Impact Factor ISSN: 1449-1907
Int J Med Sci 2017; 14(10):937-942. doi:10.7150/ijms.19565 This issue Cite
Research Paper
miR-224 Controls Human Colorectal Cancer Cell Line HCT116 Proliferation by Targeting Smad4
1. Department of General Surgery, Tongji Hospital, Tongji University School of Medicine, Shanghai, P.R. China
2. Division of Gastroenterology and Hepatology, Digestive Disease Institute, Tongji Hospital, Tongji University School of Medicine, Shanghai, P.R. China
* These three authors contributed equally to this work.
Received 2017-2-8; Accepted 2017-5-17; Published 2017-8-8
Abstract
Background: Better understanding the molecular mechanisms responsible for the genesis and progression of colorectal cancer would help advance the novel therapeutics. miR-224 has been identified to be elevated in colorectal cancer and promote human colorectal cancer cell line SW480 proliferation and invasion. However, the effect of miRNAs on cancer cell proliferation could be significantly changeable among different cell lines. HCT116 is a commonly used cell line for colorectal cancer study and the target gene responsible for the function of miR-224 in its proliferation is unclear.
Methods: miR-224 expression was determined by quantitative reverse transcription polymerase chain reactions (PCRs) in human colorectal cancer tissues compared with their corresponding matched peritumoral tissues. HCT116 cell viability and cell proliferation were determined by CCK-8, EdU incorporation assays and flow cytometry for cell cycle. Target gene of miR-224 was confirmed by Western blots and siRNA for Smad4.
Results: miR-224 was significantly increased by 29.49 fold in colorectal cancer tissues compared with their corresponding matched peritumoral tissues based on 12 colorectal cancer patients. miR-224 mimic significantly increased HCT116 cell viability, EdU positive cells rate, and decreased G1 phase cell population and increased S phase cell population. miR-224 inhibitor had opposite effects. Smad4 could be negatively regulated by miR-224 in HCT116 cells and was responsible for its effects in proliferation.
Conclusion: miR-224 mediates HCT116 cell proliferation by targeting Smad4.
Keywords: miR-224, human Colorectal Cancer Cell Line, HCT116, proliferation, Smad4.
Introduction
Colorectal cancer represents the third most common malignancy in men and the second one in women worldwide [1]. The incidence of colorectal cancer is steadily on the rise due to the lifestyle change and increasing aging population. Although several therapeutics for colorectal cancer have been developed, treatments for metastatic late-stage colorectal cancer are still limited. Palliative operation has always been considered as a main method [2]. Better understanding the molecular mechanisms responsible for the genesis and progression of colorectal cancer would help advance novel therapeutics [2].
MicroRNAs (miRNAs, miRs), 19-22 nucleotides non-coding single-stranded small molecule-RNA, can post-transcriptional regulate gene expression via control of translation or mRNA degradation followed by repression of protein synthesis [3-5]. Being the central regulators of gene expression, miRNAs have been found to play roles in regulating many essential biological processes including cell proliferation, apoptosis and migration [6-8]. Deregulated miRNAs have been linked to many diseases including cancer [9, 10]. Several miRNAs have been found to participate in the pathogenesis of colorectal cancer including miR-21, miR-451, miR-499-5p, miR-375, and miR-142-5p [9]. Besides that, miR-224 has also been identified to be elevated in colorectal cancer and promote human colorectal cancer cell line SW480 proliferation and invasion [11]. However, the effect of miRNAs on cancer cell proliferation could be significantly changeable among different cell lines [12]. HCT116 is a commonly used cell line for colorectal cancer study and the target gene responsible for the function of miR-224 in its proliferation is unclear.
Thus, in this study, we aim at determining the role of miR-224 in controlling the proliferation of a human colorectal cancer cell line HCT116. Our data demonstrate that miR-224 is significantly increased in colorectal cancer tissues as compared to the matched peritumoral tissues. miR-224 is able to control HCT116 cells proliferation and Smad4 is a target gene mediate the pro-proliferation effects of miR-224 in HCT116 cells.
Materials and Methods
Patient samples
Ethical approval for tissue collection protocol was obtained from the research ethics committee of Shanghai Tongji Hospital. All patients' written approval consent forms were required to sign before surgery. Colorectal cancer tissues and the matched peritumoral tissues were collected, at surgery and freshly frozen in liquid nitrogen, and stored at -80°C until future use.
