Impact Factor ISSN: 1449-1907
Int J Med Sci 2011; 8(6):492-500. doi:10.7150/ijms.8.492 This issue Cite
Research Paper
Translational Medicine and Reliability of Single-Nucleotide Polymorphism Studies: Can We Believe in SNP Reports or Not?
1. Onkologkliniken Sörmland, Mälarsjukhuset, Eskilstuna, Sweden
2. PACMeR, department of. Medical Oncology, Athens, Greece
3. Department of Medical Oncology, General Hospital of Lamia, Greece
4. Centre for Reproductive Medicine, Free University of Brussels, Brussels, Belgium
5. PACMeR, associate researcher, dept. Internal Medicine, Athens, Greece
6. Department of Obstetrics and Gynaecology, University General Hospital of Larisa, Greece
Received 2011-5-29; Accepted 2011-8-9; Published 2011-8-24
Abstract
Background: The number of genetic association studies is increasing exponentially. Nonetheless, genetic association reports are prone to potential biases which may influence the reported outcome.
Aim: We hypothesized that positive outcome for a determined polymorphism might be over-reported across genetic association studies analysing a small number of polymorphisms, when compared to studies analysing the same polymorphism together with a high number of other polymorphisms.
Methods: We systematically reviewed published reports on the association of glutathione s-transferase (GST) single-nucleotide polymorphisms (SNPs) and cancer outcome.
Result: We identified 79 eligible trials. Most of the studies examined the GSTM1, theGSTP1 Ile105Val mutation, and GSTT1polymorphisms (n = 54, 57 and 46, respectively). Studies analysing one to three polymorphisms (n = 39) were significantly more likely to present positive outcomes, compared to studies examining more than 3 polymorphisms (n=40) p = 0.004; this was particularly evident for studies analysing the GSTM1polymorphism (p =0.001). We found no significant associations between journal impact factor, number of citations, and probability of publishing positive studies or studies with 1-3 polymorphisms examined.
Conclusions: We propose a new subtype of publication bias in genetic association studies. Positive results for genetic association studies analysing a small number of polymorphisms (n = 1-3) should be evaluated extremely cautiously, because a very large number of such studies are inconclusive and statistically under-powered. Indeed, publication of misleading reports may affect harmfully medical decision-making and use of resources, both in clinical and pharmacological development setting.
Keywords: single-nucleotide polymorphisms, genetic association studies, publication-bias, literature bias, translational research.
Introduction
Genetic association studies investigate the relationship between gene polymorphisms and risk of disease or treatment outcome. Furthermore, due to advances in molecular targeted treatment technologies and the continual expansion of translational research, genetic association studies play a key role in the development of new therapeutical targets. For these reasons the number of genetic association studies is increasing rapidly and this trend is expected to accelerate due to the availability of mapped single-nucleotide polymorphisms (SNP) in the human genome and advances in genotyping technologies [1].
However, despite the number of genetic association studies being expected to grow exponentially over the next decade, no clear criteria are available to assess the credibility of these reports. Are all statistical significant medical reports, on SNP studies, reliable enough to drive firm conclusions and trig clinical/therapeutical applications?
Positive-outcome (also known as “publication”) bias refers to the greater likelihood of a study with positive results to be published compared with studies with negative results [2,3]. Publication bias (“false-positive” reporting) is a particular threat to the credibility of the literature, including genetic association studies, since it may affect decision-making both in clinical and pharmacological development settings.
Biologists, researchers and physicians are actually called to deal with manuscripts of translational medicine research in their daily life. However, no parameters are actually available to orient them in a correct interpretation of potential misleading sources of literature-bias.
Based on the over mentioned reflections and considering the following three facts: 1) reviewer's and editor's decision about publication of manuscripts are influenced by positive findings [2,3,4]; 2) positive studies are more possible to be published in journals with higher impact factor (IF) [3,5] and may be cited more often than negative studies [6,7]; 3) null papers are typically given low publication priority scores and may not be accepted for publication [2], we hypothesized that the pressure for publication among authors and the fierce competition for acceptance in leading journals [3,4] may lead authors firstly, to perform studies with few polymorphisms, which are less expensive, need less time to complete and secondly, to submit for publication only those studies with positive outcome.
Is it the case? If yes, what about the impact of this phenomenon on medical literature? How positive compared to negative reports correlate with publication differences in impact factor journals or citation frequency?
In our study, we thereafter tested the hypothesis that a positive outcome for a determined polymorphism might be over reported across genetic association studies analysing a small number of polymorphisms, when compared to studies analysing the same polymorphism together with a high number of other polymorphisms. We also tried to assess any differences in journal impact factor or citation frequency among positive versus negative reports.
Due to the high number of published reports on the association of GST polymorphisms and cancer outcome (mainly colorectal, breast, and lung malignancy) SNPs for GST polymorphisms were used as a substrate for analyses.
Materials and Methods
Search strategy and eligibility criteria
We electronically searched the PubMed medical literature database and ISI Web of Science from inception to June 1, 2009, without language restrictions, using the following keyword combinations: (glutathione s-transferase OR GST OR GSTT1 OR GSTM1 OR GSTP1) AND (polymorphism OR polymorphisms OR SNP OR mutation) AND (cancer OR malign* OR carcinoma OR tumor OR tumour). The electronic search was supplemented by a manual review of the references of included studies.
The studies selected for our analysis had to meet the following criteria: (a) investigate the association between at least of one GST SNP: and cancer outcome; and (b) include only patients with solid tumours.
We excluded case-control studies that examined the role of GST polymorphisms on cancer risk, studies that included patients with hematologic malignancies and studies that investigated the role of GST polymorphisms on pharmacokinetics of specific drugs.
Two investigators independently reviewed all potentially relevant articles to determine whether an article met the inclusion criteria, and disagreement was resolved by discussion between the investigators.
Data extraction
We abstracted the following information from eligible trials: authors' name, year of publication, country of origin, type of cancer, sample size, number of polymorphisms tested and results of the study.
Studies were divided into two categories based on the results reported: positive or negative study. Since there is no standardized definition of positive results [8], the following definitions for positive and negative studies were used in our study:
A study was defined as “positive” if it reported any statistical significant difference for any of the GST polymorphisms for at least one of the following outcome measures: overall survival or disease recurrence or response to treatment. In the case of lack of a clear definition, or threshold, for statistical significant difference, we defined “significance” as the presence of a P-value of <0.05 or another effect metric with 95% confidence interval (C.I.) that fell entirely on one side of the null. A study was defined as negative if there was no statistically significant difference detected between a determined GST polymorphisms and any of these outcomes.
