Impact Factor ISSN: 1449-1907
Int J Med Sci 2013; 10(5):539-547. doi:10.7150/ijms.5516 This issue Cite
Research Paper
Potential Immunogenic Polypeptides of Burkholderia pseudomallei Identified by Shotgun Expression Library and Evaluation of Their Efficacy for Serodiagnosis of Melioidosis
1. Department of Biomedical Science, Faculty of Medicine, University of Malaya.
2. Department of Medical Education, Research & Evaluation, DUKE-NUS Graduate Medical School Singapore, 8 College Road, Singapore 169857.
Received 2012-11-10; Accepted 2013-3-6; Published 2013-3-13
Abstract
The search for novel immunogenic polypeptides to improve the accuracy and reliability of serologic diagnostic methods for Burkholderia pseudomallei infection is ongoing. We employed a rapid and efficient approach to identify such polypeptides with sera from melioidosis patients using a small insert genomic expression library created from clinically confirmed local virulent isolates of B. pseudomallei. After 2 rounds of immunoscreening, 6 sero-positive clones expressing immunogenic peptides were sequenced and their identities were: benzoate 1,2-dioxygenase beta subunit, a putative 200 kDa antigen p200, phosphotransferase enzyme family protein, short chain dehydrogenase and 2 hypothetical proteins. These immunogens were then transferred to an ELISA platform for further large scale screening. By combining shotgun expression library and ELISA assays, we identified 2 polypeptides BPSS1904 (benzoate 1,2-dioxygenase beta subunit) and BPSL3130 (hypothetical protein), which had sensitivities of 78.9% and 79.4% and specificities of 88.1% and 94.8%, respectively in ELISA test, thus suggesting that both are potential candidate antigens for the serodiagnosis of infections caused by B. pseudomallei.
Keywords: melioidosis, shotgun expression library, serodiagnosis, recombinant polypeptide, ELISA.
Introduction
Burkholderia pseudomallei, the causative agent of melioidosis, is a soil saprophyte endemic to Southeast Asia and Northern Australia where infection is acquired usually by inhalation, entry through cutaneous lesions and by ingestion [1-2]. This infection has the potential for prolonged latency with recrudescence into acute and fulminating fatal infections [3]. The fatality rate in acute septicemic meliodoisis can be as high as 65% [4].
Melioidosis is difficult to diagnose clinically due to the wide range of associated symptoms and the mortality is high among patients with underlying risk factors especially diabetes and immunosuppression [5]. Definitive diagnosis of melioidosis has historically been based on the isolation of the bacterium from clinical specimens using culture techniques. Bacterial culture may take up to 4 days and reportedly can have associated problems such as inter-strain and medium dependent variability in colony morphology. While the “gold standard” for diagnosis remains the isolation and identification of the causative bacterium, serological assays have also been proven as useful tools in the presumptive diagnosis of the disease. There are many situations where patients are critically ill with fulminating sepsis or when infections are deep seated and no specimens are available, where serology may be sufficiently rapid to facilitate aggressive and appropriate treatment and management of patients. Serology also has a particular role in subacute presentations and in the screening of patients with febrile illness in endemic areas as well as personnel who have returned from areas of endemicity. Moreover, specific serodiagnosis can also be used to monitor disease activity in patients under treatment by investigating the persistence of antibody levels [6].
Several rapid serological diagnostic tests have been developed but the performance characteristics of many of these have not been adequately evaluated. The indirect hemagglutination assay (IHA) is most commonly used in disease-endemic countries [7-9], but high background titres raise problems in differentiating past and current infections [10- 11]. In addition, the IHA has been reported to have low sensitivity in sera from patients with acute septicaemia [12-13].
In Malaysia, we have previously developed an indirect immunofluorescent test (IFAT) using a whole cell antigen derived from B. pseudomallei for detection of total antibodies (IgG and IgM) in patients with possible melioidosis. This IFAT test performed better than IHA and had satisfactory sensitivity and specificity compared to culture [10]. However, the drawback is preparation of B. pseudomallei as antigen will require biosafety laboratory level 3 (BSL-3) facilities, which are not commonly available in developing countries. Subsequently a modified IFAT was developed by using B. thailandensis as antigen and this resulted in good sensitivity of 98.8% and satisfactory specificity of 92.3% [14]. However, both IFAT assays using either B. pseudomallei or B. thailandensis are based on subjective scores, are labour-intensive and do not lend themselves to large scale investigations.
ELISA studies using either crude or purified antigen preparations for the detection of antibodies to B. pseudomallei demonstrated varying results in terms of sensitivities and specificities [15-18]. Another rapid and simple test, an immunochromatographic kit, using secretory antigens had been developed but inconsistent results were observed [7-8, 19-20]. Most recently, quantitative real time PCR targeting various specific regions such as TTS1-orf1, TTSS1-orf 2 and lpxO, [21-23] and loop-mediated isothermal DNA amplification [24] have been developed for identification of B. pseudomallei. However, these assays demonstrated only moderate sensitivities and direct testing on blood specimens remain problematic.