Cell line culture
Human colorectal cancer cell line HCT116 cells were purchased from Keygen biotech Co., Ltd. and cultured in Dulbecco's modified eagle's medium (DMEM, Corning, USA) supplemented with 10% fetal bovine serum (Biolnd, Israel) and 1% penicillin streptomycin (Invitrogen), and were incubated at 37°C with 5% CO2.
miRNA and siRNA transfection
miR-224 mimic, inhibitor, and their corresponding negative controls, as well as Smad4 siRNA were all purchased from RiboBio (Guangzhou, China), Smad4-siRNA sense, 5′-GAGAAGTTCTCAAAGTTAA(dTdT)-3′; Smad4-siRNA antisense, 5′-TTAACTTTGAGAACTTCTC(dTdT) -3′. Before transfection, HCT116 cells were starved for 8 h. According to the manufacturer's instruction, Lipofectamine 2000 (Invitrogen, USA) was used to transfected miR-224 mimic (50 nM), inhibitor (100 nM), their respective negative controls and Smad4-siRNA (100 nM) into HCT116 cells for 48 h.
Cell counting kit-8 assay
Cell counting kit-8 assay (CCK-8; Dojindo, Japan) was used to determine the effects of miR-224 mimic and inhibitor on HCT116 cells viability. HCT116 cells were seeded in 96-well plates at a density of 0.2 million cells/mL and cultured in DMEM containing 10% FBS for 24 h. After starvation for 6 h with serum-free medium, cells were transfected with miR-224 mimic or inhibitor for 48h (co-incubation with CCK-8 solution 1h before the end of the transfection) in a cell culture incubator at 37 °C. The absorbance at a wavelength of 450 nm was measured using a spectrophotometer (Bio-Rad, USA).
EdU staining
Cellular proliferation rate was measured by 5-ethynyl-2'-deoxyuridine (EdU) assays. HCT116 cells were plated at 0.2 million cells/mL per well into 96-well plates, and adhered overnight. After transfection, the cells were incubated with 100 μl per well EdU for 2 hours, fixed with 4% paraformaldehyde for 30 minutes, and stained with Cell-Light™ EdU Apollo®488 In Vitro Imaging Kit (RioBio, China) according to the manufacturer's instructions.
Flow Cytometry detection of cell cycle
Cell cycle was measured using flow cytometric measurement. HCT116 cells were plated onto 12-well plates for adhere. After starvation overnight and transfected with miR-224 mimic or inhibitor for 48h, cells were harvested and fixed with pre-cooling absolute ethanol overnight at -20°C. Subsequently cells were washed with PBS, treated with RNAse (KeyGEN BioTECH) and stained with propidium iodide (PI, Sigma) solution (50µg/mL). Cell cycle was analyzed using flow cytometry (Beckman, USA).
RNA extraction and quantitative reverse transcription polymerase chain reactions
Total RNA was extracted from the collected tissues or HCT116 cells using Trizol Reagent (Invitrogen, USA) according to the manufacturer's instructions. cDNA synthesis was performed using the iScriptTM cDNA synthesis kit (Bio-Rad, USA). Then cDNA was subjected into 40 cycles of quantitative reverse transcription polymerase chain reactions (PCRs) with Takara SYBR Premix Ex TaqTM (Takara, Japan) in a CFX96TM RealTime PCR Detection System (Bio-Rad, USA). The 5S and GAPDH were used as the internal control for miR-224 and Smad4, respectively. The primer sequences were as follows. Smad4: Forward, CTCATGTGATCTATGCCCGTC; Reverse, AGGTGATACAACTCGTTCGTAGT. GAPDH: Forward, ATGACATCAAGAAGGTGGTG; Reverse, CATACCAGGAAATGAGCTTG.
Western blot analysis
Tissue or HCT116 cellular proteins were extracted used Radio-Immunoprecipitation Assay (RIPA) buffer (KeyGen, China) containing 1% phenylmethanesulfonyl fluoride (PMSF). Protein of 30 µg was added to standard 10% SDS-PAGE gel and transferred onto polyvinylidene difluoride (PVDF) membranes. After blocking, membranes were incubated with anti-Smad4 antibody (Bioworld, USA) overnight at 4°C and incubated with HRP-linked secondary antibody for 2 hour at room temperature. The ChemiDocTM XRS Plus luminescent image analyzer (Bio-Rad, USA) was used to quantify the protein bands. GAPDH was used as a loading control.
