3.2
Impact Factor
ISSN: 1449-1907
Int J Med Sci 2012; 9(8):715-724. doi:10.7150/ijms.4815 This issue Cite
Research Paper
The SCARB1 rs5888 SNP and Serum Lipid Levels in the Guangxi Mulao and Han Populations
1. Department of Cardiology, Institute of Cardiovascular Diseases, the First Affiliated Hospital, Guangxi Medical University, 22 Shuangyong Road, Nanning 530021, Guangxi, People's Republic of China
2. Department of Pathophysiology, School of Premedical Sciences, Guangxi Medical University, Nanning 530021, Guangxi, People's Republic of China
Received 2012-7-4; Accepted 2012-9-10; Published 2012-10-13
Abstract
Backgroud: The associations of scavenger receptor class B type 1 (SCARB1) rs5888 single nucleotide polymorphism (SNP) and serum lipid levels are inconsistant among diverse ethnic populations. The present study was undertaken to detect the association of rs5888 SNP and serum lipid levels in the Guangxi Mulao and Han populations.
Methods: Genotypes of the SCARB1 rs5888 SNP in 801 subjects of Mulao and 807 subjects of Han Chinese were determined by polymerase chain reaction and restriction fragment length polymorphism combined with gel electrophoresis, and then confirmed by direct sequencing.
Results: Serum apolipoprotein (Apo) B levels and the T allelic frequency were higher in Mulao than in Han. Serum high-density lipoprotein cholesterol (HDL-C) levels in Mulao were different among the genotypes, the subjects with TT genotype had lower HDL-C levels than the subjects with CC or CT genotype in female (P < 0.05). For the Han population, serum triglyceride (TG), HDL-C, ApoAI, ApoB levels and the ratio of ApoAI to ApoB in males were different among the genotypes, the T allele carriers had lower serum HDL-C, ApoAI levels and ApoAI/ApoB ratio and higher serum ApoB levels than the T allele noncarriers (P < 0.05 for all), the subjects with TT genotype had higher serum TG levels than the subjects with CC or CT genotype. Serum HDL-C levels in Mulao females and serum HDL-C, ApoAI, ApoB levels and the ApoAI/ApoB ratio in Han males were correlated with genotypes by the multiple linear regression analysis. Serum lipid parameters were also influenced by genotype-environmental interactions in Han but not in Mulao populations.
Conclusions: These results suggest that the rs5888 SNP is associated with serum HDL-C levels in Mulao females, and TG, HDL-C, ApoAI, ApoB levels and the ApoAI/ApoB ratio in Han males. The differences in serum ApoB levels between the two ethnic groups might partially attribute to different SCARB1 genotype-environmental interactions.
Keywords: scavenger receptor class B type 1 gene, single nucleotide polymorphism, lipids, apolipoproteins
Introduction
Coronary heart disease (CHD) is the leading causes of death and disability worldwide [1]. Previous studies have demonstrated an inverse correlation between CHD and serum high-density lipoprotein cholesterol (HDL-C) [2,3]. Plasma HDL-C levels are important in determining risk for CHD, four prospective American studies showed a 1-mg/dl increment in HDL-C levels was associated with a significant CHD risk decrement of 2% in men and 3% in women [4]. Serum HDL-C levels are influenced by both environmental and genetic factors. It has been reported that variation in plasma HDL-C levels is at least 50% genetically determined [5]. Many studies have focused on the associations of variants in candidate gene with plasma HDL-C levels [6-8].
The scavenger receptor class B type 1 (SCARB1) was described as the first functionally active HDL receptor capable of facilitating the selective uptake of HDL-C, and it is expressed primarily in liver and steroidogenic tissues [9]. Several previous studies have demonstrated that SCARB1 not only involves in the regulation of HDL-C levels but also implicates in the metabolism of non-HDL-C levels in mouse models [10-17]. The human SCARB1 gene encodes a 509 amino acid protein with a molecular weight of 85 kDa [13], and located on chromosome 12q24, a region showing significant linkage to plasma triglyceride (TG) and HDL-C levels [18]. The associations of variants in the SCARB1 gene and serum lipid parameters have been investigated previously in diverse populations [19-27]. Among these variants in SCARB1 gene, the single nucleotide polymorphism (SNP) of rs5888 is quite common, and has been associated with plasma HDL-C and non-HDL-C concentrations, but with different allele and genotypic frequencies in diverse ethnic groups [19,20,21,23,24,27].
China is a multi-ethnic country. There are 56 ethnic groups. Han nationality is the largest ethnic group and Mulao nationality is one of the 55 minorities with population of 207,352 according to the fifth national census statistics of China in 2000. Ninety percent of them live in the Luocheng Mulao Autonomous County, Guangxi Zhuang Autonomous Region, People's Republic of China. The history of this minority can be traced back to the Jin Dynasty (AD265-420). A previous study has shown that the genetic relationship between Mulao nationality and other minorities in Guangxi was much closer than that between Mulao and Han or Uighur nationality [28]. In a recent study, we showed that the rs5888 SNP was associated with serum lipid levels in the Guangxi Bai Ku Yao population [29]. To our knowledge, however, the association of rs5888 SNP in the SCARB1 gene and serum lipid levels has not been reported previously in the Mulao population. Therefore, the present study was to evaluate the association of rs5888 SNP and several environmental factors with serum lipid levels in the Guangxi Mulao and Han populations.
Materials and methods
Study population
A total of 801 subjects of Mulao who reside in Luocheng Mulao Autonomous County, Guangxi Zhuang Autonomous Region, People's Republic of China and 807 subjects of Han Chinese reside in the same villages were included in the study. The subjects of Mulao consisted of 379 (47.32%) males and 422 (52.68%) females, ranged in age from 16 to 86 years, with a mean age of 52.55 ± 14.78 years. The subjects of Han consisted of 355 (43.99%) males and 452 (56.01%) females, aged 16-86 years, with a mean age of 51.40 ± 15.14 years. All of the subjects were randomly selected from our previous stratified randomized cluster samples [30,31]. All study subjects were healthy rural agricultural workers, and had no evidence of diseases related to atherosclerosis, CHD and diabetes. None of them were using lipid-lowering medication. The present study was approved by the Ethics Committee of the First Affiliated Hospital, Guangxi Medical University. Informed consent was obtained from all subjects after they received a full explanation of the study.
