Impact Factor ISSN: 1449-1907
Int J Med Sci 2011; 8(4):309-314. doi:10.7150/ijms.8.309 This issue Cite
Research Paper
Monoclonal Antibodies against Nucleophosmin Mutants: Potentials for the Detection of Acute Myeloid Leukemia
1. Key Laboratory of Laboratory Medical Diagnostics, Ministry of Education, Department of Laboratory Medicine, Chongqing Medical University, Chongqing 400016, China
2. Center for Hematology, Southwest Hospital, Third Military Medical University, Chongqing 400016, China
3. Department of Laboratory Medicine, the First Affiliated Hospital, Chongqing Medical University, Chongqing 400016, China
4. Department of Hematology, the First Affiliated Hospital, Chongqing Medical University, Chongqing 400016, China
Received 2011-4-26; Accepted 2011-5-11; Published 2011-5-17
Abstract
Nucleophosmin (NPM1) gene mutations resulting in cytoplasmic delocalization of Nucleophosmin (NPMc+) are the most common genetic alteration in acute myeloid leukemia (AML). Here, we attempted to prepare monoclonal antibodies (mAbs) against NPM1 mutation A (NPM-mA) and investigated the mAbs' clinical utility in immunohistochemical detection of NPMc+AML. The pET-32a-NPM-mA vector with the whole open reading frame of the NPM-mA gene was constructed. E.coli BL21 transformed with the vector were induced to express the NPM-mA recombinant protein. BALB/c mice were immunized with the recombinant NPM-mA. Positive clones were selected by indirect ELISA and the mAbs were obtained. Immunohistochemistry was performed to detect the NPMc+ in bone marrow smears from 10 AML patients with NPM-mA. The results showed that the pET-32a-NPM-mA vector was successfully constructed and the NPM-mA recombinant protein was used to immunize the mice. Two positive clones (2G3 and 3F9) were selected. The mAbs against NPM-mA were raised, but did cross-react with wild type NPM1. The mAbs can be used to detect the cytoplasmic dislocation of NPM1 in all AMLs carrying NPM-mA. Our results show that anti-NPM-mA mAbs were produced. Though they would cross-react with wild type NPM1, the mAbs may still have potential in the detection of NPMc+AMLs.
Keywords: acute leukemia, nucleophosmin mutants, recombinant protein, monoclonal antibody
1. Introduction
Nucleophosmin (NPM1) is an ubiquitously expressed nucleo-cytoplasmic shuttling protein with prominent nucleolar localization [1, 2]. Previous studies have demonstrated that mutations of the NPM1 gene leading to aberrant cytoplasmic NPM1 expression (NPMc+) occur in about one-third of acute myeloid leukemias (AML) and 45% to 64% of AML with normal karyotype cases [3, 4]. The most common molecular variant of the NPM1 gene is mutation A, accounting for about 75-85% of cases. It is due to a duplication of TCTG tetranucleotide at the C-terminus of the NPM1 gene, which generates a nuclear export signal (NES) motif responsible for cytoplasmic accumulation of NPM1 [5-7]. Many observations indicate that the NPM1 mutation A (NPM-mA) is not only an AML-specific genetic event, but also remains stable during the course of the disease [6, 8, 9]. Meanwhile, the AML with cytoplasmic NPM1 (NPMc+AML) exhibits distinctive biological and clinical features and has been included as a new provisional entity in the 2008 World Health Organization (WHO) classification of myeloid neoplasms [5, 10-13]. Thus, the analysis of NPM1 mutations may emerge as an initial screening step in the diagnostic/prognostic work-up of AML and could also serve to monitor minimal residual disease (MRD) [14].
Over the past five years, several qualitative and quantitative molecular assays for identifying NPM1 mutations have been developed. Currently available screening of NPM1 mutations using conventional polymerase chain reaction (PCR) followed by capillary electrophoresis is rather time-consuming, technical-demanding and laborious [15]. Alternatively, the simple, inexpensive and specific immunohistochemical tests (IHC) which indirectly detect aberrant cytoplasmic accumulation of NPM1 proteins can serve as a surrogate to molecular studies [16-18]. To popularize IHC detection of cytoplasmic NPM1 in clinical diagnosis/prognosis of NPMc+AML, we need to prepare the anti-NPM-mA monoclonal antibodies (mAbs) as the primary antibody in IHC assay.
