Impact Factor ISSN: 1449-1907
Int J Med Sci 2013; 10(12):1755-1760. doi:10.7150/ijms.6749 This issue Cite
Research Paper
TNF-α -857C>T Genotype is Predictive of Clinical Response after Treatment with Definitive 5-Fluorouracil/cisplatin-based Chemoradiotherapy in Japanese Patients with Esophageal Squamous Cell Carcinoma
1. Department of Hospital Pharmacy, School of Medicine, Kobe University, Kobe 650-0017, Japan;
2. Department of Pharmacotherapy, Programs for Applied Biomedicine, Graduate School of Biomedical Sciences, Hiroshima University, Hiroshima 734-8553, Japan;
3. School of Pharmacy and Pharmaceutical Sciences, Mukogawa Women's University, Nishinomiya 663-8179, Japan;
4. Department of Gastroenterology, Kobe University Graduate School of Medicine, Kobe 650-0017, Japan;
5. Department of Medical Pharmaceutics, Kobe Pharmaceutical University, Kobe 658-8558, Japan;
6. Department of Clinical Pharmacy, Kyoto Pharmaceutical University, Kyoto 607-8414, Japan;
7. Department of Pharmaceutical Health Care, Faculty of Pharmaceutical Sciences, Himeji Dokkyo University, Himeji 670-8524, Japan.
Received 2013-5-23; Accepted 2013-10-6; Published 2013-10-15
Abstract
Background: Genotypes of tumor necrosis factor alpha (TNF-α) and its surface receptors, TNFRSF1A and TNFRSF1B, have been examined in terms of the progression, metastasis, clinical efficacy, and prognosis of various cancers; however, little is known about their effects on clinical outcome in patients with esophageal squamous cell carcinoma (ESCC). In this study, TNF-α and TNFRSF1A genotypes were retrospectively evaluated in terms of predicting clinical response, long-term survival, and severe acute toxicities in 46 male Japanese ESCC patients treated with definitive 5-fluorouracil (5-FU)/cisplatin (CDDP)-based chemoradiotherapy (CRT).
Methods: A course consisted of the continuous infusion of 5-FU at 400 mg/m2/day for days 1-5 and 8-12, the infusion of CDDP at 40 mg/m2/day on days 1 and 8, and radiation at 2 Gy/day on days 1-5, 8-12, and 15-19, with a second course being repeated after a 2-week interval. The TNF-α -1031T>C (rs1799964), -863C>A (rs1800630), -857C>T (rs1799724), -308G>A (rs1800629), -238G>A (rs361525), TNFRSF1A -609G>T (rs4149570), and 36A>G (rs767455) genotypes were evaluated.
Results: The TNF-α -857C>T genotype was found to be predictive of clinical response, i.e., complete response or not (P = 0.010, Fisher's exact test), but had no effect on long-term survival (CC-857 vs. CT-857 + TT-857, P = 0.072, Fisher's exact test, P = 0.070, Log-rank test).
Conclusions: The TNF-α -857C>T genotype was found to be predictive of clinical response and was more likely to predict long-term survival in Japanese ESCC patients receiving definitive 5-FU/CDDP-based CRT. Further clinical investigations with a larger number of patients or experiments in vitro should be performed to assess the predictive value of this genotype following CRT.
Keywords: esophageal squamous cell carcinoma, chemoradiotherapy, clinical response, prognosis, tumor necrosis factor.
Background
Tumor necrosis factor alpha (TNF-α), an important proinflammatory cytokine, is produced by activated macrophages and exerts its effects by binding to its two cognate cell surface receptors, TNFRSF1A/TNFR1 (p55/60) and TNFRSF1B/TNFR2 (p75/80). TNF-α and its superfamily members are well-known to have both beneficial and harmful effects [1]. Although TNF-α was discovered as a cytokine that could kill tumor cells, TNF-α can also lead to the proliferation, invasion, and metastasis of tumor cells [1]. The activities of TNF-α were shown to be dependent on the expression of TNFRSF1A and TNFRSF1B, which have different intracellular domains and mediate unique as well as overlapping functions, including the activation of transcriptional activator nuclear factor kappa B (NF-κB) and programmed cell death [2]. Although TNFRSF1B cannot directly mediate apoptosis, TNFRSF1A was shown to modulate TNFRSF1A-induced apoptosis, and can directly activate NF-κB [1, 3-5]. TNF-α and TNFRSF1A may play important roles in esophageal squamous cell carcinoma (ESCC), TNFRSF1A was found to be strongly expressed in an esophageal carcinoma cell line, and silencing the expression of TNFRSF1A promoted cell proliferation and downregulated the apoptotic rate [6]. In addition, 5-fluorouracil (5-FU) is commonly used in combination therapy for ESCC, and TNF-α decreased the effect of 5-FU and increased the level of proliferation in esophageal carcinoma cell lines [7].