Epidemiological survey
The survey was carried out using internationally standardized methods [32]. Information on demographics, socioeconomic status, and lifestyle factors was collected with standardized questionnaires. The alcohol information included questions about the number of liangs (about 50 g) of rice wine, corn wine, rum, beer, or liquor consumed during the preceding 12 months. Alcohol consumption was categorized into groups of grams of alcohol per day: ≤ 25 and > 25. Smoking status was categorized into groups of cigarettes per day: ≤ 20 and > 20. At the physical examination, several parameters such as height, weight, and waist circumference were measured. Sitting blood pressure was measured using a mercury sphygmomanometer on 3 separated intervals after the subjects had a 5-minute rest, and the average of the three measurements was used for the level of blood pressure. Body mass index (BMI) was calculated as weight in kg divided by the square of height in meters (kg/m2).
Biochemical analysis
Venous blood samples were obtained from all subjects after at least 12 hours of fasting. The levels of serum total cholesterol (TC), TG, HDL-C, and low-density lipoprotein cholesterol (LDL-C) in samples were determined by enzymatic methods with commercially available kits. Serum apolipoprotein (Apo) AI and ApoB levels were detected by the immunoturbidimetric immunoassay using a commercial kit [30,31].
DNA amplification and genotyping
Genomic DNA was extracted from peripheral blood leukocytes using the phenol-chloroform method [29-31]. Genotyping of the SCARB1 rs5888 SNP was performed using mutagenic primer and restriction enzyme for restriction fragment length polymorphism (RFLP). A pair of primers was designed to introduce a Hin1I restriction site (GACGCC) by changing a base from A to G. PCR amplification was performed using 5'-CCTTGTTTCTCTCCCATCCTCACTTCCTCGACGC-3' as the forward and 5'-CACCACCCCAGCCCACAGCAGC-3' (Sangon, Shanghai, People's Republic of China) as the reverse primer pairs. Each 20 μL PCR reaction mixture consisted of 1 μL of genomic DNA, 0.5 μL of each primer (10 pmol/L), 10 μL of 2 × Taq PCR Mastermix (constituent: 20 mM Tris-HCl, pH 8.3, 100 mM KCl, 3 mM MgCl2, 0.1 U Taq Polymerase/μL, 500 μM dNTP each; Tiangen, Beijing, People's Republic of China), and 8 μL of ddH2O (DNase/RNase-free). The PCR condition comprised initial denaturation at 95°C for 5 min; 33 cycles of denaturation at 95°C for 45 s, annealing at 71.5°C for 30 s, extension at 72°C for 50 s, and a final extension at 72°C for 8 min. Then 5 μL of amplification products were digested at 37°C overnight with 5 U of Hin1I restriction enzyme (Fermentas Co. Canada). After restriction enzyme digestion of the amplified DNA, the fragments were separated by electrophoresis on 3% agarose gels stained with ethidium bromide, photographed in ultraviolet light. Genotypes were scored by an experienced reader blinded to epidemiological data and serum lipid levels. Six samples (CC, CT and TT genotypes in two; respectively) detected by the PCR-RFLP were also confirmed by sequencing.
Diagnostic criteria
The normal values of serum TC, TG, HDL-C, LDL-C, ApoAI, ApoB levels, and the ratio of ApoAI to ApoB in our Clinical Science Experiment Center were 3.10-5.17, 0.56-1.70, 0.91-1.81, 2.70-3.20 mmol/L, 1.00-1.78, 0.63-1.14 g/L, and 1.00-2.50; respectively [29,30]. Hypertension was diagnosed according to the criteria of 1999 World Health Organization-International Society of Hypertension Guidelines for the management of hypertension [33]. Normal weight, overweight and obesity were defined as a BMI < 24, 24-28, and > 28 kg/m2; respectively [34].
Statistical analyses
All statistical analyses were carried out using the statistical software package SPSS 13.0 (SPSS Inc., Chicago, Illinois). Qualitative variables were expressed as raw count and percentage. Mean ± standard deviation was used for the presentation of quantitative variables. Genotypic and allelic frequencies were calculated by direct counting, and the standard goodness-of-fit test was used to test the Hardy-Weinberg equilibrium. A chi-square analysis was used to evaluate the difference in genotype distribution and sex ratio between the groups. The difference in general characteristics between Mulao and Han was tested by the Student's unpaired t-test. The association of genotypes and serum lipid parameters was tested by analysis of covariance (ANCOVA). Sex, age, BMI, blood pressure, alcohol consumption, cigarette smoking were adjusted for the statistical analysis. In order to evaluate the association of serum lipid levels with genotypes (CC = 1, CT = 2, TT = 3; or CC = 1, CT/TT = 2; or CC/CT = 1, TT = 2), multiple linear regression analysis with stepwise modeling was also performed. The genotype-environmental interactions on serum lipid levels were tested using the factorial design covariance analysis. A two-tailed P value less than 0.05 was considered statistically significant.
Results
General characteristics and serum lipid levels
Table 1 shows the general characteristics and serum lipid parameters of the study populations. As compared with the Han population, Mulao has lower BMI, and higher serum ApoB levels and the percentages of subjects who consumed alcohol. The levels of age, height, weight, blood pressure, serum TC, TG, HDL-C, LDL-C, ApoAI, the ratio of ApoAI to ApoB; the percentages of subjects who smoked cigarette; and the ratio of male to female were not different between the two ethnic groups (P > 0.05 for all).
Results of electrophoresis, genotyping and sequencing
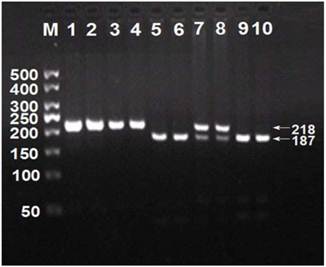
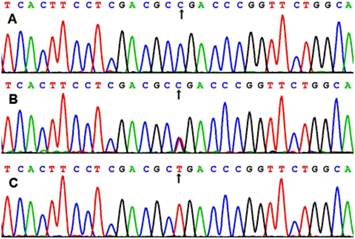
After the genomic DNA of the samples was amplified by PCR, the products of 218 bp nucleotide sequences imaged by 3.0% agarose gel electrophoresis could be found in all samples (lanes 1 and 2; Figure 1). The genotypes were called according the presence or absence of a restriction site when a C to T transversion at amino acid 350 of the SCARB1 gene. The presence of the cutting site indicates the C allele, while its absence indicates the T allele (cannot be cut). Thus, the TT genotype is homozygote for the absence of the site (band at 218 bp), CT genotype is heterozygote for the absence and presence of the site (bands at 218-, 187- and 31-bp), and CC genotype is homozygote for the presence of the site (bands at 187- and 31-bp; Figure 1). The 31 bp fragment was invisible in the gel owing to its fast migration speed. The genotypes of CC, CT and TT detected by the PCR-RFLP were also confirmed by sequencing (Figure 2); respectively.