In 1999, Cordell et al prepared the first panel of mAbs associated with NPM1 protein, two of which recognized the N-terminal portion of NPM1 present in NPM-ALK fusion protein and the third was specific for wild-type NPM1 (NPM-wt). Their main purpose was to detect the NPM-ALK fusion protein created by the t(2;5) chromosomal translocation in anaplastic large-cell lymphoma (ALCL) [19]. Nowadays, extensive detection of cytoplasmic dislocation of NPM1 by IHC has been performed using aspecific antibodies that bind both the NPM-wt and NPM-mA proteins. In IHC assay labeling with this kind of mAbs, the cytoplasmic subcellular localization of NPM1 may not be closely associated with NPM1 gene mutations probably because of NPM1 diffusion during the tissue fixation and the influence of fixatives [20]. Thus, production of anti-NPM-mA mAbs for routine diagnostic of NPMc+AML is of critical importance.
To date, most detections of cytoplasmic NPM1 by IHC have been carried out in bone marrow biopsies. However, not all hematological centers, especially in developing countries, adopt bone marrow biopsy as a frontline diagnostic procedure for AML. Hence, the ability to detect cytoplasmic NPM1 on bone marrow smears would be advantageous. In view of this, we attempted to produce the mAbs that were specific for NPM-mA protein and preliminarily explore the application of IHC labeling with these mAbs on bone marrow smears of AML patients with NPM1 mutations.
2. Materials and Methods
2.1 PCR for amplification of NPM-mA gene
According to the published sequence of the NPM-mA in GenBank (no.AY740634), a pair of specific primers were designed to amplify the ORF of NPM-mA gene from pEGFP-C1-NPM-mA vectors, which were kindly provided by Dr. B Falini (Institute of Hematology, University of Perugia, Perugia, Italy). The forward and backward primers were: 5'-CGGGATCCATCGAAGGTCGTGAAGATTCGATGGACAT-3', and 5'-CGCGCGACCGAGCGGAAGCTTCTATTTTCTTAAAGAGAC-3'. Underlined nucleotides represent the BamH I and Hind III site, respectively. PCR conditions included pre-denaturation at 98°C for 5 min; 32 cycles of denaturation at 98°C for 20 sec, annealing at 56°C for 20 sec, and extension at 72°C for 80 sec; followed by a final extension at 72°C for 5 min.
2.2 Construction of expressing vector pET-32a-NPM-mA
After being checked by using 1% agarose gel electrophoresis and retrieved utilizing the MinElute Gel Extration Kit (Tiangen, Beijing, China), the amplification products (NPM-mA gene) were cloned into the BamH I and Hind III site of the pET-32a plasmids creating fusion vectors pET-32a-NPM-mA in the presence of T4 DNA Ligase (TaKara, Tokyo, Japan). The fusion vectors were subsequently transformed into E. coli DH5α cloning vectors and E. coli BL21 (DE3) expression bacteria and then grown overnight at 37°C in Luria-Bertani (LB) medium with ampicillin (100 μg/ml). The positive expression clones were screened out by colony PCR. After extracted by a commercial kit (Huashun, Shanghai, China), pET-32a-NPM-mA was further identified by restriction enzyme digestions and DNA sequencing (Invitrogen, Shanghai, China). The positive expression BL21 (DE3) was stored in LB containing 15% glycerine at -80°C.
2.3 Expression and Purification of NPM-mA protein
Overnight culture of pET-32a-NPM-mA transformed BL21 (1 ml) was inoculated to 1000 ml LB/amp and cultured at 37°C for 3-4 h at 200 rpm until OD600 reached 0.3-0.4, then 0.1 mM IPTG (TaKara, Tokyo, Japan) was added to induce protein expression. The culture was incubated for 4 h at 37°C at 200 rpm before harvesting the cells by centrifugation (15,000×g, 20 min, 4°C) and the cell pellets were washed and lysed by sonication on ice. After centrifuged at 15,000×g for 20 min, the supernatant was analyzed by SDS-PAGE as the soluble fraction and the remaining cell pellet as the insoluble fraction to determine whether native or denaturing conditions were necessary for protein purification. The supernatant was loaded to His-Bind-Resins affinity column (Novagen, Darmstadt, Germany) to purify the fusion protein. The purified protein was dialysed against phosphate-buffered saline (PBS) overnight at 4°C and stored at -80°C before analyzed by SDS-PAGE and quantitated by using the BCA Protein Assay Kit (Beyotime, Shanghai, China).