The TNF-α and TNFRSF1A genes are highly polymorphic, and several single nucleotide polymorphisms have been identified in these genes, which may contribute to differences in gene expression levels and transcription [8-18]. Some of these polymorphisms were recently shown to be related to tumor progression, metastasis, prognosis, and survival [19-22]. However, these studies included patients treated with surgery or neoadjuvant chemoradiotherapy (CRT); therefore, the association between these TNF-α and TNFRSF1A polymorphisms and the prediction of clinical outcome with definitive CRT in ESCC remains unknown.
Predicting therapeutic responses is important in definitive CRT prior to the initiation of treatment; we previously reported a significant correlation between clinical response and survival, and showed that the TNFRSF1B 1466A/G (rs1061624) genotype could predict clinical response with definitive 5-FU/cisplatin (CDDP)-based CRT in 46 male Japanese patients with ESCC [23]. Although these findings suggest that TNF-α and its receptors may play a critical role in clinical response and survival, to the best of our knowledge, no published study has investigated associations between the polymorphisms of TNF-α and TNFRSF1A genes and the prediction of clinical outcome in ESCC patients treated with definitive CRT. The TNF-α -1031T>C (rs1799964), -863C>A (rs1800630), -857C>T (rs1799724), -308G>A (rs1800629), -238G>A (rs361525), TNFRSF1A -609G>T (rs4149570), and 36A>G (rs767455) genotypes were selected for genotyping because they affect expression or transcription and have been associated with tumor progression, metastasis, prognosis, or survival in cancer [8-22].
In this study, patients with ESCC were followed-up for 5 years after treatment with definitive 5-FU/CDDP-based CRT, and the effects of the TNF-α -1031T>C, -863C>A, -857C>T, -308G>A, -238G>A, TNFRSF1A -609G>T, and 36A>G genotypes were retrospectively evaluated in terms of predicting long-term survival, clinical response, and severe acute toxicities.
Methods
Ethics statements
Studies have been performed to evaluate the effects of genetic polymorphisms on clinical response, survival or severe acute toxicities during treatment with definitive 5-FU/CDDP-based CRT in Japanese patients with ESCC [23-27]. They were conducted with the authorization of the Institutional Review Board (IRB) and followed the Medical Research Council guidelines of Kobe University. All patients agreed to the studies and preservation of genomic DNA for future investigations, and additional studies were again authorized by the IRB and followed the guidelines of Kobe University. Written informed consent was obtained from all participants prior to genotyping.
Patients and study protocol
Forty-six ESCC patients treated with definitive 5-FU/CDDP-based CRT between October, 2003 and June, 2006 at Kobe University Hospital, Japan, were followed-up for 5 years. Female patients were excluded because of differences in TNF-α levels between males and females [28]. The demographic and clinicopathological characteristics of 46 male Japanese ESCC patients are summarized in Table 1, as reported previously [23]. Survival time was defined as the time from the initiation of treatment to death from any cause or to the last date of the confirmation of survival. Survival data were updated on June 25, 2011.
Demographic and clinicopathological characteristics of 46 Japanese patients with esophageal squamous cell carcinoma.
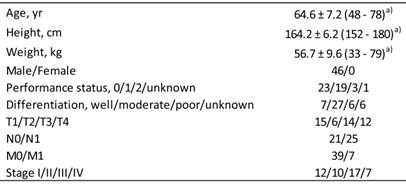

a) Values represent the mean ± SD, with the range in parentheses.