Genotyping of rs5888 SNP in the SCARB1 gene. Lane M, 50 bp marker ladder; lanes 1 and 2, PCR products (218 bp); lanes 3 and 4, TT genotype (218 bp); lanes 7 and 8, CT genotype (218-, 187- and 31-bp); and lanes 5, 6, 9 and 10, CC genotype (187- and 31-bp). The 31 bp fragment was invisible in the gel owing to its fast migration speed.
A part of the nucleotide sequence of rs5888 SNP in the SCARB1 gene. (A) CC genotype; (B) CT genotype; (C) TT genotype.
Genotypic and allelic frequencies
The genotypic and allelic frequencies of SCARB1 rs5888 SNP are shown in Table 2. The frequency of C and T alleles was 71.0% and 29.0% in Mulao, and 73.9% and 26.1% in Han; respectively. The frequency of CC, CT and TT genotypes was 49.3%, 43.5% and 7.2% in Mulao, and 55.2%, 37.4% and 7.4% in Han; respectively. The genotypic frenquency was different between Mulao and Han (P < 0.05), the allelic frequency was also borderline different between Mulao and Han (P = 0.074). There were no significant differences in genotypic and allelic frenquencies between males and females in both ethnic groups.
Genotypes and serum lipid levels
As shown in Tables 3 and 4, the levels of serum HDL-C in Mulao was different among the three genotypes (P < 0.05), the subjects with TT genotype had lower serum HDL-C levels than the subjects with CT or CC genotype in females but not in males. For the Han population, the levels of TG, HDL-C, ApoAI, ApoB and the ratio of ApoAI to ApoB in males were different among the genotypes, the T allele carriers had lower serum HDL-C, ApoAI levels and the ratio of ApoAI to ApoB, and higher serum ApoB levels than the T allele noncarriers. The subjects with TT genotype had higher serum TG levels than the subjects with CC or CT genotype. Multiple linear regression analysis showed that serum HDL-C and ApoAI levels were correlated with genotypes in the combined populations of Mulao and Han (P < 0.05, Table 5). The levels of serum HDL-C and ApoAI in Han and serum HDL-C level in Mulao were correlated with genotypes. Subgroup analysis showed that the levels of HDL-C in Mulao females and Han males were correlated with genotypes (P < 0.05 for each), serum ApoAI, ApoB levels and the ratio of ApoAI to ApoB in Han males were correlated with genotypes.
Genotype-environmental interactions on serum lipid parameters
As shown in Tables 6 and 7, the interaction of genotypes and alcohol consumption on serum TC, TG and ApoB levels in the Han population was detected. The interaction of genotypes and cigarette smoking on serum TC levels was also detected in the Han population. No significant interaction was detected between genotype and alcohol consumption or cigarette smoking on serum lipid levels in the Mulao population.
Comparison of general characteristics and serum lipid levels between Mulao and Han populations.
| Parameter | Mulao | Han Chinese | t (χ2) | P |
|---|---|---|---|---|
| Number | 801 | 807 | - | - |
| Male/female | 379/ 422 | 355/ 452 | 1.792 | 0.181 |
| Age (years) | 52.55±14.78 | 51.40±15.14 | 1.537 | 0.125 |
| Height (cm) | 155.60±8.02 | 155.10±7.85 | 1.265 | 0.206 |
| Weight (kg) | 53.27±9.33 | 54.11±9.36 | -1.809 | 0.071 |
| Body mass index (kg/m2) | 21.94±3.07 | 22.43±3.12 | -3.196 | 0.001 |
| Systolic blood pressure (mmHg) | 130.08±21.57 | 129.35±19.11 | 0.720 | 0.472 |
| Diastolic blood pressure (mmHg) | 81.26±11.55 | 81.91±11.24 | -1.148 | 0.251 |
| Pulse pressure (mmHg) | 48.82±16.19 | 47.44±14.27 | 1.818 | 0.069 |
| Cigarette smoking [n (%)] | ||||
| Nonsmoker | 591 (73.8) | 579(71.8) | ||
| < 20 cigarettes/day | 73 (9.1) | 96 (11.9) | 3.372 | 0.185 |
| ≥ 20 cigarettes/day | 137 (17.1) | 131 (16.3) | ||
| Alcohol consumption [n (%)] | ||||
| Nondrinker | 592 (73.9) | 630 (78.1) | ||
| < 25 g/day | 73 (9.1) | 95 (11.8) | 17.417 | <0.001 |
| ≥ 25 g/day | 136 (17.0) | 82 (10.2) | ||
| Total cholesterol (mmol/L) | 5.07±1.30 | 4.98±1.04 | 1.681 | 0.093 |
| Triglyceride (mmol/L) | 1.09 (0.80) | 1.07 (0.90) | -0.447 | 0.655 |
| HDL-C (mmol/L) | 1.75±0.45 | 1.74±0.53 | 0.331 | 0.756 |
| LDL-C (mmol/L) | 2.95±0.89 | 2.87±0.85 | 1.891 | 0.059 |
| Apolipoprotein (Apo) AI (g/L) | 1.33±0.40 | 1.35±0.27 | -0.978 | 0.328 |
| ApoB (g/L) | 0.99±0.57 | 0.85±0.20 | 6.431 | <0.001 |
| ApoAI/ApoB | 1.60±0.97 | 1.66±0.52 | -1.537 | 0.125 |
HDL-C, high-density lipoprotein cholesterol; LDL-C, low-density lipoprotein cholesterol. The value of triglyceride was presented as median (interquartile range). The difference between the two ethnic groups was determined by the Wilcoxon-Mann-Whitney test.