2.4 Immunizations
Five-week old female BALB/c mice initially received subcutaneous injection of purified NPM-mA fusion protein (100 μg) emulsified in an equal volume of Freund's complete adjuvant (Sigma, St. Luis, MO, USA). A second injection of the same dose of NPM-mA protein in incomplete Freund's adjuvant was administered 2 weeks later. 10-14 days after the second booster, the mice were then given NPM-mA fusion protein without adjuvant intraperitoneally. An additional intraperitoneal injection of 100 μg of antigen was given 2 days before harvesting the spleen cells. Experiments with injected mice were performed under the guidelines for care and use of experimental animals.
2.5 Cellular fusions
When the anti-NPM-mA antibodies titre of mice serum reached 1:1024 checked by indirect enzyme-linked immunosorbent assay (ELISA), myeloma cells line SP2/0 (106) were fused with splenocytes (107) by the addition of 45% polyethylene glycol (PEG-4000). Hybridomas were selected in HAT medium (Gibco, Carlsbad, CA, USA) and cultured in 96-well plates with BALB/c (8 weeks old) peritoneal macrophages cells as feeder cells at 37°C in 5% CO2 in air. When single colonies of cells were visualized, cell culture supernatants were obtained and screened for the presence of anti-NPM-mA antibodies using indirect ELISA. Selected positive hybridomas were expanded and subcloned by limiting dilution.
2.6 Purification and characterization of mAbs
After typed by mouse monoclonal antibody isotyping kit (Sigma, St. Luis, MO, USA), the prepared mAbs were purified from cell-culture supernatant by affinity chromatography. Indirect ELISA was then carried out on NPM-wt and NPM-mA coated plates to check the antigenic characterization of mAbs.
2.7 Patients
Bone marrow/peripheral blood smears were obtained from de novo AML patients, who were from Southwest Hospital of Third Military Medical University and The First Affiliated Hospital of Chongqing Medical University (Chongqing, China) between 2008 and 2009. Informed consent was obtained from all patients, and the study was approved by the ethics committees of the participating institutions. Ten positive samples with NPM-mA were selected by direct sequencing.
2.8 Immunohistochemistry
Slides were incubated with the anti-NPM antibody we prepared (1:100 in Tris-buffered saline) overnight at 4°C. Immunohistochemistry was performed using the Streptavidin-Peroxidase (SP)-9000 kit (Zhongshan, Beijing, China) according to the manufacturer's instructions. Peroxidase activity was revealed with 3-3-diaminobenzidine-copper sulphate (Sigma, St. Luis, MO, USA) to obtain brownblack granules. The subcellular distribution of NPM-mA was assessed after counterstained with hematoxylin. PBS was used as a negative control for the anti-NPM antibody.
3. Results
3.1 PCR for amplification of NPM-mA gene
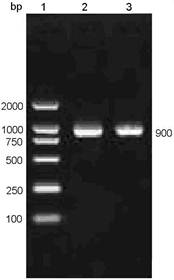
Using a pair of primers specific for NPM-mA gene, a DNA fragment of approximately 900 bp size was amplified from the pEGFP-C1-NPM-mA plasmids by PCR technique (Figure 1), which corresponded to the full length of open reading frame (ORF) of the NPM-mA gene (935 bp).
PCR amplifying the full sequence of ORF of the NPM-mA gene. The PCR products amplified with a pair of primers against the NPM-mA gene were analyzed by 1% agarose gel electrophoresis.1: DL2000 markers; 2-3: products of PCR.
3.2 Construction of recombinant vector pET-32a-NPM-mA
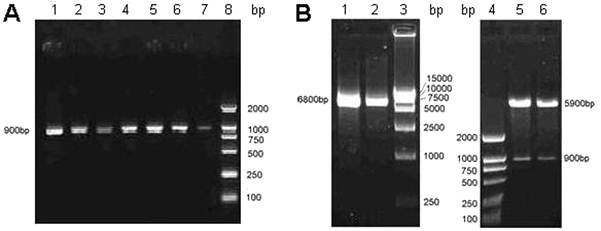
To generate a recombinant human encoding the NPM-mA protein, the pET-32a-NPM-mA vector was cloned. As shown in Figure 2, the pET-32a-NPM-mA vector was successfully constructed as verified by bacterial colony PCR (Figure 2A), restriction enzyme digestions (Figure 2B) and DNA sequencing (data not shown).