Hence, a serological test detecting antibody response to recombinant proteins of B. pseudomallei would be less complex and less expensive than molecular detection techniques, and more rapid than traditional bacterial culture. Numerous studies have been conducted to select relevant antigens for serodiagnosis using different approaches such as bioinformatic prediction tools and protein array [25-26]. The present study aimed to identify immunogenic polypeptides of B. pseudomallei recognized by the sera of patients with IFAT (serologically) proven melioidosis and to explore these immunogens for potential use in the serological diagnosis of melioidosis.
Materials and Methods
Bacterial strains and DNA extraction
Four strains of B. pseudomallei isolated from blood, swab, sputum and splenic biopsy, from patients at the University Hospital, University of Malaya (UM), Kuala Lumpur, were used for the preparation of genomic expression library. These strains had been isolated and identified by standard biochemical tests and the API 20NE system (bioMérieux, France). Genomic DNA of B. pseudomallei was extracted by using Bacterial Genomic DNA Isolation Kit (Norgen Biotek, Canada) as previously described [27].
Construction of genomic expression library
The genome expression library was constructed using the NovaTope System (Novagen, USA) according to the manufacturer's protocol. Ten micrograms of pooled DNA of the 4 clinical strains were randomly cleaved by DNase I in the presence of Mn2+ and fragments ranging in size from 50-200 bp were recovered from a 2% (w/v) agarose gel with QIAEX II Gel Extraction Kit (Qiagen, Germany). The purified DNA fragments were flush ended with T4 DNA polymerase, a single dA residue to each 3' end was then added by Tth polymerase, and ligated into pSCREEN T-vector, designed for the expression of small peptides as a carboxy-terminal fusion to the T7 bacteriophage gene 10 capsid protein. One microliter of the ligation mixture was transformed into 20 µl Escherichia coli NovaBlue (DE3) competent cells followed by selection of transformants by plating onto Luria-Bertani (LB) agar containing 50 µg/ml carbenicillin (CB) and 15 µg/ml tetracycline (Tet).
Colony screening by protein expression
Transformed colonies were picked with sterile toothpicks and streaked on fresh LB-agar containing CB and Tet and incubated overnight at 37°C. Bacterial transformants were then transferred onto nitrocellulose filters which were incubated with colony side up for 4 h at 37°C on LB-agar plates containing CB, Tet and 250 µM isopropyl-beta-thio galactopyronoside (IPTG) to induce the expression of insert targets. After induction, the filters were serially transferred to petri dishes containing 10% sodium dodecyl sulphate (SDS) for 10 minutes to disrupt cell membranes; denaturing solution for 5 minutes; neutralization solution for 10 minutes and lastly 2x saline-sodium citrate (SSC) for 15 minutes for lysis of bacteria and binding of protein to the filters. The filters were then washed in TBS [10 mM Tris-HCl pH 8.0, 150 mM NaCl] for 10 minutes before immersion in blocking solution (TBST + 1% gelatin) for 1 h with gentle agitation. Filters were washed again twice in TBST [10 mM Tris-l pH 8.0, 150 mM NaCl] 0.05% Tween-20] for 10 minutes to remove bacterial debris and then submerged in 2 different pools of sera, i.e., pooled melioidosis positive and melioidosis negative (1:1000 in blocking buffer) for 1 h. Next, filters were washed in TBST and incubated with AP-conjugated Immunopure® Protein A/G (1:10000 in blocking buffer) (Pierce). Subsequently, immunoreactive clones were detected with BCIP/NBT phosphatase substrate (Pierce). All sera used for immunoscreening were pre-absorbed with E. coli NovaBlue (DE3) cell lysates to reduce non-specific background signals. Mean signal intensity and background measurements were obtained for each clone on each filter using Image Scanner III (GE Healthcare), and analysed using ImageQuant software (GE Healthcare). The dataset for each filter was normalized using the signals of pSCREEN T-vector without insert and the ratio of signal intensities produced by melioidosis positive and negative groups was calculated. Reactive clones were recovered from the master plates and frozen as glycerol stocks.
Plasmid preparation and sequencing
Plasmids isolated from immunoreactive clones were extracted with the HiYieldTM Plasmid Mini Kit (RBC, Taiwan) and sent for sequencing (1st Base Laboratories, Malaysia) using the T7 gene 10 primer and T7 terminator primer. Protein similarity analysis of the acquired sequences was performed using BlastX based on NCBI non-redundant protein database (B. pseudomallei strains K96243, 1710, 1106, MSHR668, and MSHR305).