Statistical analysis
The Statistical Package for the Social Sciences version 18.0 software (SPSS, Chicago, IL) was used to analyze all the data. All data were shown as mean ± standard error mean (SEM). A paired T-test, an independent-sample T-test or one-way ANOVA test followed by Bonferroni's tests were used for statistical analysis. P < 0.05 was considered as statistical significance.
Results
miR-224 is increased in colorectal cancer tissues
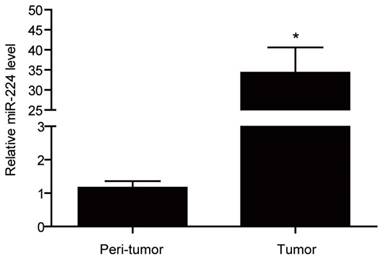
miR-224 expression levels have been frequently reported to be elevated in colorectal cancer as compared to normal colorectal tissues [13-15]. Here used qRT-PCRs, we found that that miR-224 was significantly increased by 29.49 fold in colorectal cancer tissues compared with their corresponding matched peritumoral tissues based on 12 colorectal cancer patients (Fig.1).
miR-224 is overexpressed in tissues of human colorectal cancer. qRT-PCR for miR-224 expression in the tumor versus the peri-tumortissues of colorectal cancer from a total of 12 patients (n=12 per group). *, p<0.05.
miR-224 controls HCT116 cell viability and cell proliferation
CCK-8 was used to measure cell viability of HCT116 cells transfected with miR-224 mimic and inhibitor. It was found that miR-224 mimic could significantly increase HCT116 cell viability while miR-224 inhibitor decreased that (Fig. 2A).
miR-224 promotes human colorectal cancer cell line HCT116 proliferation. (A) CCK-8 assay showed that miR-224 mimic increased the viability of HCT116 cells, while miR-224 inhibitor decreased that (n=6). (B) The EdU positive rate was increased by miR-224 mimic, while miR-224 inhibitor decreased that (n=4). Scale bar=50μm. (C)Flow cytometry for cell cycle showed that miR-224 mimic decreased G1-phase cells rate and increased S-phase cells rate, while miR-224 inhibitor increased G1-phase cells rate and decreased S-phase cells rate (n=6). *, p<0.05.
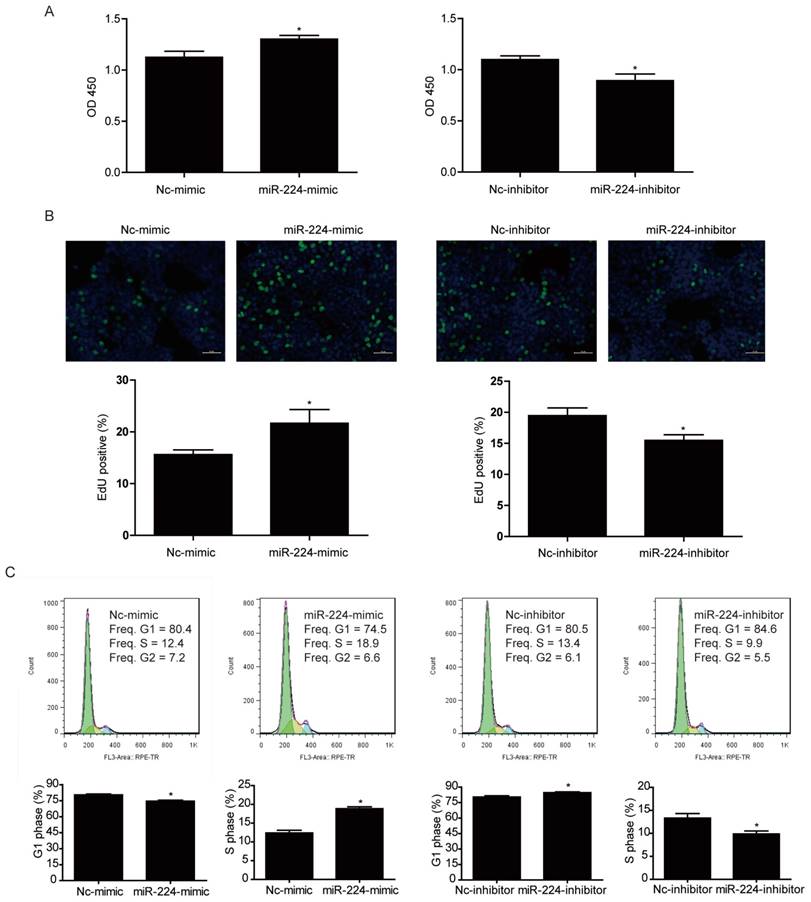
To further investigate if the effect of miR-224 in controlling cell viability was because of its effect in cell proliferation, EdU incorporation assays were used to determine the effects of miR-224 in DNA synthesis, which is an important hallmark of cancer cell growth. As indicated in Fig. 2B, miR-224 overexpression increased EdU-positive cell population while miR-224 suppression significantly decreased that, suggesting that miR-224 was able to control DNA synthesis of HCT116 cells.