Cloning of the recombinant vector pET-32a-NPM-mA. A, Bacterial colony PCR for the detection of the BL21 (DE3) clones with the target prokaryotic expression vector pET-32a-NPM-mA. 1-7: 7 colonies of bacteria selected on LB medium with ampicillin; 8: DL2000 markers. B, Double endonuclease digestion of the prokaryotic expression vector pET-32a-NPM-mA. 1-2: pET-32a-NPM-mA; 3: DL15000 markers; 4: DL2000 markers; 5-6: double digestion with the BamH I and Hind III.
3.3 Expression and purification of recombinant NPM-mA antigen
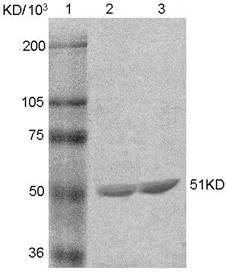
The NPM-mA fusion protein was resoluble and detected in the culture supernatants. SDS-PAGE analysis of the fusion protein is displayed in Figure 3. Expression and purification of the NPM-mA antigen were performed as described in Materials and Methods. The concentration of the purified recombinant protein was 1.95 μg/μl determined by BCA protein assay.
SDS-PAGE assay of the purified NPM-mA fusion protein. 1: protein size markers; 2-3: recombinant NPM-mA fusion protein.
3.4 Production of the anti-NPM mAbs
For selecting the clones with mAbs against NPM-mA, the supernatants of fused cells were assayed by indirect ELISA. Two clones were found to be positive in the ELISA screen of culture supernatants (designated as 2G3 and 3F9). The 2G3 clones which exhibited good growth characteristics and antibody production were subjected to subcloning. Antibodies secreted by the 2G3 clones were found to be IgG isotype. The specificity of the mAbs against NPM-mA was assessed by indirect ELISA, and the mAbs were able to react with both NPM-wt and NPM-mA.
3.5 Immunohistochemical staining for the cases with NPM1 mutations
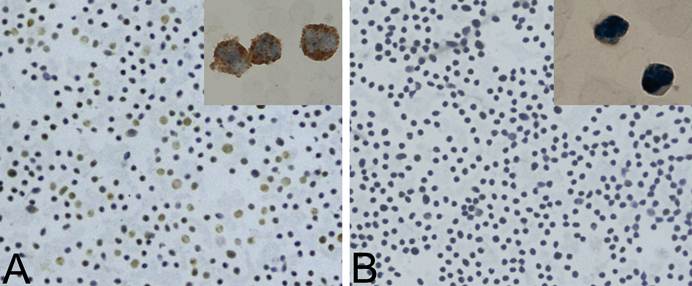
Ten AML samples had been confirmed to bear NPM-mA by direct sequencing (data not shown). To validate the mAbs against NPM-mA as a diagnostic tool for AML patients, bone marrow or peripheral blood samples were analyzed by IHC, using the 2G3 mAb. The cytoplasmic dislocation of NPM1 was observed in all 10 samples with NPM1 mutations, without staining in the cytoplasm of leukemic blasts in the negative control. Figure 4 shows representative results from a NPMc+AML patient.
Immunohistochemistry analyses of NPMc+AML samples using the 2G3 mAb. A, The cytoplasmic dislocation of NPM1 protein was observed in a representative bone marrow smear from NPMc+AML patients. Brownblack coarse granules in the cytoplasm of leukemic cells are shown. B, Negative control; the bone marrow from the same case as in (A) was stained with PBS substituting for the 2G3 mAb.
4. Discussion
Mutations involving the NPM1 gene are the most frequent genetic aberrations of AML, and define a clinically distinct subset of AML [21, 22]. NPM1 gene mutations always result in cytoplasmic dislocation of NPM1, which is the immunohistochemical hallmark of NPMc+AML. Immunohistochemistry may be a simple, rapid screening test for putative NPM1 gene alterations in a wide range of human hematological malignancies [17, 21]. The crucial step of immunohistochemical detection is to generate mAbs directed against NPM1 mutants.