Comparison of the genotypic and allelic frequencies of the rs5888 SNP between Mulao and Han populations.
| Group | n | Genotype [n (%)] | Allele [n (%)] | |||||
|---|---|---|---|---|---|---|---|---|
| CC | CT | TT | C | T | ||||
| Mulao | 801 | 395 (49.3) | 348 (43.5) | 58 (7.2) | 1138 (71.0) | 464 (29.0) | ||
| Han Chinese | 807 | 445 (55.2) | 302 (37.4) | 60 (7.4) | 1192 (73.9) | 422(26.1) | ||
| χ 2 | - | 6.243 | 3.198 | |||||
| P | - | 0.044 | 0.074 | |||||
| Mulao | ||||||||
| Male | 379 | 191 (50.4) | 161 (42.5) | 27 (7.1) | 543 (71.6) | 215 (28.4) | ||
| Female | 422 | 204(48.3) | 187 (44.3) | 31 (7.3) | 595 (70.5) | 249 (29.5) | ||
| χ 2 | - | 0.339 | 0.251 | |||||
| P | - | 0.844 | 0.616 | |||||
| Han Chinese | ||||||||
| Male | 355 | 193 (54.4) | 140 (39.4) | 22 (6.2) | 526 (74.1) | 184 (25.9) | ||
| Female | 452 | 252 (55.8) | 162 (35.8) | 38 (8.4) | 666 (73.7) | 238 (26.3) | ||
| χ 2 | - | 2.062 | 0.035 | |||||
| P | - | 0.357 | 0.852 | |||||
The genotypes of rs5888 SNP and serum lipid levels in the Mulao population.
| Genotype | n | TC (mmol/L) | TG (mmol/L) | HDL-C (mmol/L) | LDL-C (mmol/L) | ApoAI (g/L) | ApoB (g/L) | ApoAI/ ApoB |
|---|---|---|---|---|---|---|---|---|
| Mulao | ||||||||
| CC | 395 | 5.06±1.17 | 1.07 (0.82) | 1.76±0.40a | 2.92±0.83 | 1.34±0.39 | 0.99±0.57 | 1.59±0.69 |
| CT | 348 | 5.12±1.44 | 1.11 (0.85) | 1.76±0.50a | 2.99±0.93 | 1.34±0.40 | 0.99±0.55 | 1.59±0.84 |
| TT | 58 | 4.97±1.24 | 1.11 (0.76) | 1.61±0.47 | 2.92±1.01 | 1.25±0.40 | 0.98±0.65 | 1.79±2.37 |
| F | - | 0.416 | 1.580 | 3.203 | 0.483 | 1.451 | 0.024 | 1.148 |
| P | - | 0.660 | 0.454 | 0.041 | 0.617 | 0.235 | 0.976 | 0.318 |
| CC | 395 | 5.06±1.17 | 1.07 (0.82) | 1.76±0.40 | 2.92±0.83 | 1.34±0.39 | 0.99±0.57 | 1.59±0.69 |
| CT/TT | 406 | 5.10±1.42 | 1.11(0.82) | 1.74±0.50 | 2.98±0.94 | 1.32±0.40 | 0.99±0.57 | 1.62±1.18 |
| F | - | 0.197 | -0.556 | 0.427 | 0.699 | 0.367 | 0.001 | 0.277 |
| P | - | 0.657 | 0.578 | 0.514 | 0.403 | 0.545 | 0.978 | 0.599 |
| Male | ||||||||
| CC | 191 | 5.19±1.27 | 1.16(0.90) | 1.74±0.39 | 2.93±0.82 | 1.35±0.41 | 1.03±0.61 | 1.52±0.66 |
| CT | 161 | 5.22±1.60 | 1.17 (1.36) | 1.76±0.58 | 2.95±0.86 | 1.37±0.42 | 1.07±0.69 | 1.55±0.71 |
| TT | 27 | 4.99±1.20 | 0.94 (0.99) | 1.67±0.49 | 2.83±0.87 | 1.25±0.47 | 0.99±0.60 | 1.42±0.63 |
| F | - | 0.308 | 3.512 | 0.444 | 0.222 | 0.917 | 0.286 | 0.416 |
| P | - | 0.735 | 0.173 | 0.642 | 0.801 | 0.401 | 0.752 | 0.660 |
| CC | 191 | 5.19±1.27 | 1.16(0.90) | 1.74±0.39 | 2.93±0.82 | 1.35±0.41 | 1.03±0.61 | 1.52±0.66 |
| CT/TT | 188 | 5.19±1.55 | 1.15(1.12) | 1.75±0.57 | 2.93±0.86 | 1.35±0.43 | 1.06±0.66 | 1.53±0.70 |
| F | - | 0.002 | -0.204 | 0.015 | 0.000 | 0.003 | 0.169 | 0.035 |
| P | - | 0.969 | 0.838 | 0.901 | 0.999 | 0.959 | 0.681 | 0.852 |
| Female | ||||||||
| CC | 204 | 4.94±1.07 | 1.01 (0.67) | 1.78±0.40b | 2.93±0.83 | 1.34±0.38 | 0.96±0.54 | 1.65±0.71 |
| CT | 187 | 5.01±1.29c | 1.04 (0.63) | 1.76±0.43a | 3.01±0.99 | 1.31±0.38 | 0.92±0.42 | 1.64±0.94 |
| TT | 31 | 4.96±1.29 | 1.12 (0.69) | 1.56±0.44 | 3.01±1.13 | 1.24±0.34 | 0.98±0.70 | 2.07±3.18 |
| F | - | 0.200 | 0.692 | 3.927 | 0.455 | 0.928 | 0.467 | 1.950 |
| P | - | 0.819 | 0.708 | 0.020 | 0.635 | 0.396 | 0.627 | 0.144 |
| CC | 204 | 4.94±1.07 | 1.01 (0.67) | 1.78±0.40 | 2.93±0.83 | 1.34±0.38 | 0.96±0.54 | 1.65±0.71 |
| CT/TT | 218 | 5.01±1.28 | 1.06(0.64) | 1.73±0.44 | 3.01±1.01 | 1.30±0.37 | 0.92±0.47 | 1.70±1.48 |
| F | - | 0.356 | -0.804 | 1.387 | 0.912 | 1.040 | 0.470 | 0.226 |
| P | - | 0.551 | 0.421 | 0.240 | 0.340 | 0.308 | 0.494 | 0.635 |
TC, total cholesterol; TG, triglyceride; HDL-C, high-density lipoprotein cholesterol; LDL-C, low-density lipoprotein cholesterol; ApoAI, apolipoprotein AI; ApoB, apolipoprotein B
a P < 0.05, b P < 0.01 in comparison with the TT genotype of the same ethnic group
The genotypes of rs5888 SNP and serum lipid levels in the Han population.
