Impact Factor ISSN: 1449-1907
Int J Med Sci 2009; 6(4):156-159. doi:10.7150/ijms.6.156 This issue Cite
Research Paper
Gene expression analysis of human red blood cells
1. Department of Medicine, Division of Cardiology and Angiology, Universitätsklinikum Düsseldorf, Heinrich-Heine-University, 40225 Düsseldorf, Germany
2. Biologisch-Medizinisches Forschungszentrum, Heinrich-Heine-University, 40225 Düsseldorf, Germany
3. Profil Institut für Stoffwechselforschung, Hellersbergstrasse 9, 41461 Neuss, Germany
4. Klinik für Kardiologie, Herzzentrum Wuppertal, Universität Witten/ Herdecke, Wuppertal, Germany
* Both authors equally contributed in this work
Received 2009-2-5; Accepted 2009-4-27; Published 2009-4-28
Abstract
Understanding of molecular mechanisms governing the enucleating phenomena of human erythrocytes is of major importance in both fundamental and applied studies. Total RNA (n=7) from human RBCs (purity of erythrocyte preparation >99,99%) was tested using 2100 Bioanalyzer (Agilent, USA), and transcribed to cDNA. Microarray analysis was performed with the Human Genome Focus GeneChip (Affymetrix, USA), containing 8500 transcripts corresponding to 8400 human genes. Here we report that human RBCs contain typical eukaryotic RNA with 28S- and18S-rRNA standard bands. Microarray studies revealed the presence of transcripts of 1019 different genes in erythrocytic RNA. Gene Ontology analysis recognized 859 genes involved in general biological processes: 529 genes for cellular metabolism, 228 genes for signal transduction, 104 genes for development, 107 genes for immune response, 62 genes for protein localization, 53 genes for programmed cell death, and 5 genes for autophagy. A number of genes responsible for transcription, translation, RNA-stabilisation as well as for apoptosis and anti-apoptosis have been identified for the first time in circulating human RBCs. The presented data shed new light on the genetic determination of erythropoiesis, apoptosis and may have implications on the pathophysiology and diagnosis of various diseases involving red blood cells.
Keywords: red blood cell, gene expression analysis
Introduction
Human erythrocytes discard their nucleus during maturation, and are thought not to be able to synthesise proteins. Research in this field can be divided into the following: (I) gene expression analysis of erythropoietic progenitor cells; (II) biochemical characterization of nucleotide and protein synthesis during the life cycle of nucleated erythrocytes in vertebrates; (III) molecular aspects of malaria pathogenesis during RBC development; (IV) and genomic and proteomic analysis of gene expression in normal adult human erythrocytes.
Thus, current array data showed that most genes expressed in haematopoietic stem cells are developmentally regulated and associated with cell self-renewal 1 as well as survival, differentiation and/or migration/adhesion 2. Genes enriched in committed progenitors were mostly associated with haematopoietic differentiation, immune regulation, and metabolism 1. It was shown, that both the rate and extent of transcription in mature erythrocyte nuclei from chicken 3 and newt 4 were much reduced as compared to reticulocytes from this species. Previous studies of malaria pathogenesis did not shed light on gene regulation mechanisms in respect to erythrocyte development, though some regulatory elements have been proposed 5.
A part of human erythrocytic proteins were identified 6, 7, 8, 9. The number of genomic studies of RBCs is at present limited and represented either by micronucleus assay as a marker of classification of RNA- positive reticulocytes and erythrocytes 10, 11, 12 or by attempts to describe of human erythroid gene activity, planed to be finished in 5-10 years 13. At present no information regarding gene expression in human RBCs is available.
In contrast, according to recent data, there is a strong evidence that anucleate platelets contain a functional spliceosome 14, mRNAs 15, rRNA, rough endoplasmic reticulum and polyribosomes, as well as numerous translation factors including 3'-UTR RNA- and poly(A)-binding protein 16. It is therefore believed that platelets maintain functionally intact protein translational capabilities accompanied by posttranslational modifications 17. Recently we were able to detect RNA in washed human RBCs 18. In this study we used microarray technique to identify genes possibly present and translated in human RBCs.