Currently, immunohistochemistry is usually performed with mAbs that recognize wild-type and mutated NPM1 proteins. In the present study, we attempted to prepare mAbs against NPM-mA. The specific mAbs reacting with NPM-mA have the advantage of directly detecting the NPM-mA protein in leukemic cells. Firstly, some technical factors, such as NPM1 diffusion during tissue fixation and the use of different fixatives, may result in the incomplete concordance between NPMc+ and NPM mutations status in some cases [20, 23]. Furthermore, because NPM1 is a nuclear-cytoplasmic shuttling protein and highly expressive in proliferative cells, the small fraction of NPM-wt protein may pathologically present in the cytoplasm of tumor cells [24]. As a result, IHC labeling with the mAbs against NPM-mA and NPM-wt may detect the NPM-wt existing in the cytoplasm and cause false positives. In this study, we analyzed the antigen epitope of NPM-mA protein and confirmed it may exist in the C-terminal domain of the NPM-mA by using the Protean module of DNAstar analysis software. However, our results revealed that the obtained mAbs did cross-react with NPM-wt. A possible explanation is that the distinction between NPM-mA and NPM-wt is small (only a tetranucleotide insertion located at the C-terminus of NPM-mA) [25]. So the 2G3 mAb we obtained may not interat with a specific epitope generated by the NPM1 mutation. Recently, Gruszka et al [26] have raised a mAb (T26) only against NPM1 mutants by using a 19-aminoacid polypeptide immunogen (CLAVEEVLSRK) containing the unique C-terminus of the NPM-mA protein. It indicated that the specific polypeptide generated by the C-terminus of the NPM1 (type A) mutation may be an optimal immunogen.
Over the past five years, IHC detection of NPMc+ on bone marrow biopsies has been widely carried out. However, as bone marrow biopsies are not always performed for the diagnosis of AML, especially in developing countries, to detect NPMc+ on bone marrow smears would be more advantageous. IHC assay was performed using the 2G3 mAb on bone marrow/peripheral blood smears of 10 AML patients with NPM-mA, and significant correlation was found between NPMc+ and NPM1 mutations status, which is not consistent with the finding of Mattsson et al [27]. They reported that the immunocytochemical staining should not be used as a surrogate for NPM1 mutations in AML, due to the high false positive and negative rates for NPMc+ in cell smears. The possible reasons for the two different results include the different anti-NPM antibodies (2G3 mAb or NA24 mAb) and the different method used (SP method or immunoalkaline phosphatase method).
In summary, we put forward the production of mAbs that specifically recognize NPM1. Although the mAbs prove to react with NPM-mA and NPM-wt, this result provides valuable information in that the mAbs against NPM-mA cannot be raised using the recombinant NPM-mA protein as immunogen. Furthermore, the complete correlation between NPMc+ in cell smears and NPM1 mutations status has been found in clinical samples by IHC using the 2G3, which would be utilized for other potential techniques,such as immunofluorescence, flow cytometry, etc.
Acknowledgements
We would like to thank Dr. Falini B in University of Perugia for the gift of pEGFP-C1-NPM-mA vectors. This project was supported by a grant from National Natural Science Foundation of China (No. 30872418) and Natural Science Foundation Project of CQ CSTC (No. 2010BB5363).
Conflict of Interest
The authors have declared that no conflict of interest exists.
References
1. Borer RA, Lehner CF, Eppenberger HM. et al. Major nucleolar proteins shuttle between nucleus and cytoplasm. Cell. 1989;56:379-90
2. Nishimura Y, Ohkubo T, Furuichi Y. et al. Tryptophans 286 and 288 in the C-terminal region of protein B23.1 are important for its nucleolar localization. Biosci Biotechnol Biochem. 2002;66:2239-42
3. Schnittger S, Schoch C, Kern W. et al. Nucleophosmin gene mutations are predictors of favorable prognosis in acute myelogenous leukemia with a normal karyotype. Blood. 2005;106:3733-9
4. Boissel N, Renneville A, Biggio V. et al. Prevalence, clinical profile, and prognosis of NPM mutations in AML with normal karyotype. Blood. 2005;106:3618-20
5. Falini B, Nicoletti I, Martelli MF. et al. Acute myeloid leukemia carrying cytoplasmic/mutated nucleophosmin (NPMc+ AML): biologic and clinical features. Blood. 2007;109:874-85