| Genotype | n | TC (mmol/L) | TG (mmol/L) | HDL-C (mmol/L) | LDL-C (mmol/L) | ApoAI (g/L) | ApoB (g/L) | ApoAI/ ApoB |
|---|---|---|---|---|---|---|---|---|
| Han Chinese | ||||||||
| CC | 445 | 4.99±1.12 | 1.02 (0.98) | 1.74±0.46a | 2.86±0.84 | 1.36±0.28b | 0.85±0.19 | 1.67±0.50 |
| CT | 302 | 4.97±0.92 | 1.10 (0.86) | 1.77±0.64a | 2.88±0.87 | 1.35±0.26b | 0.86±0.21 | 1.68±0.57 |
| TT | 60 | 4.93±1.08 | 1.09 (0.95) | 1.59±0.34 | 2.84±0.77 | 1.25±0.18 | 0.85±0.19 | 1.52±0.36 |
| F | - | 0.086 | 1.722 | 3.141 | 0.067 | 4.780 | 0.541 | 2.715 |
| P | - | 0.918 | 0.423 | 0.044 | 0.935 | 0.009 | 0.582 | 0.067 |
| CC | 445 | 4.99±1.12 | 1.02 (0.98) | 1.74±0.46 | 2.86±0.84 | 1.36±0.28 | 0.85±0.19 | 1.67±0.50 |
| CT/TT | 362 | 4.96±0.95 | 1.10(0.86) | 1.74±0.60 | 2.87±0.86 | 1.34±0.25 | 0.86±0.21 | 1.65±0.54 |
| F | - | 0.099 | -1.212 | 0.010 | 0.038 | 1.304 | 0.865 | 0.400 |
| P | - | 0.753 | 0.225 | 0.918 | 0.845 | 0.254 | 0.353 | 0.527 |
| Male | ||||||||
| CC | 193 | 5.09±1.15 | 1.14 (1.04) a | 1.75±0.51b | 2.84±0.78 | 1.41±0.33 b | 0.88±0.19 | 1.66±0.48 b |
| CT | 140 | 5.09±0.97 | 1.16 (1.08) a | 1.66±0.35a | 2.99±0.92 | 1.35±0.29 a | 0.93±0.22 | 1.55±0.56 a |
| TT | 22 | 5.34±1.09 | 1.68 (2.61) | 1.46±0.31 | 2.94±0.75 | 1.22±0.18 | 0.94±0.20 | 1.34±0.37 |
| F | - | 0.623 | 7.939 | 5.306 | 1.228 | 4.946 | 2.668 | 5.459 |
| P | - | 0.537 | 0.019 | 0.005 | 0.294 | 0.008 | 0.071 | 0.005 |
| CC | 193 | 5.09±1.15 | 1.14 (1.04) | 1.75±0.51 | 2.84±0.78 | 1.41±0.33 | 0.88±0.19 | 1.66±0.48 |
| CT/TT | 162 | 5.12±0.99 | 1.30(1.14) | 1.64±0.35 | 2.98±0.90 | 1.34±0.28 | 0.93±0.22 | 1.53±0.54 |
| F | - | 0.105 | -1.863 | 6.110 | 2.395 | 5.376 | 5.305 | 7.213 |
| P | - | 0.746 | 0.062 | 0.014 | 0.123 | 0.021 | 0.022 | 0.008 |
| Female | ||||||||
| CC | 252 | 4.90±1.08 | 0.97 (0.86) | 1.75±0.42 | 2.86±0.89 | 1.32±0.23 | 0.82±0.19 | 1.69±0.51 |
| CT | 162 | 4.89±0.87 | 1.04 (0.74) | 1.85±0.80 | 2.81±0.82 | 1.35±0.24 | 0.81±0.18 | 1.77±0.55 |
| TT | 38 | 4.72±1.02 | 1.00 (0.75) | 1.66±0.35 | 2.82±0.79 | 1.26±0.19 | 0.79±0.16 | 1.63±0.32 |
| F | - | 0.610 | 0.311 | 2.212 | 0.213 | 2.454 | 0.669 | 1.683 |
| P | - | 0.544 | 0.856 | 0.111 | 0.808 | 0.087 | 0.513 | 0.187 |
| CC | 252 | 4.90±1.08 | 0.97 (0.86) | 1.75±0.42 | 2.86±0.89 | 1.32±0.23 | 0.82±0.19 | 1.69±0.51 |
| CT/TT | 200 | 4.85±0.90 | 1.03(0.73) | 1.82±0.74 | 2.81±0.81 | 1.33±0.23 | 0.81±0.18 | 1.74±0.52 |
| F | - | 0.228 | -0.064 | 1.502 | 0.421 | 0.277 | 0.741 | 1.060 |
| P | - | 0.634 | 0.949 | 0.221 | 0.517 | 0.599 | 0.390 | 0.304 |
TC, total cholesterol; TG, triglyceride; HDL-C, high-density lipoprotein cholesterol; LDL-C, low-density lipoprotein cholesterol; ApoAI, apolipoprotein AI; ApoB, apolipoprotein B
a P < 0.05, b P < 0.01 in comparison with the TT genotype of the same ethnic group
Correlation between genotypes and serum lipid parameters in the Mulao and Han populations.
| Lipid parameter | Relative factor | Unstandardized coefficient | Std. error | Standardized coefficient | t | P |
|---|---|---|---|---|---|---|
| Mulao and Han | ||||||
| HDL-C | Genotype (CC+CT vs TT) | -0.158 | 0.045 | -0.084 | -3.490 | <0.001 |
| ApoAI | Genotype (CC+CT vs TT) | -0.100 | 0.031 | -0.077 | -3.198 | 0.001 |
| Mulao | ||||||
| HDL-C | Genotype (CC+CT vs TT) | -0.152 | 0.059 | -0.087 | -2.562 | 0.011 |
| Han | ||||||
| HDL-C | Genotype (CC+CT vs TT) | -0.177 | 0.069 | -0.088 | -2.560 | 0.011 |
| ApoAI | Genotype (CC+CT vs TT) | -0.096 | 0.034 | -0.095 | -2.835 | 0.005 |
| Mulao/female | ||||||
| HDL-C | Genotype (CC+CT vs TT) | -0.210 | 0.075 | -0.133 | -2.844 | 0.005 |
| Han/male | ||||||
| HDL-C | Genotype | -0.115 | 0.036 | -0.158 | -3.216 | 0.001 |
| ApoAI | Genotype | -0.068 | 0.024 | -0.136 | -2.816 | 0.005 |
| ApoB | Genotype (CC vs CT+TT) | 0.054 | 0.021 | 0.129 | 2.549 | 0.011 |
| ApoAI/ApoB | Genotype | -0.127 | 0.041 | -0.152 | -3.104 | 0.002 |
TC, total cholesterol; TG, triglyceride; HDL-C, high-density lipoprotein cholesterol; LDL-C, low-density lipoprotein cholesterol; ApoAI, apolipoprotein AI; ApoB, apolipoprotein B
The interaction between genotypes and alcohol consumption on serum lipid levels in the Han population.