Preparation of purified recombinant polypeptide
Transformed colonies were chosen and cultured overnight at 37°C in 2x YT broth containing CB and Tet. The broth culture was diluted 1/20 with fresh medium and incubated at 37°C with shaking at 200rpm until the OD600 had reached the range from 0.5 to 0.7. The culture was incubated for an additional 4 h after induction with the addition of IPTG. The bacterial cells were harvested by centrifugation and resuspended in ice-cold 1 x TES buffer [0.2 M Tris-HCl, 0.5 mM EDTA, 0.5 M sucrose]. A buffer of 1/5 x TES was added and the mixture incubated on ice for 30 minutes with agitation. The supernatant containing the soluble recombinant proteins was collected by centrifugation and stored at -20°C for further use. The concentration of recombinant polypeptides was determined by the Bradford assay.
SDS-PAGE and Western blot
The recombinant polypeptide samples were separated by electrophoresis on 12% sodium dodecyl sulphate-polyacrylamide gel (SDS-PAGE) at 170 V for 1 hr. The gel was transferred to a nitrocellulose membrane using TE 70 Semi-Dry Transfer Unit (GE). Blocking, incubating and washing steps were performed as described in the section on colony screening. The presence of recombinant polypeptides specific to B. pseudomallei were detected using pooled melioidosis positive and melioidosis negative sera. The pSCREEN T with no insert was the negative control.
Serum samples
The serum samples were from suspected cases of melioidosis, sent to the University Malaya Medical Centre from 2010 to 2012 and serologically confirmed by the in-house IFAT test (10). A panel of 183 (60 melioidosis positive and 123 non-melioidosis sera i.e. IFAT negative) were divided into 8 groups as follows: Group A, sera from patients clinically and serologically (IFAT+ve) confirmed as melioidosis (n=60); Group B, sera from individuals negative (IFAT-ve) for melioidosis (n=60); Group C, healthy blood donors (n=10); Groups D to H, sera from patients who were diagnosed with Legionella pneumoniae (n=11); Leptospirosis (n=13); Chlamydophila pneumoniae (n=10); Mycoplasma pneumoniae (n=12) and Rickettsiosis (typhus group) (n=7).
Evaluation of purified recombinant polypeptides
Large scale screening using an indirect ELISA was carried out to determine the immunoreactivity of clones detected in the colony immunoblot assay. Antigens (i.e. recombinant polypeptides) of dilution range 100 ng/µL to 20 µg/mL and IFAT confirmed melioidosis positive and negative sera of 1:100 to 1:3,000 dilutions were optimized and the optimal concentrations were chosen for the subsequent ELISA assay, i.e., 20 µg/mL for each antigen and 1:500 dilutions for sera. Ninety-six well microtiter plates (Costar, Corning) were coated and incubated overnight at 4°C, with 100 µL/well of antigens in 1x PBS (pH 8.4). The plates were then washed 3 times with PBS containing 0.05% Tween 20 (1x PBST) to remove unbound antigen and blocked with 100 µL of 1% gelatin in PBST for 2 h at 37°C. The plates were again washed using 1x PBST and the wells incubated with 100 µL of sera at 37°C for 2 h followed by another washing step and addition of AP-conjugated Immunopure® Protein A/G ( 1:5,000 in blocking buffer) (Pierce) at 37°C for 2 h. A final washing step was performed before addition of 100 µL of p-Nitrophenyl Phosphate (pNPP). The plates were incubated in the dark at 37°C for 30 minutes and 100 µL of 2N H2SO4 added to stop the reaction. Mean optical density (OD) at 415 nm was determined using an iMark reader (Biorad). Each sample was run in duplicate wells and results determined by the mean value of the OD.
Statistical analyses
The cutoff for ELISA was determined by using the mean OD for the IFAT confirmed melioidosis negative group (Group B) plus 3 standard deviations (M + 3SD). The sensitivity, specificity, positive and negative predictive values were calculated.
Results
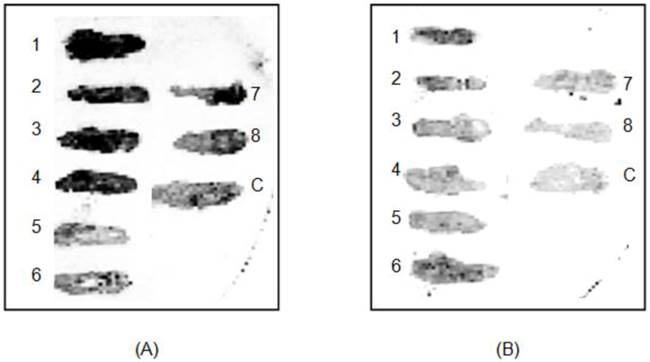
A small diversely sized genomic expression library was successfully constructed using local clinical strains of B. pseudomallei carrying random fragmented inserts of 50-200 bp. From the resulting library, 480 transformants were taken at random and screened with pooled positive (P: IFAT confirmed positive) and pooled negative sera (N: IFAT confirmed negative) (Fig 1). Each pool consisted of 10 samples, and 4 sets of pooled sera were prepared, i.e., Pos1, Neg1, Pos2 and Neg2. The first screening step was an attempt to identify a broad panel of targets, and immunoreactive clones with signal intensity ratios (P/N) of two-fold and above using sera Pos1 and Neg1, resulted in the selection of 79 clones. These reactive clones were isolated and retested for reactivity with another set of pooled sera (Pos2 and Neg2) and finally the panel was narrowed down to 20 clones with signal ratio intensities (P/N) of at least 3-fold and above. Sequencing of these inserts revealed 6 unique clones: a benzoate 1,2-dioxygenase beta subunit, a putative 200 kDa antigen p200, phosphotransferase enzyme family protein, short chain dehydrogenase and 2 hypothetical proteins (Table 1).