Regarding number of polymorphisms examined, eligible studies were divided into two groups: those which examined the association between 1-3 polymorphisms and cancer outcome and those which examined the association of more than 3 polymorphisms.
The Impact Factor (IF) of each journal was extracted from Journal Citation Reports (Institute for Scientific Information, JCR-ISI) [9]. When a journal was not included in the citation index, we set 0 as IF.
The number of citations was obtained though the Science Citation Index [10]. For each published article, all citations of that article from publication to the time of the search were identified. The number of citations per year from the year of publication to the study period was calculated for each article (citations per year).
Statistical analysis
Associations were tested using the chi-square statistic or Fisher's exact test with significance set at P < 0.05. The null hypothesis is that there is no difference in the proportions of positive and negative studies analysing a determined polymorphisms between the studies examining it within 1-3 polymorphisms versus studies examining the same polymorphism among more than 3 polymorphism.
Since the distribution of IF and citation frequency were not normal (Shapiro-Wilk test < 0.05) [11], we used nonparametric tests (Mann-Whitney tests) [12] to study the difference in IF and in frequency of citations per year between groups.
To better examine the possibility of a bias for positive results in studies examined 1-3 polymorphisms, logistic regression analysis, with adjustment for sample size, was used to calculate the odds ratio (OR) of reporting positive results in 2 study groups.
All statistical analyses were done using the SPSS software (SPSS Inc., Chicago, IL, USA, version 11.5). All tests were two-sided with a significance level of 0.05.
Results
Description of studies
A total of 4695 studies were identified from the combined searches. We scanned titles and abstracts for mention of GST polymorphisms associated with cancer outcome in either the title or the abstract. We retrieved 121 potentially eligible articles in full text [Figure 1].
Flowchart diagram of study selection.
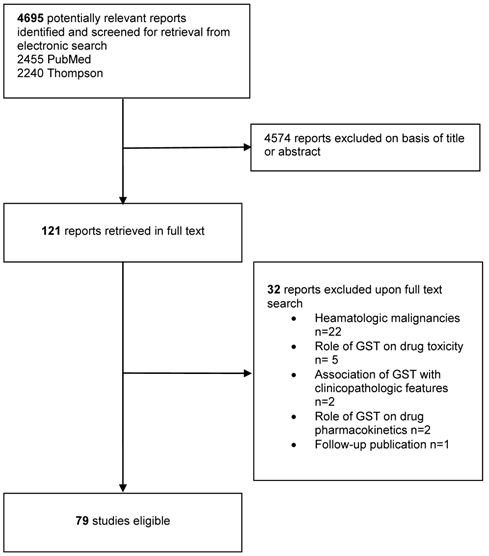
A total of 79 articles that fulfilled the inclusion criteria were found [13-91]. A total of 10 GST polymorphisms were analysed: GSTM1, GSTP1 Ile105Val, GSTP1 Ala114Val, GSTP1 Thr110Ser, GSTP1 Asp147Tyr, GSTT1, GSTM3, GSTA1, GSTO1, GSTO2. Of those, 39 examined 1-3 polymorphisms [13,14,16,19,20,23,24,26-28,31,36,37,39-41,45,46,48,50,53,56,60-62,64,65,68,69,72-74,76-79,81,89,91] and 40 more than 3 polymorphisms [15,17,18,21,22,25,29,30,32-35,38,42-44,47,49,51,52,54,55,57-59,63,66,67,70,71,75,80,82-88,90]. Fifty four studies examined the GSTM1 polymorphism [13,14,16,17,19-27,30-33,35,37,38,43-46,48,49,51-63,65,68,69,70,72-75,77,78,82,83,85-87], 57 the GSTP1 Ile105Val polymorphism [13-16,18,21,22,26,28-30,32-49,51-55,57-61,64,66,67,71,74,76-85,87-90], 46 the GSTT1 polymorphism [13,14,16,17,19, 21-23,25,27,32,33,35,37,38,43-46,48-63,65,70,72,74,75,77,78,82,83,85,87]. Only a small number of reports were available for other polymorphisms: GSTP1 Ala114Val (n=7) [15,32,40,42,66,80,85], GSTP1 Thr110Ser (n=1) [15], GSTP1 Asp147Tyr (n=1) [15], GSTM3 (n=2) [26,63], GSTA1 (n=1) [91], GSTO1 (n=1) [38], GSTO2 (n=1) [44].
Single studies characteristics for each of the 79 eligible studies are reported in the appendix Table, while general characteristics for the eligible studies are summarized in Table 1.
Characteristics of eligible genetic association studies
| Characteristic | No of studies (%) |
|---|---|
| Country of origin | |
| USA | 21 (26.5) |
| United Kingdom | 10 (13) |
| Germany | 6 (8) |
| India | 5 (6) |
| South Korea | 5 (6) |
| Other | 32 (40.5) |
| Type of cancer | |
| Breast | 15 (19) |
| Colorectal | 14 (18) |
| Lung | 12 (15) |
| Ovarian | 10 (13) |
| Other | 28 (35) |
| Sample size | |
| < 50 patients | 6 (7.5) |
| 50-150 patients | 33 (42) |
| 150-250 patients | 18 (23) |
| 250-500 patients | 17 (21.5) |
| 500-1000 patients | 2 (2.5) |
| > 1000 patients | 3 (3.5) |
| No of polymorphisms examined | |
| 1-3 polymorphisms | 39 (49) |
| > 3 polymorphisms | 40 (51) |
| Type of GST examined | |
| GSTM1 present/null | 54 (68) |
| GSTT1 present/null | 46 (58) |
| GSTP1 Ile105Val | 57 (72) |
| GSTP1 Ala114Val | 7 (9) |
| GSTM3 A*/A* or A*/B* or B*/B* | 2 (2.5) |
| GSTA1 A*/A* or A*/B* or B*/B* | 1 (1) |
| GSTP1 Thr110Ser | 1 (1) |
| GSTP1 Asp147Tyr | 1 (1) |
| GSTO1 Ala140Asp | 1 (1) |
| GSTO2 Asn142Asp | 1 (1) |
Association between the outcome of studies and number of polymorphisms tested
When a given polymorphism was analysed, studies reporting 1-3 polymorphisms were significantly more likely to present positive outcomes (n= 29; 74%) compared to studies evaluating the polymorphism across more than 3 polymorphisms (n= 17; 42.5%) (P-value = 0.004); this was particularly evident for studies analysing GSTM1 polymorphism (n= 13 vs. 2, P-value = 0.001), but it does not reach statistical significant differences for studies analysing GSTT1 and GSTP1 polymorphisms (P-value = 0.685 and 0.147 respectively) [Table 2].