A course consisted of the continuous infusion of 5-FU at 400 mg/m2/day for days 1-5 and 8-12, the infusion of CDDP at 40 mg/m2/day on days 1 and 8, and external beam radiotherapy using megavoltage (≥6 MV) X-rays was administered at 2 Gy/day on days 1 to 5, 8 to 12, and 15 to 19 (a total dose of 60 Gy in 30 fractions), with a second course being repeated after a 2-week interval [29, 30]. The clinical target volume (CTV) for 60 Gy irradiation included the primary tumor plus a 5-cm craniocaudal margin, and metastatic lymph nodes plus a 1-cm margin. Planning target volume was defined as CTV plus 5- to 20-mm margins for uncertainty. Elective nodal irradiation (40 Gy) to the mediastinal and perigastric lymph nodes in all cases, cervical lymph nodes for an upper thoracic primary tumor, and celiac lymph nodes for a lower thoracic primary tumor was also performed. Three-dimensional computed tomography or X-ray simulation was performed, which allowed for two-dimensional anterior-posterior opposed fields and a bilateral oblique boost. Heterogeneity-uncorrected doses were used. Salvage surgery, endoscopic treatment, or another regimen of chemotherapy was scheduled if disease progression/recurrence was observed. Clinical response and severe acute toxicities; leucopenia, stomatitis, and cheilitis, were evaluated in a previous study [23]. Associations between the TNF-α and TNFRSF1A genotypes and clinical response, long-term survival, and severe acute toxicities were evaluated in this study. It should be noted that the patients participating in this study were the same as those in our previous report [23].
Data analysis and statistics
All values reported are the mean ± standard deviation (SD). Fisher's exact test and the log-rank test were used to evaluate the effects of genotypes on long-term survival. P values of less than 0.05 (two tailed) were considered to be significant.
Genomic DNA extraction and genotyping
Genomic DNA was isolated from whole blood using a Taq-Man® Sample-to-SNP™ kit (Applied Biosystems, Foster City, CA, USA), according to the manufacturer's directions, and stored at -80°C until analysis. TNF-α and TNFRSF1A genotypes were determined in a TaqMan® MGB probe-based polymerase chain reaction (PCR) using the 7900HT™ real-time PCR system (Applied Biosystems) and TaqMan® SNP genotyping assays (Applied Biosystems). PCR was carried out according to the manufacturer's protocol. Negative controls containing H2O instead of DNA were added for each set of reactions to detect contamination.
Results
The 5-year survival rate was 68.2 % (15/22) for patients with CR and 20.8 % (5/24) for those with non-CR, and survival depended on clinical response, i.e., CR or non-CR (P = 0.0025, Fisher's exact test; P = 0.0004, Log-rank test). A significant difference was observed in lymph node metastasis (P = 0.036, Fisher's exact test) between patients who survived 5 years or more and those who survived less than 5 years. The 5-year survival rate was 61.9 % (13/21) in N0.
Table 2 shows the association between the TNF-α and TNFRSF1A genotypes, and TNF-α -1031T>C, -863C>A, -857C>T, -308G>A, -238G>A, TNFRSF1A -609G>T, and 36A>G genotypes and clinical response in ESCC patients. Among the 7 genotypes evaluated, the TNF-α -857C>T genotype had a significant effect on clinical response (P = 0.010, Fisher's exact test).
Effect of the TNF-α and TFNRSF1A genotypes on clinical response in 46 Japanese patients with esophageal squamous cell carcinoma.
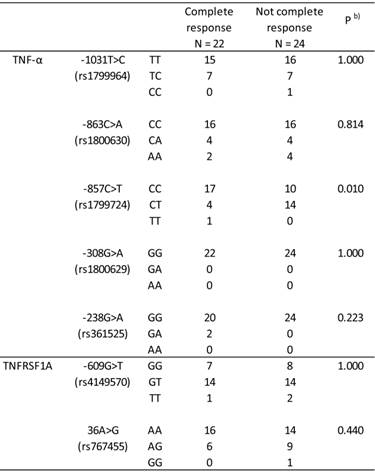
Genotype frequencies followed the Hardy-Weinberg principle. b) Complete response vs. not complete response.
The TNF-α and TNFRSF1A genotypes, and TNF-α -1031T>C, -863C>A, -857C>T, -308G>A, -238G>A, TNFRSF1A -609G>T, and 36A>G genotypes had no effect on 5-year survival (data not shown). The 5-year survival rates were 55.6 % (15/27) and 26.3 % (5/19) in patients with CC-857 and CT-857 + TT-857, respectively, and the TNF-α -857C>T genotype had no significant effect on 5-year survival (CC-857 vs. CT-857 + TT-857, P = 0.072, Fisher's exact test).