Besides EdU incorporation assays, we also used flow cytometry to determine HCT116 cell cycle distribution. We found that miR-224 overexpression decreased G1 phase cell population and increased S phase cell population while miR-224 inhibition increased G1 phase cell population and decreased S phase cell population (Fig.2C), further supporting that miR-224 was able to promote HCT116 cell proliferation.
Smad4 is a target gene of miR-224
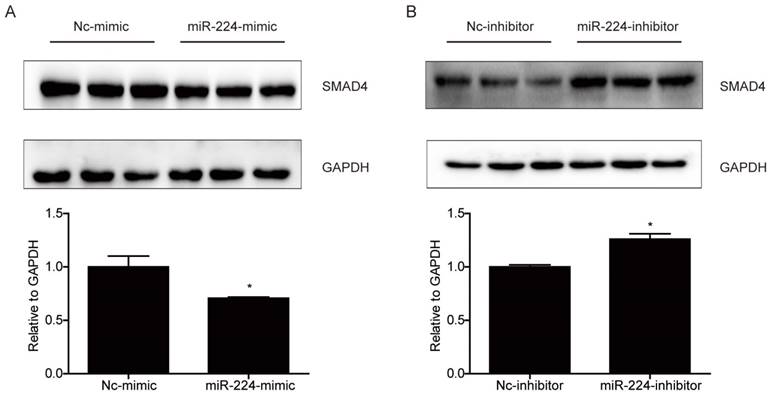
Smad4 is a reported target gene of miR-224 [11], however, due to the cell specific effects of miRNAs [12], it is unclear if Smad4 is a target gene of miR-224 and responsible for its effects in HCT116 cells. Based on Western blot, we determined the effects of miR-224 in endogenous expressions of Smad4 in HCT116 cells first. We found that the protein level of Smad4 could be endogenously negatively regulated by miR-224 in HCT116 cells (Fig.3). To further determine if Smad4 mediates the effects of miR-224 in HCT116 cells, siRNA to Smad4 was transfected into HCT116 cells, and qRT-PCR was used to confirm the effects of Smad4 siRNA in knock-downing Smad4 (Fig.4A). miR-224 inhibitor and Smad4 were co-transfected, and EdU assays indicated that Smad4 siRNA could reverse the proliferation suppression effects of miR-224 in HCT116 cells (Fig.4B). Thus, Smad4 is a target gene of miR-224, mediating its effects in proliferation.
Smad4 is a target gene of miR-224 in HCT116 cells. (A) Smad4 protein level was decreased by miR-224 mimic in HCT116 cells (n=3). (B) Smad4 protein level was increased by miR-224 inhibitor in HCT116 cells (n=3). *, p<0.05.
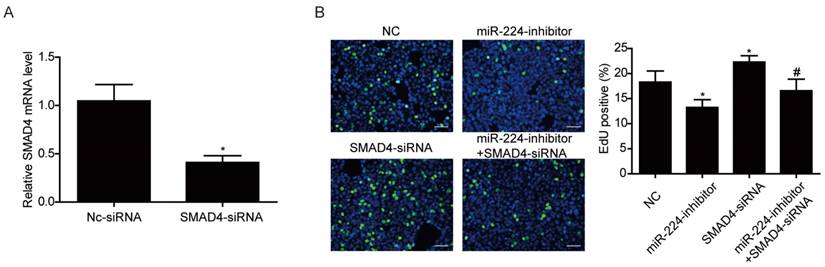
Smad4 mediates the effects of miR-224 in HCT116 cells proliferation. (A) qRT-PCR showed that Smad4 siRNA significantly downregulated Smad4 in HCT116 cells (n=4). (B) Silencing Smad4 restored the inhibitory effect of miR-224 inhibitor on proliferation in HCT116 cells. (n=4). Scale bar=50 μm.*, p<0.05 vs NC. #, p<0.05 vs miR-224 inhibitor group.