Outcome of eligible studies (positive-negative) according to number of polymorphisms tested
| Polymorphisms | No of studies (%) | P-value | |
|---|---|---|---|
| Positive outcome (%) | Negative outcome (%) | ||
| any GST analysed | |||
| 1-3 polymorphisms tested | 29 (74) | 10 (26) | 0.004 |
| > 3 polymorphisms tested | 17 (42.5) | 23 (57.5) | |
| GSTM1 present/null | |||
| 1-3 polymorphisms tested | 13 (48) | 14 (52) | 0.001 |
| > 3 polymorphisms tested | 2 (7) | 25 (93) | |
| GSTT1 present/null | |||
| 1-3 polymorphisms tested | 4 (19) | 17 (81) | 0.685 |
| > 3 polymorphisms tested | 6 (24) | 19 (76) | |
| GSTP1 Ile105Val | |||
| 1-3 polymorphisms tested > 3 polymorphisms tested | 11 (46) 9 (27) | 13 (54) 24 (73) | 0.147 |
| GSTP1 Ala114Val | |||
| 1-3 polymorphisms tested | 1 (100) | 0 (0) | 0.286 |
| > 3 polymorphisms tested | 1 (17) | 5 (83) | |
Logistic regression analysis for studies examined any GST polymorphism revealed that the OR for positive outcome, when comparing studies with 1-3 polymorphisms tested to studies with more than 3 polymorphisms tested, was 3.906 (95% CI, 1.506 to 10.204, P-value = 0.005) after adjustment for sample size.
Association of outcome of studies, IF and citation frequency
There were no significant associations between the impact factor (range: 0.0 - 17.157) and positive studies or studies (P-value = 0.415) with 1-3 polymorphisms examined (P-value = 0.341) [Table 3].
Impact factor and Citations per Year in studies regarding outcome and number of polymorphisms
| Impact Factor | Citations Per Year | |||||
|---|---|---|---|---|---|---|
| Mean (+/- SD) | Median (range) | P-value | Mean +/- SD | Median (range) | P-value | |
| Any GST tested | ||||||
| Positive outcome | 4.848 (4.170) | 4.154 (0.0 - 17.157) | 3.62 (4.24) | 2.00 (0 - 17.33) | ||
| Negative outcome | 5.099 (5.431) | 3.508 (0.0 - 17.157) | 0.415 | 2.40 (3.14) | 1.17 (0 - 11.67) | 0.185 |
| 1-3 polymorphisms tested | 3.855 (2.743) | 3.551 (0.0 - 14.933) | 2.66 (3.22) | 1.67 (0 -16.375) | ||
| > 3 polymorphisms tested | 6.023 (5.879) | 3.508 (0.0 - 17.157) | 0.341 | 3.50 (4.31) | 1.70 (0 - 17.33) | 0.986 |
| GSTM1 present/null | ||||||
| Positive outcome | 3.260 (2.421) | 2.919 (0.0 - 7.514) | 2.89 (2.65) | 2.00 (0 - 8.33) | ||
| Negative outcome | 5.212 (5.335) | 3.508 (0.0 - 17.157) | 0.369 | 3.32 (4.42) | 1.67 (0 - 17.33) | 0.736 |
| 1-3 polymorphisms tested | 3.777 (3.138) | 2.970 (0.0 - 14.933) | 2.87 (3.61) | 1.67 (0 - 16.375) | ||
| > 3 polymorphisms tested | 5.563 (5.889) | 2.970 (0.0 - 17.157) | 0.522 | 3.51 (4.37) | 2.00 (0 - 17.33) | 0.893 |
| GSTT1 present/null | ||||||
| Positive outcome | 5.224 (4.816) | 3.883 (0.0 - 17.157) | 2.39 (2.85) | 1.14 (0 - 8.33) | ||
| Negative outcome | 4.906 (5.138) | 3.289 (0.0 - 17.157) | 0.454 | 3.80 (4.54) | 1.95 (0 - 17.33) | 0.343 |
| 1-3 polymorphisms tested | 3.876 (3.368) | 3.069 (0.0 - 14.933) | 3.17 (3.99) | 1.67 (0 - 16.375) | ||
| > 3 polymorphisms tested | 5.899 (5.986) | 3.508 (0.0 - 17.157) | 0.420 | 3.74 (4.48) | 2.40 (0 - 17.33) | 0.854 |
| GSTP1 Ile105Val | ||||||
| Positive outcome | 5.472 (3.966) | 4.846 (1.932 - 17.157) | 4.52 (5.44) | 2.04 (0 - 17.33) | ||
| Negative outcome | 5.107 (5.117) | 3.508 (0.0 - 17.157) | 0.162 | 2.71 (3.43) | 1.38 (0 - 11.67) | 0.238 |
| 1-3 polymorphisms tested | 4.078 (3.010) | 3.738 (0.0 -14.933) | 3.05 (3.69) | 1.78 (0 - 16.375) | ||
| > 3 polymorphisms tested | 6.077 (5.532) | 3.551 (0,843 - 17.157) | 0.352 | 3.51 (4.64) | 1.25 (0 - 17.33) | 0.461 |
We failed to retrieve information about citation frequency from 8 studies [59,61,62,63,73,74,79,89] . The median citations per year for the remaining 71 studies was 1.67 (range: 0 - 17.33). Citations per year were not significantly associated with either the study outcome (P-value = 0.185) or the number of polymorphisms tested (P-value = 0.986) [Table 3].