Representative clones (1-8) immunoscreened with pooled melioidosis positive sera (A); melioidosis negative (B); control strain, E. coli NovaBlue (DE3) containing the pSCREEN T-vector with no insert (C).

Characteristics of the 6 recombinant polypeptides of B. pseudomallei.
| Locus tag | Putative identity | Predicted functiona | Cellular | Accession | Amino acid sequence | ||
|---|---|---|---|---|---|---|---|
| location a | no. | Identity (%) | Coverage (%) | E-value | |||
| BPSS1904 | benzoate 1,2-dioxygenase beta subunit | Secondary metabolites biosynthesis, transport, and catabolism | C | YP_111910 | 100 | 39.7 | 8.0E-5 |
| BPSL3130 | hypothetical protein | Function unknown | CM | YP_109723 | 100 | 33.9 | 0.00 |
| BURPS1710b_0454 | 200 kDa antigen p200, putative | Cell envelope biogenesis, outer membrane / carbohydrate transport and metabolism | C | YP_331868 | 98 | 98 | 8E-31 |
| BPSS1856 | phosphotransferase enzyme family protein | Function unknown | C | YP_111860 | 100 | 99 | 3E-17 |
| BPSS0897 | short chain dehydrogenase | Secondary metabolites biosynthesis, transport, and catabolism | C | YP_ 110906 | 98 | 99 | 3E-21 |
| BPSS1757 | hypothetical protein | Function unknown | C | YP_ 105144 | 100 | 98 | 3E-27 |
a As predicted by PSORTb V3.0 .Cytoplasm (C), cytoplasmic membrane (CM).
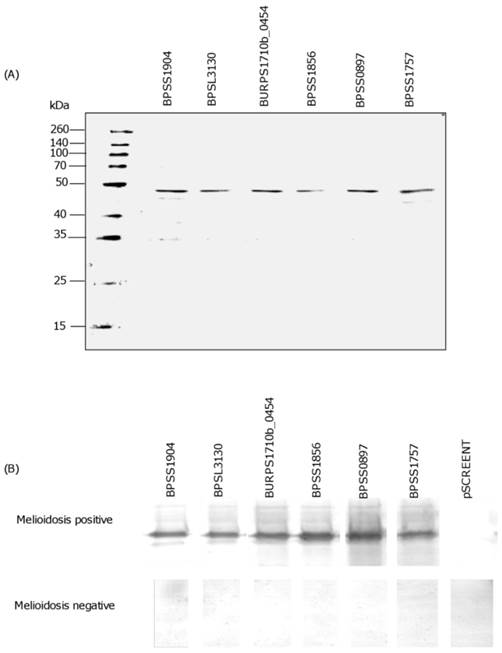
Each recombinant polypeptide was found to have a predicted molecular weight in the range 46 to 48 kDa (Fig 2A). Western blot analysis with patients' sera further confirmed the expression of each recombinant immunogenic polypeptide. A clear and strong signal for positive reactivity using melioidosis positive sera when compared to the negative sera (Fig 2B), indicated that the candidate polypeptides identified by the immunoscreening were indeed immunogenic and specifically expressed during human infection. The negative control, pSCREEN T with no insert did not react with either melioidosis positive or negative sera (Fig 2B).
SDS-PAGE and Western blot of candidate polypeptide antigens. SDS-PAGE showing purified recombinant polypeptides (~46-48 kDa) (A). Western blot imunodetection using melioidosis positive and negative serum samples. Note absence of band in both sera using negative control (pSCREEN T) (B).
Validation of the 6 recombinant polypeptides was carried out by using serum samples from Groups A-H in an alternative analytical platform, i.e. indirect ELISA. The results of the 6 ELISA tests were analysed using the in-house IFAT as a reference test (sensitivity and specificity equal to 100%), and cutoff point was established as the mean A415 + 3SD of the IFAT confirmed melioidosis negative sera. The cutoff values were 0.123, 0.133, 0.145, 0.136, 0.198 and 0.122 for BPSS1904, BPSL3130, BURPS1710b_0454, BPSS1856, BPSS0897 and BPSS1757, respectively. These values allowed us to discriminate between positive and negative sera and the percentages of positive sera were determined (Table 2), followed by estimation of the performance of the ELISAs (Table 3). The overall performance of the 6 ELISA tests were moderate to satisfactory, with sensitivities and specificities ranging from 75% to 94.8%. It was evident that BPSS1904 and BPSL3130 recognized a large proportion of the melioidosis positive sera (54/60) and (45/60) but there were also cross reactions with some of the disease control sera as shown in Table 2. Thus among the 6 recombinant polypeptides, BPSS1904 and BPSL3130 have shown to be superior as both had sensitivities of 78.9% and 79.4%, specificities of 88.1% and 94.8%, giving a positive predictive value of approximately 75% and 90%, and a negative predictive value of 90.2% and 88.6% (Table 3).