Discussion
Colorectal cancer represents the third most common cancer and is widely detected in the world, causing 700,000 deaths yearly [9]. The first-line treatment and second-line chemotherapy for colorectal cancer patients have been well-established [9]. However, novel therapies are needed to increase treatment efficacy [9]. Here we provide data that miR-224 controls HCT116 cell proliferation by targeting Smad4, suggesting that suppression of miR-224 could be a novel therapy for colorectal cancer.
miRNAs have been reported to be related to a variety of cancers [16]. Recent evidences proved that miR-224 was obviously aberrant up-regulated in a number of cancers including hepatocellular carcinoma [17], breast cancer [18], lung cancer [19] and colorectal cancer [13-15]. miR-224 has been found to be able to promote aggressive phenotype of colorectal cancer [20], and was also related to the progression and response to chemotherapy of colorectal cancer [21]. The clinical and biological significance of miR-224 has been reported in colorectal cancer metastasis. However, miR-224 has also been found to suppress colorectal cancer cell migration [13]. miR-224 could promote human colorectal cancer cell line proliferation [11]. However, due to the cell specific effects of miRNA [12], and HCT116 is a commonly used cell line for colorectal cancer study, it would be highly interested to explore the target gene responsible for the function of miR-224 in its proliferation. In this study, we found that miR-224 was able to induce DNA synthesis and transition of G1-to-S phase in HCT116 cells.
Smad family proteins play a key role in the process of transmitting TGF-β signal from cell surface receptors to the nucleus [22]. TGF-β signal plays a critical role in colorectal cancer and Smad4 is important in the function of TGF-β shift [22]. Smad4 knockout mice have been found to develop cancers in the gastrointestinal tract [23]. Smad4 has been reported to be a target gene of miR-224 in human colorectal cancer cell line SW480 [11]. However, the effect of miRNAs on cancer cell proliferation could be significantly changeable among different cell lines. As HCT116 is a widely used cell line for colorectal cancer study and the target gene responsible for the function of miR-224 in its proliferation is unclear. Here we provide direct evidence that Smad4 could be endogenous negatively regulated in HCT116 cells and as Smad4 siRNA could block the effects of miR-224 inhibitor, suggesting that Smad4 is a target gene of miR-224, responsible for the effects of miR-224 in controlling proliferation.
In summary, our study shows that miR-224 promotes HCT116 cells proliferation by targeting Smad4. Inhibition of miR-224 might be a novel therapeutic strategy for colorectal cancer.
Abbreviations
miRNAs, microRNAs; CCK-8, cell counting kit-8 assay; EdU, 5-ethynyl-2'-deoxyuridine; PCRs, quantitative reverse transcription polymerase chain reactions.
Acknowledgements
This work was supported by the grants from National Natural Science Foundation of China (81400635 to F. Wang, 81370586 to B. Ge), Health and Family Planning Commission of Shanghai (201540072 to Q. Huang), Shanghai Medical Guide Project from Shanghai Science and Technology Committee (14411971500 to F. Wang), and grants from Chinese Foundation for Hepatitis Prevention and Control (TQGB20140141 to F. Wang), Shanghai Science and Technology Committee (15411962600 to B. Ge).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M. et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359-86
2. Chu XY, Chen LB, Wang JH, Su QS, Yang JR, Lin Y. et al. Overexpression of survivin is correlated with increased invasion and metastasis of colorectal cancer. J Surg Oncol. 2012;105:520-8
3. Alevizos I, Illei GG. MicroRNAs as biomarkers in rheumatic diseases. Nat Rev Rheumatol. 2010;6:391-8