Discussion
To our knowledge, this is the first study examined the potential role of number of polymorphisms tested on publication bias. We found that the positive outcome for a given polymorphism might be over reported across genetic association studies analysing a small number of polymorphisms (n = 1-3) when compared to studies analysing the same polymorphism within a higher number of polymorphisms. This was particularly evident for GSTM1 polymorphism. We, therefore, propose a new subtype of publication bias in genetic association studies regarding the number of polymorphisms tested. Thereafter positive results for genetic association studies analysing a small number of polymorphisms (n = 1-3) should be evaluated cautiously and considered at a lower level of evidence.
There are several possible explanations for this finding. First, the pressure for publication among authors and the competition for acceptance in journals are fierce [3,4] and may lead the authors to perform studies with few polymorphisms, which need less time to complete, and to submit for publication only those with positive outcome. Researchers are generally more enthusiastic about projects that have positive results and are more likely to complete them and submit them for publication [4]. On the other hand, authors of studies with negative results are disappointed due to the feeling that null papers are typically given low publication priority scores and may not be accepted for publication [92]. Moreover, it is possible that reviewer's and editor's decision about publication are influenced by positive findings [2,3,4].
An additional potential explanation for publication bias is that positive studies are more possible to be published in journals with higher IF [3,5]. Nonetheless , when we investigated this theory by comparing the IF from journals published positive studies versus those published negative we found no significant association between IF and positive studies. We further tried to investigate the magnitude of the reported publication bias on scientific knowledge. Since there is no way to measure the impact of published articles on medical knowledge we may estimate their impact indirectly by calculating how frequently other authors cite them. Our results are encouraging since we found that studies with positive results or with 1-3 polymorphisms were not cited more frequently compared with studies with negative results or with > 3 polymorphisms. In literature, there are controversies concerning the citation frequency among positive or negative studies. Previous studies in other medical fields demonstrated that trials with a positive outcome were cited significantly more often than trials with a negative outcome [6,7] while other studies find no association [93].
We further documented that studies analysing a low number or a higher number of polymorphisms has the same probability to be published in impact factor journals.
Biologists, researchers and physicians are actually called to deal with translational medicine research manuscripts in their daily life. However, no parameters are actually available to orient them in a correct interpretation of potential misleading sources of literature-bias. The existence of such type of bias in genetic association studies might lead to incorrect conclusions about the usefulness of certain polymorphisms as prognostic genetic markers. It may also have direct impact in medical research by guiding researchers and funding sources in investigating insignificant genes. It is, therefore, extremely important to minimize this bias in medical literature since it may lead to severe decision-making consequences both in clinical and pharmacological development settings. We, therefore, propose that researchers should perform studies that examine many polymorphisms; however, for researchers who investigate one or a few SNPs, they should publish their study regardless of the outcome. It should be emphasized, however, that the transition from single SNP studies to genome-wide association studies (GWAS)--of cohorts sufficiently large in size so as to guarantee ample statistical power--represents a new era in human genetics, which has now arrived and offers an opportunity to overcome biases related to under-powered SNP studies on cohorts that are too small. There are several drawbacks in the study which should be discussed. First, one can oppose that there is problem in generalizing these data to all genetic association studies. This is correct. Anyhow, since our study outcomes were pre-specified and include all available evidence (79 studies) in this topic, our data are extremely possible to be solid enough to support our hypotheses. Confirmatory studies on polymorphisms of other genes are anyhow needed to depict the strength of our hypothesis. Second, one could argue that the observed higher rate of positive outcome in studies with small numbers of polymorphisms might reflect the investigation of SNPs that have been found to be “positive” in previous studies. Moreover, because several thresholds for the definition of statistical significance on genetic association studies have been proposed [94] whereas no consensus has been reached, there is the risk that some of these studies could be misclassified as “positive.
Conclusion: publication bias due to the number of polymorphism tested is a potential threat in medical literature. Positive results for genetic association studies analysing a small number of polymorphisms (n = 1-3) should be evaluated cautiously and considered at a lower level of evidence. Biologists, researchers and physicians dealing with translational medicine research should be aware of this potential threat for “false-positive” reports.