ELISA screening for melioidosis and disease control sera using the 6 recombinant polypeptides.
| Disease group | Percentage positivity | |||||
|---|---|---|---|---|---|---|
| BPSS1904 | BPSL3130 | BURPS1710b_0454 | BPSS1856 | BPSS0897 | BPSS1757 | |
| A Melioidosis positive (n=60) | 75.0 (45/60) | 90.0 (54/60) | 66.7 (40/60) | 66.7 (40/60) | 60.0 (36/60) | 33.3 (20/60) |
| B Melioidosis negative (n=60) | 1.7 (1/60) | 1.7 (1/60) | 0/60 | 1.7 (1/60) | 0/60 | 3.3 (2/60) |
| C Healthy control (n=10) | 0/60 | 0/60 | 0/60 | 0/60 | 0/60 | 0/60 |
| D Legionella pneumoniae (n=11) | 36.4 (4/11) | 36.4 (4/11) | 27.3 (3/11) | 36.4 (4/11) | 27.3 (3/11) | 0/11 |
| E Leptospirosis (n=13) | 15.4 (2/13) | 7.7 (1/13) | 7.7 (1/13) | 15.4 (2/13) | 7.7 (1/13) | 0/13 |
| F Chlamydophila pneumoniae (n=10) | 20.0 (2/10) | 20.0 (2/10) | 0/10 | 20.0 (2/10) | 0/10 | 10.0 (1/10) |
| G Mycoplasma pneumoniae (n=12) | 16.7 (2/12) | 33.3 (4/12) | 25.0 (3/12) | 25.0 (3/12) | 25.0 (3/12) | 0/12 |
| H Rickettsiosis (typhus group) (n=7) | 14.3 (1/7) | 28.6 (2/7) | 0/7 | 0/7 | 0/7 | 0/7 |
Sensitivity, specificity, positive and negative predictive values of the 6 B. pseudomallei recombinant polypeptides.
| Percentage | BPSS1904 | BPSL3130 | BURPS1710b_0454 | BPSS1856 | BPSS0897 | BPSS1757 |
|---|---|---|---|---|---|---|
| Sensitivity | 78.9 | 79.4 | 85.1 | 76.9 | 83.7 | 87 |
| (65.8-88.2) | (67.5-87.9) | (71-93.3) | (62.8-87) | (68.7-92.7) | (65.3-96.6) | |
| Specificity | 88.1 | 94.8 | 85.3 | 84.7 | 82.9 | 75 |
| (80.8-93.0) | (88.5-97.9) | (78-90.6) | (77.1-90.2) | (75.4-88.5) | (67.4-81.3) | |
| Positive predictive value | 75 | 90 | 66.7 | 66.7 | 60 | 33.3 |
| (61.9-84.9) | (78.8-95.9) | (53.2-78) | (53.2-78) | (46.5-72.2) | (22-46.8) | |
| Negative predictive value | 90.2 | 88.6 | 94.3 | 90.2 | 94.3 | 97.5 |
| (83.2-94.6) | (81.3-93.4) | (88.2-97.5) | (83.2-94.6) | (88.2-97.5) | (92.5-99.4) |
( ): Confidence interval 95%.
Discussion
The absence of a well-established universally acceptable antigen has made the serologic diagnosis of melioidosis difficult and controversial. It is of paramount importance that the ideal antigen should consist of all or most of the relevant immunogens that are recognized by patients infected with B. pseudomallei. Although several immunogenic proteins in B. pseudomallei have been identified such as GroEL [28], OmpA [29] and Elongation factor-Tu [30], these immunogens have not been further evaluated as potential serodiagnostic markers.
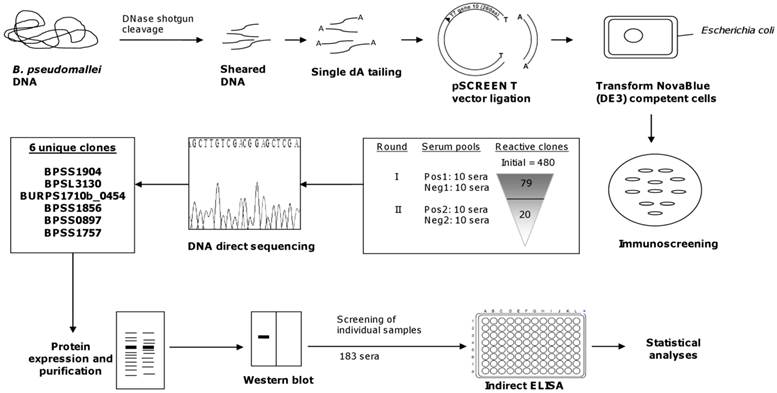
With the availability of the complete genome sequences of B. pseudomallei strains K96243, 1710b, 1106a, MSHR668, and MSHR305, we searched for potential immunogens among all genome-encoded proteins [31-32]. The effectiveness of the shotgun expression library with an average insert size of 50 to 200 bp expressing partial ORF which we used, permitted the identification of immunogenic polypeptides that reacted with pooled sera of melioidosis patients (Fig 3). This approach had been used successfully in the identification of diagnostically useful immunodominant polypeptides of M. tuberculosis [33].