4. Xu J, Zhao J, Evan G, Xiao C, Cheng Y, Xiao J. Circulating microRNAs: novel biomarkers for cardiovascular diseases. J Mol Med (Berl). 2012;90:865-75
5. Suzuki HI, Miyazono K. Dynamics of microRNA biogenesis: crosstalk between p53 network and microRNA processing pathway. J Mol Med (Berl). 2010;88:1085-94
6. Tao L, Bei Y, Chen P, Lei Z, Fu S, Zhang H. et al. Crucial Role of miR-433 in Regulating Cardiac Fibrosis. Theranostics. 2016;6:2068-83
7. Wang H, Bei Y, Huang P, Zhou Q, Shi J, Sun Q. et al. Inhibition of miR-155 Protects Against LPS-induced Cardiac Dysfunction and Apoptosis in Mice. Mol Ther Nucleic Acids. 2016;5:e374
8. Shi J, Bei Y, Kong X, Liu X, Lei Z, Xu T. et al. miR-17-3p Contributes to Exercise-Induced Cardiac Growth and Protects against Myocardial Ischemia-Reperfusion Injury. heart. 2017;7:665
9. Moridikia A, Mirzaei H, Sahebkar A, Salimian J. MicroRNAs: Potential Candidates for Diagnosis and Treatment of Colorectal Cancer. J Cell Physiol. 2017
10. O'Bryan S, Dong S, Mathis JM, Alahari SK. The roles of oncogenic miRNAs and their therapeutic importance in breast cancer. Eur J Cancer. 2017;72:1-11
11. Zhang GJ, Zhou H, Xiao HX, Li Y, Zhou T. Up-regulation of miR-224 promotes cancer cell proliferation and invasion and predicts relapse of colorectal cancer. Cancer Cell Int. 2013;13:104
12. Yamashita R, Sato M, Kakumu T, Hase T, Yogo N, Maruyama E. et al. Growth inhibitory effects of miR-221 and miR-222 in non-small cell lung cancer cells. Cancer Med. 2015;4:551-64
13. Ling H, Pickard K, Ivan C, Isella C, Ikuo M, Mitter R. et al. The clinical and biological significance of MIR-224 expression in colorectal cancer metastasis. Gut. 2016;65:977-89
14. Wang Y, Ren J, Gao Y, Ma JZ, Toh HC, Chow P. et al. MicroRNA-224 targets SMAD family member 4 to promote cell proliferation and negatively influence patient survival. PLoS One. 2013;8:e68744
15. Liao WT, Li TT, Wang ZG, Wang SY, He MR, Ye YP. et al. microRNA-224 promotes cell proliferation and tumor growth in human colorectal cancer by repressing PHLPP1 and PHLPP2. Clin Cancer Res. 2013;19:4662-72
16. Morris MR, Latif F. The epigenetic landscape of renal cancer. Nat Rev Nephrol. 2017;13:47-60
17. Wang Y, Lee CG. Role of miR-224 in hepatocellular carcinoma: a tool for possible therapeutic intervention? Epigenomics. 2011;3:235-43
18. Huang L, Dai T, Lin X, Zhao X, Chen X, Wang C. et al. MicroRNA-224 targets RKIP to control cell invasion and expression of metastasis genes in human breast cancer cells. Biochem Biophys Res Commun. 2012;425:127-33
19. Cui R, Kim T, Fassan M, Meng W, Sun HL, Jeon YJ. et al. MicroRNA-224 is implicated in lung cancer pathogenesis through targeting caspase-3 and caspase-7. Oncotarget. 2015;6:21802-15
20. Li T, Lai Q, Wang S, Cai J, Xiao Z, Deng D. et al. MicroRNA-224 sustains Wnt/beta-catenin signaling and promotes aggressive phenotype of colorectal cancer. J Exp Clin Cancer Res. 2016;35:21
21. Amankwatia EB, Chakravarty P, Carey FA, Weidlich S, Steele RJ, Munro AJ. et al. MicroRNA-224 is associated with colorectal cancer progression and response to 5-fluorouracil-based chemotherapy by KRAS-dependent and -independent mechanisms. Br J Cancer. 2015;112:1480-90
22. Massague J. TGFbeta in Cancer. Cell. 2008;134:215-30
23. Yang G, Yang X. Smad4-mediated TGF-beta signaling in tumorigenesis. Int J Biol Sci. 2010;6:1-8
Author contact
Corresponding authors: Dr. Bujun Ge, Department of General Surgery, Tongji Hospital, Tongji University School of Medicine, 389 Xin Cun Road, Shanghai 200065, P.R. China Tel: 0086-21-66111428; Fax: 0086-21-66111428 E-mail: gebujuncom Dr. Qi Huang, Department of General Surgery, Tongji Hospital, Tongji University School of Medicine, 389 Xin Cun Road, Shanghai 200065, P.R. China Tel: 0086-21-66111428; Fax: 0086-21-66111428 E-mail: 1410675edu.cn

 Global reach, higher impact
Global reach, higher impact