Conflict of Interest
There is no conflict of interest and no financial interest to declare.
References
1. Ioannidis JP. Genetic associations: False or true? Trends Mol Med. 2003;9:135-138
2. Dickersin K, Min YI, Meinert CL. Factors influencing publication of research results. Follow-up of applications submitted to two institutional review boards. JAMA. 1992;267:374-8
3. Easterbrook PJ, Berlin JA, Gopalan R. et al. Publication bias in clinical research. Lancet. 1991;337:867-72
4. Begg CB, Berlin JA. Publication bias: a problem in interpreting medical data. J R Stat SOC A. 1988;151:419-63
5. Littner Y, Mimouni FB, Dollberg S, Mandel D. Negative results and impact factor: A lesson from neonatology. Arch Pediatr Adolesc Med. 2005;159:1036-1037
6. Ravnskov U. Cholesterol lowering trials in coronary heart disease: frequency of citation and outcome. BMJ. 1992;305:15-9
7. Kjaergard LL, Gluud C. Citation bias of hepato-biliary randomized clinical trials. Journal of Clinical Epidemiology. 2002;55:407-410
8. Olson CM. Publication bias. Acad Emerg Med. 1994;1:207-209
9. Thomson Reuters. http://www.isiknowledge.com/jcr
10. Thomson Reuters. http://www.isiknowledge.com
11. Shapiro SS, Wilk MB. "An analysis of variance test for normality (complete samples)". Biometrika. 1965;52(3-4):591-611
12. Mann HB, Whitney DR. On a test of whether one of two random variables is stochastically larger than the other. Annals of Mathematical Statistics. 1947;18:50-60
13. Agalliu I, Lin DW, Salinas CA, Feng Z, Stanford JL. Polymorphisms in the glutathione S-transferase M1, T1, and P1 genes and prostate cancer prognosis. Prostate. 2006;66:1535-41
14. Beeghly A, Katsaros D, Chen H, Fracchioli S, Zhang Y, Massobrio M, Risch H, Jones B, Yu H. Glutathione S-transferase polymorphisms and ovarian cancer treatment and survival. Gynecol Oncol. 2006;100:330-7
15. Booten R, Ward T, Heighway J, Ashcroft L, Morris J, Thatcher N. Glutathione-S-transferase P1 isoenzyme polymorphisms, platinum-based chemotherapy, and non-small cell lung cancer. J Thorac Oncol. 2006;1:679-83
16. Cabelguenne A, Loriot MA, Stucker I, Blons H, Koum-Besson E, Brasnu D, Beaune P, Laccourreye O, Laurent-Puig P, De Waziers I. Glutathione-associated enzymes in head and neck squamous cell carcinoma and response to cisplatin-based neoadjuvant chemotherapy. Int J Cancer. 2001;93:725-30
17. Chacko P, Joseph T, Mathew BS, Rajan B, Pillai MR. Role of xenobiotic metabolizing gene polymorphisms in breast cancer susceptibility and treatment outcome. Mutat Res. 2005;581:153-63
18. Chung HH, Kim MK, Kim JW, Park NH, Song YS, Kang SB, Lee HP. XRCC1 R399Q polymorphism is associated with response to platinum-based neoadjuvant chemotherapy in bulky cervical cancer. Gynecol Oncol. 2006;103:1031-7
19. Csejtei A, Tibold A, Varga Z, Koltai K, Ember A, Orsos Z, Feher G, Horvath OP, Ember I, Kiss I. GSTM, GSTT and p53 polymorphisms as modifiers of clinical outcome in colorectal cancer. Anticancer Res. 2008;28:1917-22
20. Ge H, Lam WK, Lee J, Wong MP, Yew WW, Lung ML. Analysis of L-myc and GSTM1 genotypes in Chinese non-small cell lung carcinoma patients. Lung Cancer. 1996;15:355-66
21. Geisler SA, Olshan AF, Cai J, Weissler M, Smith J, Bell D. Glutathione S-transferase polymorphisms and survival from head and neck cancer. Head Neck. 2005;27:232-42
22. Goekkurt E, Hoehn S, Wolschke C, Wittmer C, Stueber C, Hossfeld DK, Stoehlmacher J. Polymorphisms of glutathione S-transferases (GST) and thymidylate synthase (TS)--novel predictors for response and survival in gastric cancer patients. Br J Cancer. 2006;94:281-6
23. Gonlugur U, Pinarbasi H, Gonlugur TE, Silig Y. The association between polymorphisms in glutathione S-transferase (GSTM1 and GSTT1) and lung cancer outcome. Cancer Invest. 2006;24:497-501
24. Goto I, Yoneda S, Yamamoto M, Kawajiri K. Prognostic significance of germ line polymorphisms of the CYP1A1 and glutathione S-transferase genes in patients with non-small cell lung cancer. Cancer Res. 1996;56:3725-30
25. Haque AK, Au W, Cajas-Salazar N, Khan S, Ginzel AW, Jones DV, Zwischenberger JB, Xie J. CYP2E1 polymorphism, cigarette smoking, p53 expression, and survival in non-small cell lung cancer: a long term follow-up study. Appl Immunohistochem Mol Morphol. 2004;12:315-22
26. Holley SL, Rajagopal R, Hoban PR, Deakin M, Fawole AS, Elder JB, Elder J, Smith V, Strange RC, Fryer AA. Polymorphisms in the glutathione S-transferase mu cluster are associated with tumour progression and patient outcome in colorectal cancer. Int J Oncol. 2006;28:231-6
27. Howells RE, Holland T, Dhar KK, Redman CW, Hand P, Hoban PR, Jones PW, Fryer AA, Strange RC. Glutathione S-transferase GSTM1 and GSTT1 genotypes in ovarian cancer: association with p53 expression and survival. Int J Gynecol Cancer. 2001;11:107-12
28. Howells RE, Dhar KK, Hoban PR, Jones PW, Fryer AA, Redman CW, Strange RC. Association between glutathione-S-transferase GSTP1 genotypes, GSTP1 over-expression, and outcome in epithelial ovarian cancer. Int J Gynecol Cancer. 2004;14:242-50
29. Huang MY, Fang WY, Lee SC, Cheng TL, Wang JY, Lin SR. ERCC2 2251A>C genetic polymorphism was highly correlated with early relapse in high-risk stage II and stage III colorectal cancer patients: a preliminary study. BMC Cancer. 2008;8:50
30. Huang ZH, Hua D, Du X. Polymorphisms in p53, GSTP1 and XRCC1 predict relapse and survival of gastric cancer patients treated with oxaliplatin-based adjuvant chemotherapy. Cancer Chemother Pharmacol. 2009 In press