Study design
Cell isolation
Whole blood was taken from healthy human volunteers (n=7) and collected in tubes containing Natrium heparin. RBCs were isolated as described earlier 18. In short, an open syringe without piston was closed at the tip, filled with whole blood and kept upright, and then centrifugated at 800g for 20 min at 4°C. The resulting plasma was discarded. After opening the syringe at the tip, about 2/3 of the sedimented RBCs were allowed to carefully drop out of the syringe. Special attention was paid to not disturb the WBC layer above the RBCs. This simple method to purify RBCs was superior to methods using density gradients or magnetic beads.
RNA isolation and probe synthesis
Total RNA from RBCs was purified using reagents provided in the PAXgene Blood RNA Kit (Qiagen, Germany) and tested with RNA LabChip Kit by 2100 Bioanalyzer (Agilent, USA). cDNA was synthesised from 5µg total RNA using the SuperScript Double-Stranded cDNA Synthesis Kit (Invitrogen, USA) and purified according to the manufactures' instruction (GeneChip Sample Cleanup Module, Affymetrix, USA). Biotin-labeled cRNA was synthesized using the BioArray HighYield RNA Transcript Labeling Kit (Enzo Life Sciences, USA) and purified using GeneChip Sample Cleanup Module (Affymetrix, USA). Yield and size distribution of the labeled transcripts were determined with NanoDrop (Kisker, Germany) and 2100 Bioanalyzer (Agilent, USA). Fragmentation was carried out using the fragmentation buffer from GeneChip Sample Cleanup Module (Affymetrix, USA).
Microarray and gene ontology analysis
10 µg of fragmented cRNA were hybridised to the Human Genome Focus Array (Affymetrix, USA). After hybridisation, GeneChips were automatically stained with streptavidin-phycoerythrin by using a fluidic station (Affymetrix, USA). Microarrays were scanned by GeneChip Scanner (Affymetrix, USA). The resulting images were processed by the accompanying software (Microarray Suite 5.0; Affymetrix, USA). A global scaling approach was used to normalize signal intensities (TGT value = 500). Genes that were present in all 7 arrays are reported and subjected to further analysis. For classification of the resulting genes the gene ontology browser (Netaffx, Affymetrix, USA) was used.
Results and discussion
Characterization of total RNA of human RBCs
The purity of erythrocyte fraction achieved 99,99997% and was confirmed by Pappenheim staining of blood slides, flow cytometry (MÖLAB, Germany) and FACS analysis (Cytomics FC 500 CXP, Beckman Coulter, Germany) using labeling with human leucocyte- (CD45) or platelet specific (CD42) antibodies (Table 1).
The purity of human RBCs fraction tested by independent methods.
| blood cell count (n=5) | RBCs (cells/µL) | white blood cells (cells/µL) | platelets (cells/µL) |
|---|---|---|---|
| cytometer | 6600000±2000 | n.d.* | n.d.* |
| FACS | 6300000±1520 | n.d.* | n.d.* |
| blood slide | 6400000±120 | 0±0 | 2±1 |
| *according to the resolution options of the cytometer n.d. correspond to < 100 cells/µL | |||
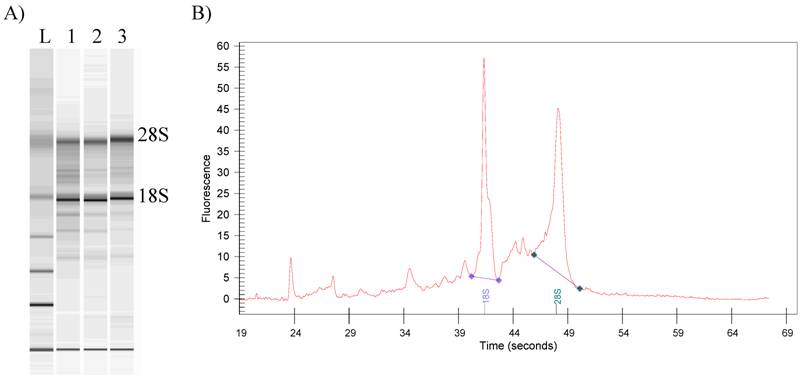
Human erythrocyte lack a nucleus and are thought to be void of protein synthesis. In contrast, we have found that total RNA from human RBCs resembles typical eukaryotic RNA with 5S-80S sedimentation distributions, and contains standard 28S- and18S-rRNA bands (Fig. 1). Total RNA from nucleated avian erythrocytes was discovered to have from 5 to 60 S sedimentation rates 3. Identification of each unique RNA-class within the RNA pool as well as genetic mechanisms from both nucleated and anucleate erythrocytes awaits future studies.
