Limongelli G, Pacileo G, Marino B, Bernal JE, De Luca A, Dallapiccola B. Search of somatic GATA4 and NKX2.5 gene mutations in sporadic septal heart defects. Eur J Med Genet. 2011;54:306-9
31. Liu XY, Wang J, Zheng JH, Bai K, Liu ZM, Wang XZ, Liu X, Fang WY, Yang YQ. Involvement of a novel GATA4 mutation in atrial septal defects. Int J Mol Med. 2011;28:17-23
32. Wang J, Fang M, Liu XY, Xin YF, Liu ZM, Chen XZ, Wang XZ, Fang WY, Liu X, Yang YQ. A novel GATA4 mutation responsible for congenital ventricular septal defects. Int J Mol Med. 2011;28:557-64
33. Yang YQ, Li L, Wang J, Liu XY, Chen XZ, Zhang W, Wang XZ, Jiang JQ, Liu X, Fang WY. A novel GATA4 loss-of-function mutation associated with congenital ventricular septal defect. Pediatr Cardiol. 2012;33:539-46
34. Kodo K, Nishizawa T, Furutani M, Arai S, Yamamura E, Joo K, Takahashi T, Matsuoka R, Yamagishi H. GATA6 mutations cause human cardiac outflow tract defects by disrupting semaphorin-plexin signaling. Proc Natl Acad Sci U S A. 2009;106:13933-8
35. Lin X, Huo Z, Liu X, Zhang Y, Li L, Zhao H, Yan B, Liu Y, Yang Y, Chen YH. A novel GATA6 mutation in patients with tetralogy of Fallot or atrial septal defect. J Hum Genet. 2010;55:662-7
36. Maitra M, Koenig SN, Srivastava D, Garg V. Identification of GATA6 sequence variants in patients with congenital heart defects. Pediatr Res. 2010;68:281-5
37. Zheng GF, Wei D, Zhao H, Zhou N, Yang YQ, Liu XY. A novel GATA6 mutation associated with congenital ventricular septal defect. Int J Mol Med. 2012;29:1065-71
38. Nemer G, Qureshi ST, Malo D, Nemer M. Functional analysis and chromosomal mapping of Gata5, a gene encoding a zinc finger DNA-binding protein. Mamm Genome. 1999;10:993-9
39. Heicklen-Klein A, McReynolds LJ, Evans T. Using the zebrafish model to study GATA transcription factors. Semin Cell Dev Biol. 2005;16:95-106
40. Laforest B, Nemer M. GATA5 interacts with GATA4 and GATA6 in outflow tract development. Dev Biol. 2011;358:368-78
41. Posch MG, Boldt LH, Polotzki M, Richter S, Rolf S, Perrot A, Dietz R, Ozcelik C, Haverkamp W. Mutations in the cardiac transcription factor GATA4 in patients with lone atrial fibrillation. Eur J Med Genet. 2010;53:201-3
42. Yang YQ, Wang MY, Zhang XL, Tan HW, Shi HF, Jiang WF, Wang XH, Fang WY, Liu X. GATA4 loss-of-function mutations in familial atrial fibrillation. Clin Chim Acta. 2011;412:1825-30
43. Jiang JQ, Shen FF, Fang WY, Liu X, Yang YQ. Novel GATA4 mutations in lone atrial fibrillation. Int J Mol Med. 2011;28:1025-32
44. Wang J, Sun YM, Yang YQ. Mutation spectrum of the GATA4 gene in patients with idiopathic atrial fibrillation. Mol Biol Rep. 2012;39:8127-35
45. Yang YQ, Wang J, Wang XH, Wang Q, Tan HW, Zhang M, Shen FF, Jiang JQ, Fang WY, Liu X. Mutational spectrum of the GATA5 gene associated with familial atrial fibrillation. Int J Cardiol. 2012;157:305-7
46. Yang YQ, Wang XH, Tan HW, Jiang WF, Fang WY, Liu X. Prevalence and spectrum of GATA6 mutations associated with familial atrial fibrillation. Int J Cardiol. 2012;155:494-6
47. Yang YQ, Li L, Wang J, Zhang XL, Li RG, Xu YJ, Tan HW, Wang XH, Jiang JQ, Fang WY, Liu X. GATA6 loss-of-function mutation in atrial fibrillation. Eur J Med Genet. 2012;55:520-6
48. Haïssaguerre M, Jaïs P, Shah DC, Takahashi A, Hocini M, Quiniou G, Garrigue S, Le Mouroux A, Le Métayer P, Clémenty J. Spontaneous initiation of atrial fibrillation by ectopic beats originating in the pulmonary veins. N Engl J Med. 1998;339:659-66
49. Nattel S. Basic electrophysiology of the pulmonary veins and their role in atrial fibrillation: precipitators, perpetuators, and perplexers. J Cardiovasc Electrophysiol. 2003;14:1372-5
50. Mommersteeg MT, Brown NA, Prall OW, de Gier-de Vries C, Harvey RP, Moorman AF, Christoffels VM. Pitx2c and Nkx2-5 are required for the formation and identity of the pulmonary myocardium. Circ Res. 2007;101:902-9
51. Mommersteeg MT, Hoogaars WM, Prall OW, de Gier-de Vries C, Wiese C, Clout DE, Papaioannou VE, Brown NA, Harvey RP, Moorman AF, Christoffels VM. Molecular pathway for the localized formation of the sinoatrial node. Circ Res. 2007;100:354-62
52. Zhang Y, Rath N, Hannenhalli S, Wang Z, Cappola T, Kimura S, Atochina-Vasserman E, Lu MM, Beers MF, Morrisey EE. GATA and Nkx factors synergistically regulate tissue-specific gene expression and development in vivo. Development. 2007;134:189-98
Author contact
Corresponding author: Xing-Yuan Liu, Tel: +8621-56051080; Fax: +8621-66371663; Email: liuxingyuan789com. Yi-Qing Yang, Tel: +8621-62821990; Fax: +8621-62821105; Email: yang99yang66com.

 Global reach, higher impact
Global reach, higher impact