31. Jatoi A, Martenson JA, Foster NR, McLeod HL, Lair BS, Nichols F, Tschetter LK, Moore DF Jr, Fitch TR, Alberts SR; North Central Cancer Treatment Group (N0044). Paclitaxel, carboplatin, 5-fluorouracil, and radiation for locally advanced esophageal cancer: phase II results of preliminary pharmacologic and molecular efforts to mitigate toxicity and predict outcomes: North Central Cancer Treatment Group (N0044). Am J Clin Oncol. 2007;30:507-13
32. Jiao L, Bondy ML, Hassan MM, Chang DZ, Abbruzzese JL, Evans DB, Smolensky MH, Li D. Glutathione S-transferase gene polymorphisms and risk and survival of pancreatic cancer. Cancer. 2007Mar1;109(5):840-8
33. Kim EJ, Jeong P, Quan C, Kim J, Bae SC, Yoon SJ, Kang JW, Lee SC, Jun Wee J, Kim WJ. Genotypes of TNF-alpha, VEGF, hOGG1, GSTM1, and GSTT1: useful determinants for clinical outcome of bladder cancer. Urology. 2005;65:70-5
34. Kim K, Kang SB, Chung HH, Kim JW, Park NH, Song YS. XRCC1 Arginine194Tryptophan and GGH-401Cytosine/Thymine polymorphisms are associated with response to platinum-based neoadjuvant chemotherapy in cervical cancer. Gynecol Oncol. 2008;111:509-15
35. Kim HS, Kim MK, Chung HH, Kim JW, Park NH, Song YS, Kang SB. Genetic polymorphisms affecting clinical outcomes in epithelial ovarian cancer patients treated with taxanes and platinum compounds: a Korean population-based study. Gynecol Oncol. 2009;113:264-9
36. Kweekel DM, Gelderblom H, Antonini NF, Van der Straaten T, Nortier JW, Punt CJ, Guchelaar HJ. Glutathione-S-transferase pi (GSTP1) codon 105 polymorphism is not associated with oxaliplatin efficacy or toxicity in advanced colorectalcancer patients. Eur J Cancer. 2009;45:572-8
37. Lee JM, Wu MT, Lee YC, Yang SY, Chen JS, Hsu HH, Huang PM, Kuo SW, Lee CJ, Chen CJ. Association of GSTP1 polymorphism and survival for esophageal cancer. Clin Cancer Res. 2005;11:4749-53
38. Lima MM Jr, Oliveira MN, Granja F, Trindade AC, De Castro Santos LE, Ward LS. Lack of association of GSTT1, GSTM1, GSTO1, GSTP1 and CYP1A1 polymorphisms for susceptibility and outcome in Brazilian prostate cancer patients. Folia Biol (Praha). 2008;54:102-8
39. Liu B, Wei J, Zou Z, Qian X, Nakamura T, Zhang W, Ding Y, Feng J, Yu L. Polymorphism of XRCC1 predicts overall survival of gastric cancer patients receiving oxaliplatin-based chemotherapy in Chinese population. Eur J Hum Genet. 2007;15:1049-53
40. Lu C, Spitz MR, Zhao H, Dong Q, Truong M, Chang JY, Blumenschein GR Jr, Hong WK, Wu X. Association between glutathione S-transferase pi polymorphisms and survival in patients with advanced nonsmall cell lung carcinoma. Cancer. 2006;106:441-7
41. Luscombe CJ, French ME, Liu S, Saxby MF, Farrell WE, Jones PW, Fryer AA, Strange RC. Glutathione S-transferase GSTP1 genotypes are associated with response to androgen ablation therapy in advanced prostate cancer. Cancer Detect Prev. 2002;26:376-80
42. Marsh S, Paul J, King CR, Gifford G, McLeod HL, Brown R. Pharmacogenetic assessment of toxicity and outcome after platinum plus taxane chemotherapy in ovarian cancer: the Scottish Randomised Trial in Ovarian Cancer. J Clin Oncol. 2007;25:4528-35
43. Matthias C, Harréus U, Strange R. Influential factors on tumor recurrence in head and neck cancer patients. Eur Arch Otorhinolaryngol. 2006Jan;263(1):37-42
44. Morari EC, Lima AB, Bufalo NE, Leite JL, Granja F, Ward LS. Role of glutathione-S-transferase and codon 72 of P53 genotypes in epithelial ovarian cancer patients. J Cancer Res Clin Oncol. 2006;132:521-8
45. Nagle CM, Chenevix-Trench G, Spurdle AB, Webb PM. The role of glutathione-S-transferase polymorphisms in ovarian cancer survival. Eur J Cancer. 2007;43:283-90
46. Okcu MF, Selvan M, Wang LE, Stout L, Erana R, Airewele G, Adatto P, Hess K, Ali-Osman F, Groves M, Yung AW, Levin VA, Wei Q, Bondy M. Glutathione S-transferase polymorphisms and survival in primary malignant glioma. Clin Cancer Res. 2004Apr15;10(8):2618-25
47. Okuno T, Tamura T, Yamamori M, Chayahara N, Yamada T, Miki I, Okamura N, Kadowaki Y, Shirasaka D, Aoyama N, Nakamura T, Okumura K, Azuma T, Kasuga M, Sakaeda T. Favorable genetic polymorphisms predictive of clinical outcome of chemoradiotherapy for stage II/III esophageal squamous cell carcinoma in Japanese. Am J Clin Oncol. 2007;30:252-7
48. Ott K, Lordick F, Becker K, Ulm K, Siewert J, Höfler H, Keller G. Glutathione-S-transferase P1, T1 and M1 genetic polymorphisms in neoadjuvant-treated locally advanced gastric cancer: GSTM1-present genotype is associated with better prognosis in completely resected patients. Int J Colorectal Dis. 2008;23:773-82
49. Pillot GA, Read WL, Hennenfent KL, Marsh S, Gao F, Viswanathan A, Cummings K, McLeod HL, Govindan R. A phase II study of irinotecan and carboplatin in advanced non-small cell lung cancer with pharmacogenomic analysis: final report. J Thorac Oncol. 2006;1:972-8
50. Rajagopal R, Deakin M, Fawole AS, Elder JB, Elder J, Smith V, Strange RC, Fryer AA. Glutathione S-transferase T1 polymorphisms are associated with outcome in colorectal cancer. Carcinogenesis. 2005;26:2157-63
51. Rajkumar T, Samson M, Rama R, Sridevi V, Mahji U, Swaminathan R, Nancy NK. TGFbeta1 (Leu10Pro), p53 (Arg72Pro) can predict for increased risk for breast cancer in south Indian women and TGFbeta1 Pro (Leu10Pro) allele predicts response to neo-adjuvant chemo-radiotherapy. Breast Cancer Res Treat. 2008;112:81-7