Analysis of total human RNA of RBCs (Bioanalyzer 2100 Agilent, USA): A) L- RNA 6000 ladder (Ambion, USA); 1-3 - total RNA of human RBCs from different donors; B) typical electropherogram of total RNA of human RBCs
Microarray analysis of RNA from human RBCs
Recent proteomic studies of RBCs based on 1D/2D-electrophoresis 6, 7 or mass spectrometry assay 8, 9, allowed to recognize 272 proteins. Our data generated from microarray studies (n=7) evidence the presence of transcripts for 1019 genes in RNA of human RBCs including the above mentioned 272 proteins. The complete array dataset with genes reported has been deposited in the Gene Expression Omnibus database (accession number - GSE3674).
It was found 529 genes for cellular metabolism (among them 96 genes for protein biosynthesis), 228 genes for signal transduction (among them 112 genes for intracellular signalling cascade), 104 genes for development, 107 genes for immune response, 62 genes for protein localization, and only 53 genes for programmed cell death as well as 5 genes for autophagy. The function of remainder (160 genes) is yet unknown.
In our work the percentage of genes sorted according to key developmental functions corresponds to results presented by Kakhniashvili and Tyang 8, 9. Interestingly, human RBCs contain 40-50% of genes encoding cell cycle processes (including 3-5% of genes for transcription/translation) as compared to only 10-20% of genes responsible for self-destruction processes.
For the first time we report about the presence of genes in human RBCs encoding initiation, activation and regulation of transcription and translation (for instance RNA polymerises I,II,III, zinc/PHD finger- DNA-binding proteins, cysteinyl, lysyl-tRNA synthetase), important RNA-stabilising factor - poly(A) binding protein, anti-apoptotic proteins (for instance beclin 1, reticulon 4, BCL2, IAP) together with genes for RNA degradation (for example ribonuclease T2) as well as genes encoding typical apoptotic proteins such as cyclooxygenase, apoptotic protease activating factor, caspase 8. Other authors were able to show a protein synthesis in human platelets by megakaryocyte-derived mRNAs 19. The finding of RNA in anucleate cells like erythrocytes support the idea of nucleus independent protein synthesis and supports data 20 about possible mechanism of globin m-RNA stability in human RBCs.
Further experiments are needed to understand the mechanisms and the biological meaning of these findings. But gene expression profiling of human erythrocyte could be an important key for understanding the machinery of anucleate protein synthesis and its meaning in the pathophysiology of diseases.
Acknowledgements
This work was supported by the Deutsche Forschungsgemeinschaft, Sonderforschungsbereich 612 (to M. Kelm) and Ke405/4-3 (to M. Kelm). The indispensable technical assistance of Katharina Lysaja is gratefully acknowledged.
Conflict of Interest
The authors have declared that no conflict of interest exists.