Diagrammatic representation of shotgun approach for B. pseudomallei.
The pooling of sera from melioidosis patients has enabled us to compensate for variations in the immune responses of individual patients. A stepwise selection process identified 6 immunogenic polypeptides, i.e., BPSS1904, BPSL3130, BURPS1710b_0454, BPSS1856, BPSS0897 and BPSS1757. These polypeptides are involved in secondary metabolite biosynthesis, cell envelope biogenesis as well as hypothetical proteins of unknown function(s). Protein topology analysis of these proteins revealed that the majority of the identified immunogens involved in human infection were cytoplasmic localized proteins (Table 1) which is in accordance with a previous published report [34].
These 6 immunogenic polypeptides described above may possibly be potential virulence factors that mediate important host-pathogen interactions. Interestingly, 2 immunogens (BPSS1904 and BPSS0897) that have been previously identified to be involved in bacterial adaptation processes and BURPS1710b_0454 is predicted to regulate cell envelope biogenesis. BPSS1904 was reported to play an essential role in the survival of B. pseudomallei during the early stationary phase of growth [35] whereas BPSS0897 was coexpressed with T3SS-associated genes in response to salt stress [36]. However, the relative importance of these remain largely unknown, and therefore it would be valuable to further investigate in vivo or in vitro models that could give insights into B. pseudomallei virulence.
It is noteworthy that 3 immunogens BPSL3130, with unknown functions were detected: both BPSS1856 and BPSS1757 proteins have PKc-like and gatB-Yqey related domains based on conserved domain database [37]. BPSS1856 has homology to Streptomyces coelicolor putative phosphotransferase and may be involved in carbon metabolism [38]. These 3 polypeptides shown to be immunogenic during immunoscreening deserve further investigation as diagnostic candidates. To the best of our knowledge, none of these 6 immunogenic polypeptides have been evaluated as antigens for serodiagnosis of melioidosis.
Although the immunoscreening approach is rapid, the cocktail of recombinant polypeptides may have contained contaminants with low impurities derived from E. coli NovaBlue (DE3) or pSCREEN T vector and could have influenced the specificity of the assay. Indeed, the colony immunoblot assay is cumbersome and poorly standardised. For these reasons, the 6 recombinant polypeptides were purified and transferred to the ELISA format for further evaluation using individual samples.
Recent studies investigating the potential of ELISAs based on detection of antibody to various antigen preparations of B. pseudomallei have provided differing results with poor, moderate and good sensitivities and specificities [16-18]. In the present study, the highest diagnostic indices of ELISA were observed for BPSS1904 (sensitivity, 78.9%; specificity, 88.1%) and BPSL3130 (sensitivity, 79.4%; specificity, 94.8%), both also gave the best results in terms of positive and negative predictive values. These results compare favourably with the sensitivities and specificities of previous studies using an affinity-purified antigen (sensitivity, 82%; specificity, 72%) and a crude B. pseudomallei antigen (sensitivity, 81%; specificity, 70%) [39].
Overall, BPSS1904 and BPSL3130 have shown good potential as candidate antigens for serodiagnosis although the sensitivities were only moderate. Therefore we suggest using these 2 recombinant immunogenic polypeptides or synthetic peptides as a combination antigen to further enhance the sensitivity of antibody detection. Other investigators have shown that using 2 purified proteins (EPS plus LPS) demonstrated increased sensitivity compared to results when the antigens were used separately [39]. Cross-reactivity with disease-control sera, have been reported previously in other studies [14, 19]. A possible explanation for this observation is that immunodominant epitopes might be shared with other microorganisms or indicative of past exposure to bacteria related to B. pseudomallei.
In summary, the shotgun expression library approach was shown to be useful, whereby we have identified 6 immunogens that were able to react specifically with sera from melioidosis patients. Two of these recombinant polypeptides in combination, would probably be the superior potential candidate antigen for serodiagnostic purposes based on their ELISA performance and further field testing would be desirable.