52. Stoehlmacher J, Park DJ, Zhang W, Yang D, Groshen S, Zahedy S, Lenz HJ. A multivariate analysis of genomic polymorphisms: prediction of clinical outcome to 5-FU/oxaliplatin combination chemotherapy in refractory colorectal cancer. Br J Cancer. 2004;91:344-54
53. Sweeney C, Nazar-Stewart V, Stapleton PL, Eaton DL, Vaughan TL. Glutathione S-transferase M1, T1, and P1 polymorphisms and survival among lung cancer patients. Cancer Epidemiol Biomarkers Prev. 2003Jun;12(6):527-33
54. Wrensch M, Wiencke JK, Wiemels J, Miike R, Patoka J, Moghadassi M, McMillan A, Kelsey KT, Aldape K, Lamborn KR, Parsa AT, Sison JD, Prados MD. Serum IgE, tumor epidermal growth factor receptor expression, and inherited polymorphisms associated with glioma survival. Cancer Res. 2006;66:4531-41
55. Yang P, Yokomizo A, Tazelaar HD, Marks RS, Lesnick TG, Miller DL, Sloan JA, Edell ES, Meyer RL, Jett J, Liu W. Genetic determinants of lung cancer short-term survival: the role of glutathione-related genes. Lung Cancer. 2002;35:221-9
56. Rigual NR, Anderson GR, Loree TR, Wiseman S, Alrawi S, Stoler DL. Molecular prognosticators and genomic instability in papillary thyroid cancer. Laryngoscope. 2005;115:1479-85
57. Ruzzo A, Graziano F, Kawakami K, Watanabe G, Santini D, Catalano V, Bisonni R, Canestrari E, Ficarelli R, Menichetti ET, Mari D, Testa E, Silva R, Vincenzi B, Giordani P, Cascinu S, Giustini L, Tonini G, Magnani M. Pharmacogenetic profiling and clinical outcome of patients with advanced gastric cancer treated with palliative chemotherapy. J Clin Oncol. 2006;24:1883-91
58. Ruzzo A, Graziano F, Loupakis F, Rulli E, Canestrari E, Santini D, Catalano V, Ficarelli R, Maltese P, Bisonni R, Masi G, Schiavon G, Giordani P, Giustini L, Falcone A, Tonini G, Silva R, Mattioli R, Floriani I, Magnani M. Pharmacogenetic profiling in patients with advanced colorectal cancer treated with first-line FOLFOX-4 chemotherapy. J Clin Oncol. 2007;25:1247-54
59. Sanyal S, Ryk C, De Verdier PJ, Steineck G, Larsson P, Onelöv E, Hemminki K, Kumar R. Polymorphisms in NQO1 and the clinical course of urinary bladder neoplasms. Scand J Urol Nephrol. 2007;41:182-90
60. Stoehlmacher J, Park DJ, Zhang W, Groshen S, Tsao-Wei DD, Yu MC, Lenz HJ. Association between glutathione S-transferase P1, T1, and M1 genetic polymorphism and survival of patients with metastatic colorectal cancer. J Natl Cancer Inst. 2002;94:936-42
61. Sreeja L, Syamala V, Hariharan S, Syamala VS, Raveendran PB, Sivanandan CD, Madhavan J, Ankathil R. Glutathione S-transferase M1, T1 and P1 polymorphisms: susceptibility and outcome in lung cancer patients. J Exp Ther Oncol. 2008;7:73-85
62. Medeiros R, Pereira D, Afonso N, Palmeira C, Faleiro C, Afonso-Lopes C, Freitas-Silva M, Vasconcelos A, Costa S, Osório T, Lopes C. Platinum/paclitaxel based chemotherapy in advanced ovarian carcinoma: glutathione S-transferase genetic polymorphisms as predictive biomarkers of disease outcome. Int J Clin Oncol. 2003;8:156-61
63. Worrall SF, Corrigan M, High A, Starr D, Matthias C, Wolf CR, Jones PW, Hand P, Gilford J, Farrell WE, Hoban P, Fryer AA, Strange RC. Susceptibility and outcome in oral cancer: preliminary data showing an association with polymorphism in cytochrome P450 CYP2D6. Pharmacogenetics. 1998;8:433-9
64. Kweekel DM, Koopman M, Antonini NF, Van der Straaten T, Nortier JW, Gelderblom H, Punt CJ, Guchelaar HJ. GSTP1 Ile105Val polymorphism correlates with progression-free survival in MCRC patients treated with or without irinotecan: a study of the Dutch Colorectal Cancer Group. Br J Cancer. 2008;99:1316-21
65. Ambrosone CB, Sweeney C, Coles BF, Thompson PA, McClure GY, Korourian S, Fares MY, Stone A, Kadlubar FF, Hutchins LF. Polymorphisms in glutathione S-transferases (GSTM1 and GSTT1) and survival after treatment for breast cancer. Cancer Res. 2001;61:7130-5
66. Bewick MA, Conlon MS, Lafrenie RM. Polymorphisms in manganese superoxide dismutase, myeloperoxidase and glutathione-S-transferase and survival after treatment for metastatic breast cancer. Breast Cancer Res Treat. 2008;111:93-101
67. Braun MS, Richman SD, Quirke P, Daly C, Adlard JW, Elliott F, Barrett JH, Selby P, Meade AM, Stephens RJ, Parmar MK, Seymour MT. Predictive biomarkers of chemotherapy efficacy in colorectal cancer: results from the UK MRC FOCUS trial. J Clin Oncol. 2008;26:2690-8
68. Kondratieva TV, Imyanitov EN, Togo AV, Zaitseva OA, Yatsuk OS, Romanenko SM, Bersnev VP, Hanson KP. L-MYC and GSTM1 polymorphisms are associated with unfavourable clinical parameters of gliomas. J Exp Clin Cancer Res. 2000;19:197-200
69. Lallas TA, McClain SK, Shahin MS, Buller RE. The glutathione S-transferase M1 genotype in ovarian cancer. Cancer Epidemiol Biomarkers Prev. 2000;9:587-90
70. DeMichele A, Aplenc R, Botbyl J, Colligan T, Wray L, Klein-Cabral M, Foulkes A, Gimotty P, Glick J, Weber B, Stadtmauer E, Rebbeck TR. Drug-metabolizing enzyme polymorphisms predict clinical outcome in a node-positive breast cancer cohort. J Clin Oncol. 2005;23:5552-9
71. Goode EL, Dunning AM, Kuschel B, Healey CS, Day NE, Ponder BA, Easton DF, Pharoah PP. Effect of germ-line genetic variation on breast cancer survival in a population-based study. Cancer Res. 2002;62:3052-7
72. Khedhaier A, Remadi S, Corbex M, Ahmed SB, Bouaouina N, Mestiri S, Azaiez R, Helal AN, Chouchane L. Glutathione S-transferases (GSTT1 and GSTM1) gene deletions in Tunisians: susceptibility and prognostic implications in breast carcinoma. Br J Cancer. 2003;89:1502-7
73. Lizard-Nacol S, Coudert B, Colosetti P, Riedinger JM, Fargeot P, Brunet-Lecomte P. Glutathione S-transferase M1 null genotype: lack of association with tumour characteristics and survival in advanced breast cancer. Breast Cancer Res. 1999;1:81-7