| Alcohol | Lipid | CC (n = 445) | CT (n = 302) | TT (n = 60) | P for trend | P for interaction |
|---|---|---|---|---|---|---|
| Nondrinker | 4.91±1.10 | 4.93±0.86 | 4.73±1.03 | 0.423 | ||
| < 25 g/d | TC | 4.96±0.62 | 5.12±0.93 | 6.00±0.92 | 0.002 | 0.016 |
| ≥25 g/d | 5.51±1.52 | 5.28±1.25 | 4.91±0.93 | 0.515 | ||
| Nondrinker | 0.99(0.89) | 1.05(0.79) | 1.01(0.71) | 0.973 | ||
| < 25 g/d | TG | 0.97(1.05) | 1.30(0.99) | 4.26(6.59) | 0.031 | 0.003 |
| ≥25 g/d | 1.18(1.34) | 1.47(1.97) | 1.68(4.98) | 0.182 | ||
| Nondrinker | 0.83±0.19 | 0.85±0.20 | 0.80±0.16 | 0.349 | ||
| < 25 g/d | ApoB | 0.85±0.13 | 0.95±0.19 | 1.11±0.20 | <0.001 | 0.002 |
| ≥25 g/d | 0.95±0.22 | 0.92±0.28 | 0.82±0.12 | 0.483 |
TC, total cholesterol; TG, triglyceride; ApoB, apolipoprotein B; The value of triglyceride was presented as median (interquartile range).
The interaction between genotypes and cigarette smoking on serum lipid levels in the Han population.
| Cigarette smoking | Lipid | CC(n = 445) | CT(n = 302) | TT(n = 60) | P for trend | P for interaction |
|---|---|---|---|---|---|---|
| Nonsmoker | 4.92±1.03 | 5.01±0.93 | 4.67±1.00 | 0.105 | ||
| < 20 cigarettes/d | TC | 4.94±0.82 | 4.81±0.74 | 4.60±0.42 | 0.563 | 0.007 |
| ≥20 cigarettes/d | 5.34±1.53 | 4.97±1.03 | 5.82±0.96 | 0.071 |
Discussion
The present study shows that serum ApoB levels were higher in Mulao than in Han. There were no significant differences in the levels of TC, TG, HDL-C, LDL-C, ApoAI and the ratio of ApoAI to ApoB between the two ethnic groups. It is well known that dyslipidemia is a complex trait caused by both environmental and genetic factors. Mulao is a genetic feature distinctive nationality. The engagements of Mulao nationality were family-arranged in childhood, usually with the girl being four or five years older than the boy. Cousin marriage was very popular. Engagement and marriage were marked by bride-wealth payments. Marriage ceremonies were held when the girl reached puberty. She remained with her natal family until her first child was born. Till then she was free to join the young men and women who came together for responsive singing, flirtations, and courtships at festival times. Divorce and remarriage were permitted, with little restriction. Therefore, we believe that the hereditary characteristics and genotypes of some lipid metabolism-related genes in this population may be different from those in Han Chinese.
The genotypic and allelic frequencies of the rs5888 SNP in diverse racial/ethnic groups are different, which can be found on the International HapMap project (http://hapmap.ncbi.nlm.nih.gov/cgi-perl/gbrowse/hapmap24_B36). The frequency of CC, CT and TT genotypes was 20.7%, 53.4% and 25.9% in CEU (Utah residents with ancestry from northern and western Europe), 76.3%, 23.7% and 0% in YRI (Yoruba in Ibadan, Nigeria), 65.9%, 29.5% and 4.5% in JPT (Japanese in Tokyo, Japan), 57.8%, 35.6% and 6.7% in CHB (Han Chinese in Beijing, China). The frequency of C and T alleles was 47.4% and 52.6% in CEU, 88.1% and 11.9% in YRI, 80.7% and 19.3% in JPT, 75.6% and 24.4% in CHB. In several previous studies, the C allele frequency of rs5888 SNP was also varied from 40% to 83% in diverse ethnic populations [19-27]. In the present study, we showed that there was significant difference in genotypic frequency of SCARB1 rs5888 SNP between the two ethnic groups. The frequency of CC genotype was lower in Mulao than in Han. The frequency of C allele was borderline lower in Mulao than in Han (71.0% vs. 73.9%; P = 0.074). These results suggest that the associations of the rs5888 SNP in the SCARB1 gene and lipid phenotypes may have a racial/ethnic difference.
Several ethnic distinct populations have previously reported the association of rs5888 SNP in the SCARB1 gene and serum lipid levels. There are consistent observations of a protective phenotype of the rs5888 SNP and increased serum HDL-C levels in some studies [19,21,23] but not in others [20,22,27]. In our study, however, we found that the TT genotype was associated with lower serum HDL-C levels in Mulao females and lower serum HDL-C and ApoAI levels in Han males, which was opposite to the results of several provious studies [19,21,23]. The associations of rs5888 genotypes and serum non-HDL-C were also found in several provious studies, but the results are inconsistent. Acton et al. [20] and Osgood et al. [23] showed that the T allele carriers have significant lower LDL-C levels in females. In a research on subjects with heterozygous familial hypercholesterolemia, however, Tai et al. [27] found that the exon 8 (rs5888) SNP was associated with increased levels of TC, very-low-density lipoprotein cholesterol (VLDL-C), LDL-C and TG. Morabia et al. [21] also showed that the subjects with TT genotype had higher levels of TC and LDL-C than the subjects with CC genotype in females. These deleterious effects were similar to the results of our present study. We showed that the levels of TG, ApoB and the ratio of ApoAI to ApoB were different among the genotypes in Han males, the T allele carriers had lower ApoAI/ApoB ratio and higher ApoB levels than the T allele noncarriers. The subjects with TT genotype had higher levels of serum TG than the subjects with CC or CT genotype. But these results were not observed in the Mulao population.