References
1. Terskikh AV, Miyamoto T, Chang C, Diatchenko L, Weissman IL. Gene expression analysis of purified hematopoietic stem cells and committed progenitors. Blood. 2003;102:94-101
2. Georgantas RW3, Tanadve V, Malehorn M. et al. Microarray and serial analysis of gene expression analyses identify known and novel transcripts overexpressed in hematopoietic stem cells. Cancer Res. 2004;64:4434-4441
3. Wiersma PA, Cox GS. Synthesis of messenger-like RNA in avian erythrocyte nuclei. Arch Biochem Biophys. 1985;242:90-103
4. Grasso JA, Chromey NC, Moxey CF. Biochemical characterization of RNA and protein synthesis in erythrocyte development. The Journal of Cell Biology. 1977;73:206-222
5. Rayner JC, Tran TM, Corredor V. et al. Dramatic difference in diversity between Plasmodium falciparum and Plasmodium vivax reticulocyte binding-like genes. Am j Trop Med Hyg. 2005;72:666-674
6. Copeland BR, Todd SA, Furlong CE. High resolution two-dimensional gel electrophoresis of human erythrocyte membrane proteins. High resolution two-dimensional gel electrophoresis of human erythrocyte membrane proteins. Am J Hum Genet. 1982;34:15-31
7. Low TY, Seow TK, Chung MC. Separation of human erythrocyte membrane associated proteins with one-dimensional and two-dimensional gel electrophoresis followed by identification with matrix-assisted laser desorption/ionization-time of flight mass spectrometry. Proteomics. 2002;2:1229-1239
8. Kakhniashvili DG, Bulla LAJr, Goodman SR. The human erythrocyte proteome: analysis by ion trap mass spectrometry. Mollecular und Cellular Proteomics. 2004;3:501-509
9. Tyan YC, Jong SB, Liao JD. et al. Proteomic profiling of erythrocyte proteins by proteolytic digestion chip and identification using two-dimensional electrospray ionization tandem mass spectrometry. Journal of Proteome Research. 2005;4:748-757
10. Shelby MD. A simplified and rapid method for scoring micronucleated erythrocytes in human blood. Mutat.Res. 2002;515:1
11. Dertinger SD, Camphausen K, Macgregor JT. et al. Three-color labeling method for flow cytometric measurement of cytogenetic damage in rodent and human blood. Environ.Mol.Mutagen. 2004;44:427-435
12. Grawe J, Biko J, Lorenz R. et al. Evaluation of the reticulocyte micronucleus assay in patients treated with radioiodine for thyroid cancer. Mutat.Res. 2005;583:12-25
13. Miller JL. A genome-based approach for the study of erythroid biology and disease. Blood Cells.Mol.Dis. 2004;32:341-343
14. Denis MM, Tolley ND, Bunting M. et al. Escaping the nuclear confines: signal-dependent pre-mRNA splicing in anucleate platelets. Cell. 2005;122:379-391
15. Gnatenko DV, Dunn JJ, McCorkle SR. et al. Transcript profiling of human platelets using microarray and serial analysis of gene expression. Blood. 2003;101:2285-2293
16. Weyrich AS, Lindemann S, Tolley ND. et al. Change in protein phenotype without a nucleus: translational control in platelets. Semin Thromb Hemost. 2004;30:491-498
17. Macaulay IC, Carr P, Gusnanto A. et al. Platelet genomics and proteomics in human health and disease. J Clin Invest. 2005;115:3370-3377
18. Kleinbongard P, Schulz R, Rassaf T. et al. Red blood cells express a functional endothelial nitric oxide synthase. Blood. 2006;107:2943-2951
19. Newman PJ, Gorski J, White GC2. et al. Enzymatic amplification of platelet-specific messenger RNA using the polymerase chain reaction. J Clin Invest. 1988;82:739-743
20. Waggoner SA, Liebhaber SA. Regulation of α-Globin mRNA Stability. Exp.Biol.Med. 2003;228:387-395
Author contact
Correspondence to: Thomas W. Jax, MD, PhD, Profil Institut für Stoffwechselforschung, Hellersbergstrasse 9, 41461 Neuss, Germany. thomas.jaxde. Tel: ++49 - 2131 - 4018 - 0; Fax. ++49 - 2131 - 4018 - 577

 Global reach, higher impact
Global reach, higher impact