Acknowledgements
This study was supported by University of Malaya Research Grant RG409/12HTM and Higher Education Malaysia High Impact Research grant (UM/MoHE HIR grant E000044-20001).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Cheng AC, Currie BJ. Melioidosis: epidemiology, pathophysiology, and management. Clin Microbiol Rev. 2005;18:383-416
2. Galyov EE, Brett PJ, DeShazer D. Molecular insights into Burkholderia pseudomallei and Burkholderia mallei pathogenesis. Annu Rev Microbiol. 2010;64:495-517
3. Puthucheary SD. Melioidosis in Malaysia. Med J Malaysia. 2009;64:266-274
4. Puthucheary SD, Parasakthi N, Lee MK. Septicaemic melioidosis: a review of 50 cases from Malaysia. Trans R Soc Trop Med Hyg. 1992;86:683-685
5. Puthucheary SD, Vadivelu J. Human Melioidosis. Singapore: Singapore University Press. 2002
6. Vadivelu J, Puthucheary SD. Diagnostic and prognostic value of an immunofluorescent assay for melioidosis. Am J Trop Med Hyg. 2000;62:297-300
7. O'Brien M, Freeman K, Lum G, Cheng AC, Jacups SP, Currie BJ. Further evaluation of a rapid diagnostic test for melioidosis in an area of endemicity. J Clin Microbiol. 2004;42:2239-2240
8. Chuah SC, Gilmore G, Norton RE. Rapid serological diagnosis of melioidosis: an evaluation of a prototype immunochromatographic test. Pathology. 2005;37:169-171
9. Cheng AC, O'Brien M, Freeman K, Lum G, Currie BJ. Indirect hemagglutination assay in patients with melioidosis in northern Australia. Am J Trop Med Hyg. 2006;74:330-334
10. Vadivelu J, Puthucheary SD, Gendeh GS, Parasakthi N. Serodiagnosis of melioidosis in Malaysia. Singapore Med J. 1995;36:299-302
11. Wuthiekanun V, Chierakul W, Langa S, Chaowagul W, Panpitpat C, Saipan P, Thoujaikong T. et al. Development of antibodies to Burkholderia pseudomallei during childhood in melioidosis-endemic northeast Thailand. Am J Trop Med Hyg. 2006;74:1074-1075
12. Appassakij H, Silpapojakul KR, Wansit R, Pornpatkul M. Diagnostic value of the indirect hemagglutination test for melioidosis in an endemic area. Am J Trop Med Hyg. 1990;42:248-253
13. Dharakul T, Songsivilai S, Anuntagool N, Chaowagul W, Wongbunnate S, Intachote P, Sirisinha S. Diagnostic value of an antibody enzyme-linked immunosorbent assay using affinity-purified antigen in an area endemic for melioidoisis. Am J Trop Med Hyg. 1997;56:418-423
14. Puthucheary SD, Anuar AS, Tee TS. Burkholderia thailandensis whole cell antigen cross-reacts with B. pseudomallei antibodies from patients with melioidosis in an immunofluorescent assay. Southeast Asian J Trop Med Public Health. 2010;41:395-400
15. Sirisinha S, Anuntagool N, Dharakul T, Ekpo P, Wongratanacheewin S, Naigowit P, Petchclai B. et al. Recent developments in laboratory diagnosis of melioidosis. Acta Trop. 2000;74:234-235
16. Chenthamarakshan V, Kumutha MV, Vadivelu J, Puthucheary SD. Distribution of immunoglobulin classes and IgG subclasses against a culture filtrate antigen of Burkholderia pseudomallei in melioidosis patients. J Med Microbiol. 2001;50:55-61
17. Allwood EM, Logue CA, Hafner GJ, Ketheesan N, Norton RE, Peak IR, Beacham IR. Evaluation of recombinant antigens for diagnosis of melioidosis. FEMS Immunol Med Microbiol. 2008;54:144-153
18. Druar C, Yu F, Barnes JL, Okinaka RT, Chantratita N, Beg S, Stratilo CW. et al. Evaluating Burkholderia pseudomallei Bip proteins as vaccine and Bip antibodies as detection agents. FEMS Immunol Med Microbiol. 2008;52:78-87
19. Cuzzubbo AJ, Chenthamarakshan V, Vadivelu J, Puthucheary SD, Rowland D, Devine PL. Evaluation of a new commercially available immunogloblulin M and immunoglobulin G immunochromatographic test for diagnosis of melioidosis infection. J Clin Microbiol. 2000;38:1670-1671
20. Cheng AC, Peacock SJ, Limmathurotsakul D, Wongsuvan G, Chierakul W, Amornchai P, Getchalarat N. et al. Prospective evaluation of a rapid immunochromogenic cassette test for the diagnosis of melioidosis in northeast Thailand. Trans R Soc Trop Med Hyg. 2006;100:64-67
21. Meumann EM, Novak RT, Gal D, Kaestli ME, Mayo M, Hanson JP, Spencer E. et al. Clinical evaluation of a type III secretion system real-time PCR assay for diagnosis melioidosis. J Clin Microbiol. 2006;44:3028-3030