Studies have shown that mice with attenuated expression of SCARB1 display elevated concentrations of LDL-C as well as HDL-C [11,12,17]. In human, a novel 11-base pair deletion mutation in the promoter region of the SCARB1 gene was associated with increased plasma HDL-C levels in Chinese Taiwanese [25]. Furthermore, a recent research reported that the rs5888 variant in the SCARB1 was significantly associated with reduced SCARB1 protein expression and function in vitro [35]. Therefore, our subjects with the rs5888 mutation seem to be similar to the conditions of mice with attenuated expression of SCARB1. So it is easy to understand the rs5888 SNP is associated with higher serum ApoB and TG levels in our study. However, our results for the association of rs5888 and serum HDL-C levels differ from what would be expected on the mice model with attenuated expression of the SCARB1 that showed increased serum HDL-C concentrations. In another large study, Osgood et al. [23] also did not find consistent results with the results in mice model. Although they found that the T allele was associated with increased HDL-C, the T allele was also associated with decreased LDL-C levels, instead of increased LDL-C levels. The reasons for these contradictory results with mice model and the diverse results in different populations are not well known. The possible reasons might include that the rs5888 SNP does not affect the amino acid sequence of the protein, and is in linkage disequilibrium with a functional mutation in the SCARB1 gene, or alternatively with another nearby functional variant at the chromosomal region of 12q24. Furthermore, some studies showed that the SCARB1 rs5888 SNP has age-related effects on cholesterol transport lipoproteins [19,21]. In addition, serum lipid levels were also influenced by many other lipid-related genes [8], environmental factors [36] and gene-environmental interactions [37,38]. Therefore, the diverse results may due to the differences in age, gene-gene, as well as gene-environmental interactions.
Environmental factors and gene-environmental interactions have been shown to play an important role in determining serum lipid levels [36-38]. In the present study, we also found that serum lipid parameters were influenced by the gene-environmental interactions. However, the gene-environmental interactions on serum lipid phenotypes were different between the Mulao and Han population. In the Han population, the interaction of genotypes and alcohol consumption was shown to influence serum TC, TG amd ApoB levels, and the interaction of genotypes and cigarette smoking was detected to influence serum TC levels. But no significant interaction between genotypes and alcohol consumption or cigarette smoking on serum lipid levels was detected in the Mulao population. These findings suggest that the difference in serum ApoB levels between the Mulao and Han population may partly attributed to the differences in SCARB1 genotype-environmental interactions.
Conclusion
The present study shows that the T allele frequency of rs5888 SNP and serum ApoB levels were higher in Mulao than in Han Chinese. The levels of HDL-C were correlated with genotypes in Mulao females, the subjects with TT genotype had lower serum HDL-C levels than the subjects with CT or TT genotype. The levels of TG, HDL-C, ApoA1, ApoB and ApoA1/ApoB ratio in Han were also correlated with genotypes in males, the T allele carriers had lower levels of HDL-C, ApoAI and the ApoAI/ApoB ratio, and higher levels of ApoB than the subjects with T allele noncarriers. The subjects with TT genotype had higher levels of TG than the subjects with CC or CT genotype. Serum lipid parameters were also influenced by gene-environmental interactions in the Han population. These results suggest that the differences in serum ApoB levels between the two enthnic groups might partially attribute to the differences in SCARB1 genotype-environmental interactions.
Acknowledgements
This study was supported by the National Natural Science Foundation of China (No: 30960130).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Lopez AD, Mathers CD, Ezzati M. et al. Global and regional burden of disease and risk factors, 2001: systematic analysis of population health data. Lancet. 2006;367:1747-57
2. Wilson PW. High-density lipoprotein, low-density lipoprotein and coronary artery disease. Am J Cardiol. 1990;66:7A-10A
3. Castelli WP, Garrison RJ, Wilson PW. et al. Incidence of coronary heart disease and lipoprotein cholesterol levels. The Framingham Study. JAMA. 1986;256:2835-8
4. Gordon DJ, Probstfield JL, Garrison RJ. et al. High-density lipoprotein cholesterol and cardiovascular disease. Four prospective American studies. Circulation. 1989;79:8-15
5. Heller DA, de Faire U, Pedersen NL. et al. Genetic and environmental influences on serum lipid levels in twins. N Engl J Med. 1993;328:1150-6
6. Chen SN, Cilingiroglu M, Todd J. et al. Candidate genetic analysis of plasma high-density lipoprotein-cholesterol and severity of coronary atherosclerosis. BMC Med Genet. 2009;10:111
7. Boes E, Coassin S, Kollerits B. et al. Genetic-epidemiological evidence on genes associated with HDL cholesterol levels: a systematic in-depth review. Exp Gerontol. 2008;44:136-60
8. Weissglas-Volkov D, Pajukanta P. Genetic causes of high and low serum HDL-cholesterol. J Lipid Res. 2010;51:2032-57
9. Acton S, Rigotti A, Landschulz KT. et al. Identification of scavenger receptor SR-BI as a high density lipoprotein receptor. Science. 1996;271:518-20
10. Kozarsky KF, Donahee MH, Rigotti A. et al. Overexpression of the HDL receptor SR-BI alters plasma HDL and bile cholesterol levels. Nature. 1997;387:414-7
11. Rigotti A, Trigatti BL, Penman M. et al. A targeted mutation in the murine gene encoding the high density lipoprotein (HDL) receptor scavenger receptor class B type I reveals its key role in HDL metabolism. Proc Natl Acad Sci USA. 1997;94:12610-5
12. Varban ML, Rinninger F, Wang N. et al. Targeted mutation reveals a central role for SR-BI in hepatic selective uptake of high density lipoprotein cholesterol. Proc Natl Acad Sci USA. 1998;95:4619-24