6. Liso A, Bogliolo A, Freschi V. et al. In human genome, generation of a nuclear export signal through duplication appears unique to nucleophosmin (NPM1) mutations and is restricted to AML. Leukemia. 2008;22:1285-9
7. Falini B, Bolli N, Shan J. et al. Both carboxy-terminus NES motif and mutated tryptophan(s) are crucial for aberrant nuclear export of nucleophosmin leukemic mutants in NPMc+ AML. Blood. 2006;107:4514-23
8. Chou WC, Tang JL, Lin LI. et al. Nucleophosmin mutations in de novo acute myeloid leukemia: the age-dependent incidences and the stability during disease evolution. Cancer Res. 2006;66:3310-6
9. Palmisano M, Grafone T, Ottaviani E. et al. NPM1 mutations are more stable than FLT3 mutations during the course of disease in patients with acute myeloid leukemia. Haematologica. 2007;92:1268-9
10. Falini B, Martelli MP, Bolli N. et al. Acute myeloid leukemia with mutated nucleophosmin (NPM1): is it a distinct entity? Blood. 2011;117:1109-20
11. Becker H, Marcucci G, Maharry K. et al. Favorable prognostic impact of NPM1 mutations in older patients with cytogenetically normal de novo acute myeloid leukemia and associated gene- and microRNA-expression signatures: a Cancer and Leukemia Group B study. J Clin Oncol. 2010;28:596-604
12. Vardiman JW, Thiele J, Arber DA. et al. The 2008 revision of the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia: rationale and important changes. Blood. 2009;114:937-51
13. Pasqualucci L, Liso A, Martelli MP. et al. Mutated nucleophosmin detects clonal multilineage involvement in acute myeloid leukemia: Impact on WHO classification. Blood. 2006;108:4146-55
14. Papadaki C, Dufour A, Seibl M. et al. Monitoring minimal residual disease in acute myeloid leukaemia with NPM1 mutations by quantitative PCR: clonal evolution is a limiting factor. Br J Haematol. 2009;144:517-23
15. Noguera NI, Ammatuna E, Zangrilli D. et al. Simultaneous detection of NPM1 and FLT3-ITD mutations by capillary electrophoresis in acute myeloid leukemia. Leukemia. 2005;19:1479-82
16. Falini B, Mason DY. Proteins encoded by genes involved in chromosomal alterations in lymphoma and leukemia: clinical value of their detection by immunocytochemistry. Blood. 2002;99:409-26
17. Falini B, Mecucci C, Tiacci E. et al. Cytoplasmic nucleophosmin in acute myelogenous leukemia with a normal karyotype. N Engl J Med. 2005;352:254-66
18. Falini B, Martelli MP, Bolli N. et al. Immunohistochemistry predicts nucleophosmin (NPM) mutations in acute myeloid leukemia. Blood. 2006;108:1999-2005
19. Cordell JL, Pulford KA, Bigerna B. et al. Detection of normal and chimeric nucleophosmin in human cells. Blood. 1999;93:632-42
20. Konoplev S, Huang X, Drabkin HA. et al. Cytoplasmic localization of nucleophosmin in bone marrow blasts of acute myeloid leukemia patients is not completely concordant with NPM1 mutation and is not predictive of prognosis. Cancer. 2009;115:4737-44
21. Thiede C, Koch S, Creutzig E. et al. Prevalence and prognostic impact of NPM1 mutations in 1485 adult patients with acute myeloid leukemia (AML). Blood. 2006;107:4011-20
22. Falini B, Sportoletti P, Martelli MP. Acute myeloid leukemia with mutated NPM1: diagnosis, prognosis and therapeutic perspectives. Curr Opin Oncol. 2009;21:573-81
23. Falini B, Martelli MP, Pileri SA. et al. Molecular and alternative methods for diagnosis of acute myeloid leukemia with mutated NPM1: flexibility may help. Haematologica. 2010;95:529-34
24. Ochs R, Lischwe M, O'Leary P. et al. Localization of nucleolar phosphoproteins B23 and C23 during mitosis. Exp Cell Res. 1983;146:139-49
25. Falini B, Bolli N, Liso A. et al. Altered nucleophosmin transport in acute myeloid leukaemia with mutated NPM1: molecular basis and clinical implications. Leukemia. 2009;23:1731-43
26. Gruszka AM, Lavorgna S, Consalvo MI. et al. A monoclonal antibody against mutated nucleophosmin1 for the molecular diagnosis of acute myeloid leukemias. Blood. 2010;116:2096-102
27. Mattsson G, Turner SH, Cordell J. et al. Can cytoplasmic nucleophosmin be detected by immunocytochemical staining of cell smears in acute myeloid leukemia? Haematologica. 2010;95:670-3
Author contact
Corresponding author: Ling Zhang, Department of Laboratory Medicine, Chongqing Medical University, 1#, Yixueyuan Road, Chongqing, 400016, China. Tel: +86 023-68485223, Fax: +86 023-68485005; Email: lingzhangedu.cn

 Global reach, higher impact
Global reach, higher impact