22. Supaprom C, Wang D, Leelayuwat C, Thaewpia W, Susaengrat W, Koh V, Ooi EE. et al. Development of real-time PCR assays and evaluation of their potential use for rapid detection of Burkholderia pseudomallei in clinical blood specimens. J Clin Microbiol. 2007;45:2894-2901
23. Kaestli M, Richardson LJ, Colman RE, Tuanyok A, Price A, Price EP, Browers JR. et al. Comparison of Taqman PCR assays for detection of the, melioidosis agent Burkholderia pseudomallei in clinical specimens. J Clin Microbiol. 2012;50:2059-2062
24. Chantratita N, Meumann E, Thanwisai A, Limmathurotsakul D, Wuthiekanun V, Wannapasni S, Tumapa S. et al. Loop-mediated isothermal amplification method targeting the TTS1 gene cluster for detection of Burkholderia pseudomallei and diagnosis of melioidosis. J Clin Microbiol. 2008;46:568-573
25. Thompson DB, Crandall K, Harding SV, Smither SJ, Kitto GB, Titball RW, Brown KA. In silico analysis of potential diagnostic targets from Burkholderia pseudomallei. Trans R Soc Trop Med Hyg. 2008;102:S61-S65
26. Felgner PL, Kayala MA, Vigil A, Burk C, Nakajima-Sasaki R, Pablo J, Molina DM. et al. A Burkholderia pseudomallei protein microarray reveals serodiagnostic and cross-reactive antigens. Prot Natl Acad Sci USA. 2009;106:13499-13504
27. Puthucheary SD, Puah SM, Chua KH. Molecular characterization of clinical isolates of Aeromonas species from Malaysia. PLoS ONE. 2012;7:e30205
28. Woo PC, Leung PK, Wong SS, Ho PL, Yuen KY. groEL encodes a highly antigenic protein in Burkholderia pseudomallei. Clin Diagn Lab Immunol. 2001;8:832-836
29. Hara Y, Mohamed R, Nathan S. Immunogenic Burkholderia pseudomallei outer membrane protein as potential candidate vaccine targets. PLoS One. 2009;4:e6496
30. Nieves W, Heang J, Asakrah S, Höner zu Bentrup K, Roy CJ, Morici LA. Immunospecific responses to bacterial elongation factor Tu during Burkholderia infection and immunization. PLoS One. 2010;5:e14361
31. Holden MT, Titball RW, Peacock SJ, Cerdeño-Tárraga AM, Atkins T, Crossman LC, Pitt T. et al. Genomic plasticity of the causative agent of melioidosis, Burkholderia pseudomallei. Proc Natl Acad Sci USA. 2004;28:14240-14245
32. Tuanyok A, Leadem BR, Auerbach RK, Beckstron-Stemberg SM, Beckstrom-Stemberg JS, Mayo M, Wuthiekanun V. et al. Genomic islands from five strains of Burkholderia pseudomallei. BMC Genomics. 2008;27:566
33. Bisen PS, Garo SK, Tiwari RP, Tagore PR, Chandra R, Karbik R, Thaker N. et al. Analysis of the shotgun expression library of the Mycobacterium tuberculosis genome for immunodominant polypeptides: potential use in serodiagnosis. Clin Diagn Lab Immunol. 2003;10:1051-1058
34. Su YC, Wan KL, Mohamed R, Nathan S. A genome level survey of Burkholderia pseudomallei immunome expressed during human infection. Microbes Infect. 2008;10:1335-1345
35. Wongtrakoongate P, Roytrakul S, Yasothornsrikul S, Tungpradabkul S. A proteome reference map of the causative agent of melioidosis Burkholderia pseudomallei. J Biomed Biotechnol. 2011;2011:530926
36. Pumirat P, Cuccui J, Stabler RA, Stevens JM, Muangsombut V, Singsuksawat E, Stevens MP. et al. Global transcriptional profiling of Burkholderia pseudomallei under salt stress reveals differential effects on the Bsa type III secretion system. BMC Microbiol. 2010;10:171
37. Marchler-Bauer A, Lu S, Anderson JB, Chitsaz F, Derbyshire MK, DeWeese-Scott C, Fong JH. et al. CDD: a Conserved Domain Database for the functional annotation of proteins. Nucleic Acid Res. 2011;39:D225-229
38. Kamionka A, Parche S, Nothaft H, Siepelmeyer J, Jahreis K, Titgemeyer F. The phosphotransferase system of Streptomyces coelicolor. Eur J Biochem. 2002;269:2143-2150
39. Chantratita N, Wuthiekanun V, Thanwisai A, Limmathurotsakul D, Cheng AC, Chierakul W, Day NP. et al. Accuracy of enzyme-linked immunosorbent assay using crude and purified antigens for serodiagnosis of melioidosis. Clin Vaccine Immunol. 2007;14:110-113
Author contact
Corresponding author: Kek Heng Chua, Department of Biomedical Science, Faculty of Medicine, University of Malaya, 50603 Kuala Lumpur, Malaysia. Tel: 603-79676607 Fax: 603-79676600 E-mail: khchuaedu.my.

 Global reach, higher impact
Global reach, higher impact