13. Calvo D, Gomez-Coronado D, Lasuncion MA. et al. CLA-1 is an 85-kD plasma membrane glycoprotein that acts as a high-affinity receptor for both native (HDL, LDL, and VLDL) and modified (OxLDL and AcLDL) lipoproteins. Arterioscler Thromb Vasc Biol. 1997;17:2341-9
14. Swarnakar S, Temel RE, Connelly MA. et al. Scavenger receptor class B, type I, mediates selective uptake of low density lipoprotein cholesteryl ester. J Biol Chem. 1999;274:29733-9
15. Wang N, Arai T, Ji Y. et al. Liver-specific overexpression of scavenger receptor BI decreases levels of very low density lipoprotein ApoB, low density lipoprotein ApoB, and high density lipoprotein in transgenic mice. J Biol Chem. 1998;273:32920-6
16. Ueda Y, Royer L, Gong E. et al. Lower plasma levels and accelerated clearance of high density lipoprotein (HDL) and non-HDL cholesterol in scavenger receptor class B type I transgenic mice. J Biol Chem. 1999;274:7165-71
17. Huszar D, Varban ML, Rinninger F. et al. Increased LDL cholesterol and atherosclerosis in LDL receptor-deficient mice with attenuated expression of scavenger receptor B1. Arterioscler Thromb Vasc Biol. 2000;20:1068-73
18. Feitosa ME, Rice T, Borecki IB. et al. Pleiotropic QTL on chromosome 12q23-q24 influences triglyceride and high-density lipoprotein cholesterol levels: the HERITAGE family study. Hum Biol. 2006;78:317-27
19. Roberts CG, Shen H, Mitchell BD. et al. Variants in scavenger receptor class B type I gene are associated with HDL cholesterol levels in younger women. Hum Hered. 2007;64:107-13
20. Acton S, Osgood D, Donoghue M. et al. Association of polymorphisms at the SR-BI gene locus with plasma lipid levels and body mass index in a white population. Arterioscler Thromb Vasc Biol. 1999;19:1734-43
21. Morabia A, Ross BM, Costanza MC. et al. Population-based study of SR-BI genetic variation and lipid profile. Atherosclerosis. 2004;175:159-68
22. McCarthy JJ, Lewitzky S, Reeves C. et al. Polymorphisms of the HDL receptor gene associated with HDL cholesterol levels in diabetic kindred from three populations. Hum Hered. 2003;55:163-70
23. Osgood D, Corella D, Demissie S. et al. Genetic variation at the scavenger receptor class B type I gene locus determines plasma lipoprotein concentrations and particle size and interacts with type 2 diabetes: the framingham study. J Clin Endocrinol Metab. 2003;88:2869-79
24. Hong SH, Kim YR, Yoon YM. et al. Association between HaeIII polymorphism of scavenger receptor class B type I gene and plasma HDL-cholesterol concentration. Ann Clin Biochem. 2002;39:478-81
25. Hsu LA, Ko YL, Wu S. et al. Association between a novel 11-base pair deletion mutation in the promoter region of the scavenger receptor class B type I gene and plasma HDL cholesterol levels in Taiwanese Chinese. Arterioscler Thromb Vasc Biol. 2003;23:1869-74
26. McCarthy JJ, Lehner T, Reeves C. et al. Association of genetic variants in the HDL receptor, SR-B1, with abnormal lipids in women with coronary artery disease. J Med Genet. 2003;40:453-8
27. Tai ES, Adiconis X, Ordovas JM. et al. Polymorphisms at the SRBI locus are associated with lipoprotein levels in subjects with heterozygous familial hypercholesterolemia. Clin Genet. 2003;63:53-8
28. Xu L, Deng QY, Li SF. et al. Genetic analysis of Mulao nationality using 15 short tandem repeats. Zhonghua Yi Xue Yi Chuan Xue Za Zhi. 2008;25:96-100
29. Wu DF, Yin RX, Hu XJ. et al. Association of rs5888 SNP in the scavenger receptor class B type 1 gene and serum lipid levels. Lipids Health Dis. 2012;11:50
30. Li Q, Yin RX, Yan TT. et al. Association of the GALNT2 gene polymorphisms and several environmental factors with serum lipid levels in the Mulao and Han populations. Lipids Health Dis. 2011;10:160
31. Yan TT, Yin RX, Li Q. et al. Sex-specific association of rs16996148 SNP in the NCAN/CILP2/PBX4 and serum lipid levels in the Mulao and Han populations. Lipids Health Dis. 2011;10:248
32. People's Republic of China--United States Cardiovascular and Cardiopulmonary Epidemiology Research Group. An epidemiological study of cardiovascular and cardiopulmonary disease risk factors in four populations in the People's Republic of China. Baseline report from the P.R.C.-U.S.A. Collaborative Study. Circulation. 1992;85:1083-96
33. Ruixing Y, Weixiong L, Hanjun Y. et al. Diet, lifestyle, and blood pressure of the middle-aged and elderly in the Guangxi Bai Ku Yao and Han populations. Am J Hypertens. 2008;21:382-7
34. Zhou B. Effect of body mass index on all-cause mortality and incidence of cardiovas-cular diseases—report for meta-analysis of prospective studies open optimal cut-off points of body mass index in Chinese adults. Biomed Environ Sci. 2002;15:245-52
35. Constantineau J, Greason E, West M. et al. A synonymous variant in scavenger receptor, class B, type I gene is associated with lower SR-BI protein expression and function. Atherosclerosis. 2010;210:177-82
36. Ruixing Y, Yuming C, Shangling P. et al. Effects of demographic, dietary and other lifestyle factors on the prevalence of hyperlipidemia in Guangxi Hei Yi Zhuang and Han populations. Eur J Cardiovasc Prev Rehabil. 2006;13:977-84
37. Ruixing Y, Yiyang L, Meng L. et al. Interactions of the apolipoprotein C-III 3238C>G polymorphism and alcohol consumption on serum triglyceride levels. Lipids Health Dis. 2010;9:86
38. Liu WY, Yin RX, Zhang L. et al. Interactions of the LIPG 584C>T polymorphism and alcohol consumption on serum lipid levels. Alcohol. 2011;45:681-7
Author contact
![]() Corresponding author: Prof. Rui-Xing Yin, Department of Cardiology, Institute of Cardiovascular Diseases, the First Affiliated Hospital, Guangxi Medical University, 22 Shuangyong Road, Nanning 530021, Guangxi, People's Republic of China. Tel: +86-771-5326125; Fax: +86-771-5353342; e-mail: yinruixingcom.cn.
Corresponding author: Prof. Rui-Xing Yin, Department of Cardiology, Institute of Cardiovascular Diseases, the First Affiliated Hospital, Guangxi Medical University, 22 Shuangyong Road, Nanning 530021, Guangxi, People's Republic of China. Tel: +86-771-5326125; Fax: +86-771-5353342; e-mail: yinruixingcom.cn.

 Global reach, higher impact
Global reach, higher impact