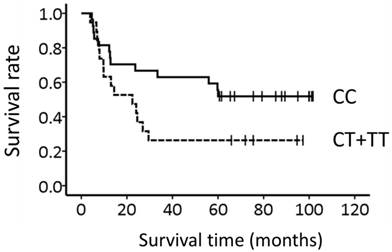
Figure 1 shows the association between the TNF-α -857C>T genotype and survival. The median overall survival time (±SE) was more than 60 months and 22.4 ± 8.0 months for patients with CC-857 and CT-857 + TT-857, respectively; however, the genotype could not predict survival (P = 0.070, Log-rank test).
Association of the TNF-α -857C>T genotype with long-term survival in 46 Japanese patients with esophageal squamous cell carcinoma. Lines: patients with CC-857 (N = 27) and CT-857 + TT-857 (N = 19). Survival appeared to be associated with the genotype (P = 0.070, Log-rank test).
The TNF-α and TNFRSF1A genotypes were not associated with severe acute leucopenia, stomatitis, or cheilitis (data not shown). In addition, these genotypes had no effect on the TNM classification or disease stage. These genotype frequencies followed the Hardy-Weinberg principle, analyzed by Fisher's exact test.
Discussion
We examined the effects of 7 selected polymorphisms in TNF-α and TNFRSF1A on clinical response, long-term survival, and severe acute toxicities with definitive 5-FU/CDDP-based CRT in Japanese ESCC patients. We identified the TNF-α -857C>T genotype as a new predictive genetic marker of clinical response to treatment; patients with the T-allele at position -857 in the TNF-α promoter had a poorer response, whereas no such association was found for TNF-α -1031T>C, -863C>A, -308G>A, -238G>A, TNFRSF1A -609G>T, or 36A>G (Table 2). Clinical response was significantly correlated with long-term survival, whereas the TNF-α -857C>T genotype was not. Long-term survival was determined by lymph node metastasis. The TNF-α -857C>T genotype was independent of clinicopathological factors, TNM classification, and disease stage. None of the other polymorphisms of TNF-α and TNFRSF1A had any effect on clinical response, long-term survival, or severe acute leucopenia, stomatitis, and cheilitis.
The TNF-α gene is located on Chr 6p21, within the class III region of the major histocompatibility complex, between HLA-B and HLA-DR. TNF-α performs multiple functions in immunity, inflammation, differentiation, apoptosis, and the control of cell proliferation via NF-κB through the distinct receptors, TNFRSF1A and TNFRSF1B [1]. When a member of the TNF superfamily binds with TNFRSF1A, it results in receptor trimerization and the clustering of intracellular death domains as the extrinsic pathway of apoptosis, which leads to the induction of apoptosis via the activation of NF-κB and caspases [31]. In a previous in vitro study, TNF-α was shown to strongly increase the activity of NF-κB, decrease the effect of 5-FU, and increase the level of proliferation [7]. Several polymorphisms have been identified in the TNF-α gene, which may contribute to differences in gene expression levels [8-15]. These results suggest the possibility that if polymorphisms in the promoter region of TNF-α increase TNF-α production, tolerance to 5-FU may increase via the activation of NF-κB. In previous reports, the TNF-α -1031C allele, -863A allele, and -857T allele increased TNF-α production [11, 14], and patients in our study with the T-allele at position -857 in the TNF-α promoter had a poorer response than that of CC-857 patients. Our results are consistent with those of previous studies in which the TNF-α -857C>T genotype could predict clinical response. Therefore, the TNF-α -857C>T genotype has been proposed to be predictive of clinical response caused by increase tolerance to 5-FU. On the other hand, TNF-α -1031T>C and -863C>A were not associated with clinical response. This may have been because the effect of activation with TNF-α polymorphisms differed depending on the cell lines used and ethnicity, and a combination of these polymorphisms [8-15, 32]. Furthermore, TNF-α production was significantly increased by the TNF-α -857T allele alone, whereas the TNF-α -1031C allele and -863A allele required both mutations to increase TNF-α production in Japanese subjects [11, 32]. These findings suggest that the influence of the TNF-α -857C>T polymorphism on Japanese patients may be higher than that of the -1031T>C and -863C>A polymorphisms. In a previous study, the TNF-α -857T allele showed better overall survival in gastric cancer [20], which was in contrast with our findings. However, this difference may exist due to the therapeutic methods used, surgical treatment, drugs administered, and different types of cancer. Therefore, the effect of the TNF-α -857C>T genotype may be important in Japanese ESCC patients with definitive CRT.