74. Nedelcheva Kristensen V, Andersen TI, Erikstein B, Geitvik G, Skovlund E, Nesland JM, Borresen-Dale AL. Single tube multiplex polymerase chain reaction genotype analysis of GSTM1, GSTT1 and GSTP1: relation of genotypes to TP53 tumor status and clinicopathological variables in breast cancer patients. Pharmacogenetics. 1998;8:441-7
75. Petros WP, Hopkins PJ, Spruill S, Broadwater G, Vredenburgh JJ, Colvin OM, Peters WP, Jones RB, Hall J, Marks JR. Associations between drug metabolism genotype, chemotherapy pharmacokinetics, and overall survival in patients with breast cancer. J Clin Oncol. 2005;23(25):6117-25
76. Sweeney C, McClure GY, Fares MY, Stone A, Coles BF, Thompson PA, Korourian S, Hutchins LF, Kadlubar FF, Ambrosone CB. Association between survival after treatment for breast cancer and glutathione S-transferase P1 Ile105Val polymorphism. Cancer Res. 2000;60:5621-4
77. Syamala VS, Sreeja L, Syamala V, Raveendran PB, Balakrishnan R, Kuttan R, Ankathil R. Influence of germline polymorphisms of GSTT1, GSTM1, and GSTP1 in familial versus sporadic breast cancer susceptibility and survival. Fam Cancer. 2008;7:213-20
78. Yang G, Shu XO, Ruan ZX, Cai QY, Jin F, Gao YT, Zheng W. Genetic polymorphisms in glutathione-S-transferase genes (GSTM1, GSTT1, GSTP1) and survival after chemotherapy for invasive breast carcinoma. Cancer. 2005;103:52-8
79. Sun XF, Ahmadi A, Arbman G, Wallin A, Asklid D, Zhang H. Polymorphisms in sulfotransferase 1A1 and glutathione S-transferase P1 genes in relation to colorectal cancer risk and patients' survival. World J Gastroenterol. 2005;11:6875-9
80. Wu X, Gu J, Wu TT, Swisher SG, Liao Z, Correa AM, Liu J, Etzel CJ, Amos CI, Huang M, Chiang SS, Milas L, Hittelman WN, Ajani JA. Genetic variations in radiation and chemotherapy drug action pathways predict clinical outcomes in esophageal cancer. J Clin Oncol. 2006;24:3789-98
81. Le Morvan V, Smith D, Laurand A, Brouste V, Bellott R, Soubeyran I, Mathoulin-Pelissier S, Robert J. Determination of ERCC2 Lys751Gln and GSTP1 Ile105Val gene polymorphisms in colorectal cancer patients: relationships with treatment outcome. Pharmacogenomics. 2007;8:1693-703
82. Kalikaki A, Kanaki M, Vassalou H, Souglakos J, Voutsina A, Georgoulias V, Mavroudis D. DNA repair gene polymorphisms predict favorable clinical outcome in advanced non-small-cell lung cancer. Clin Lung Cancer. 2009;10:118-23
83. Gordon MA, Gil J, Lu B, Zhang W, Yang D, Yun J, Schneider S, Groshen S, Iqbal S, Press OA, Rhodes K, Lenz HJ. Genomic profiling associated with recurrence in patients with rectal cancer treated with chemoradiation. Pharmacogenomics. 2006;7:67-88
84. Paré L, Marcuello E, Altés A, del Río E, Sedano L, Salazar J, Cortés A, Barnadas A, Baiget M. Pharmacogenetic prediction of clinical outcome in advanced colorectal cancer patients receiving oxaliplatin/5-fluorouracil as first-line chemotherapy. Br J Cancer. 2008;99:1050-5
85. Ashton LJ, Murray JE, Haber M, Marshall GM, Ashley DM, Norris MD. Polymorphisms in genes encoding drug metabolizing enzymes and their influence on the outcome of children with neuroblastoma. Pharmacogenet Genomics. 2007;17:709-17
86. Przygodzki RM, Bennett WP, Guinee DG Jr, Khan MA, Freedman A, Shields PG, Travis WD, Jett JR, Tazelaar H, Pairolero P, Trastek V, Liotta LA, Harris CC, Caporaso NE. p53 mutation spectrum in relation to GSTM1, CYP1A1 and CYP2E1 in surgically treated patients with non-small cell lung cancer. Pharmacogenetics. 1998;8:503-11
87. Goekkurt E, Al-Batran SE, Hartmann JT, Mogck U, Schuch G, Kramer M, Jaeger E, Bokemeyer C, Ehninger G, Stoehlmacher J. Pharmacogenetic analyses of a phase III trial in metastatic gastroesophageal adenocarcinoma with fluorouracil and leucovorin plus either oxaliplatin or cisplatin: a study of the arbeitsgemeinschaft internistische onkologie. J Clin Oncol. 2009;27:2863-73
88. Keam B, Im SA, Han SW, Ham HS, Kim MA, Oh DY, Lee SH, Kim JH, Kim DW, Kim TY, Heo DS, Kim WH, Bang YJ. Modified FOLFOX-6 chemotherapy in advanced gastric cancer: Results of phase II study and comprehensive analysis of polymorphisms as a predictive and prognostic marker. BMC Cancer. 2008;8:148
89. Ruwali M, Pant MC, Shah PP, Mishra BN, Parmar D. Polymorphism in cytochrome P450 2A6 and glutathione S-transferase P1 modifies head and neck cancer risk and treatment outcome. Mutat Res. 2009 In press
90. Huang MY, Wang YH, Chen FM, Lee SC, Fang WY, Cheng TL, Hou MF, Wang JY, Lin SR. Multiple Genetic Polymorphisms of GSTP1 313AG, MDR1 3435CC, and MTHFR 677CC highly correlated with early relapse of breast cancer patients in Taiwan. Ann Surg Oncol. 2008;15:872-80
91. Sweeney C, Ambrosone CB, Joseph L, Stone A, Hutchins LF, Kadlubar FF, Coles BF. Association between a glutathione s-transferase A1 promoter polymorphism and survival after breast cancer treatment. Int J Cancer. 2003;103:810-814
92. Dickersin K, Min YI, Meinert CL. Factors influencing publication of research results. J. Am. Med. Assoc. 1992;267:374-378
93. Callaham M, Wears RL, Weber E. Journal prestige, publication bias, and other characteristics associated with citation of published studies in peer-reviewed journals. JAMA. 2002;287(21):2847-2850
94. Tan NC, Mulley JC, Berkovic SF. Genetic association studies in epilepsy: "the truth is out there". Epilepsia. 2004Nov;45(11):1429-42
Author contact
Corresponding author: Davide Mauri, Dept. Med. Oncol., Gen. Hosp. of Lamia, Daravelia 27, TK 35100, Lamia, Greece. dvd.mauricom

 Global reach, higher impact
Global reach, higher impact