The G to A nucleotide change at -308 and -238 altered the transcription factor binding site, resulting in the increased production of TNF-α [8, 9]. Furthermore, the -308A allele has been associated with poor performance status, lesions, and large-size tumors of head and neck squamous cell carcinoma [19], and also with the prevalence of the T4 stage of disease and distant metastasis of gastric cancer [21]. In our study, TNF-α -308G>A and -238G>A were not associated with clinical response, survival, or clinicopathological factors. Although our samples had few variants to evaluate the association between TNF-α -308G>A and -238G>A, and clinical outcomes, these frequencies were almost the same as previous reports on Japanese patients [11, 32]. These 2 polymorphisms may not have had a strong influence on Japanese ESCC patients with definitive CRT. More studies with a larger sample size are needed to clarify the effects of these 2 polymorphisms. On the other hand, in spite of the clinical response associated with survival, the TNF-α -857C>T genotype could not predict survival, which may have been due to the small sample size (N = 46). Here, 5 of 7 patients with CR appeared to have died because of late toxicities and other cancer deaths. This may have affected the association between the TNF-α -857C>T genotype and long-term survival.
The TNFRSF1A -609G/T and 36A/G genotypes were also not associated with clinical response or long-term survival. TNFRSF1A may play an apoptosis-mediating role in esophageal carcinoma [6], and TNFRSF1A -609G/T was previously shown to increase TNFRSF1A mRNA expression in subjects including patients with invasive pulmonary aspergillosis [17]. However, another report showed the decreased expression of TNFRSF1A -609G/T in hepatocellular carcinoma cells [16]; therefore, further studies are needed to clarify the effect of TNFRSF1A -609G/T. In addition, the TNFRSF1A -609T allele was associated with poor overall survival and disease-free survival in non-small-cell lung cancer [22]; however, approximately half of these patients did not receive chemotherapy. TNFRSF1A 36A>G was previously suggested to be a potential exonic splicing enhancer site for three splicing factors, SRp55, SF2/ASF2, and SF2/ASF1 [18]; however, its effect on cancer remains unknown. TNFRSF1B has a higher binding affinity to TNF-α than TNFRRSF1A [2], and NF-κB activity was mediated not only by TNFRSF1A, but also by TNFRSF1B. The expression levels of the TNFRSF1B gene were higher in the colorectal cancer specimens of non-responding patients to 5-FU than in those from responding patients [33]. In addition, a high concentration of TNF-α induced responses through TNFRSF1A, which transmits signals promoting growth inhibition and cell death, while a low concentration stimulated responses through the TNFRSF1B-NF-κB signaling pathway, which stimulates cell proliferation [34, 35]. Moreover, the results of our previous study showed the relationship between TNFRSF1B 1466A>G and clinical response in ESCC [23], and suggested that the relationship of TNF-α and TNFRSF1B to clinical response in ESCC with definitive CRT may be more important than that of TNFRSF1A to TNF-α concentrations and the effect of 5-FU.
In conclusion, the TNF-α -857C>T genotype was shown to be a new predictive genetic marker of clinical response and may be predictive of long-term survival following treatment with definitive 5-FU/CDDP-based CRT. This is the first study to identify the association between TNF-α polymorphisms and clinical response with definitive CRT in Japanese ESCC patients. Further clinical studies with a larger number of cases, in which the relationship between long-term survival and clinical response with polymorphism-dependent TNF-α expression levels, including the TNF-α and TNFRSF1B genotypes, is examined, are needed to clarify the effects of this genotype.
Authors' contributions
AK, TT, TN, TA, and MH conceived, designed, and coordinated the study. HO, IM, TO, and TT evaluated clinical outcomes. MY, MF, and NO carried out the genotyping study. AK, MY, and KN performed data management. HO, TH, KO, and MH prepared the manuscript. All authors read and approved the final manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Aggarwal BB. Signalling pathways of the TNF superfamily: a double-edged sword. Nat Rev Immunol. 2003;3:745-56
2. Tartaglia LA, Goeddel DV. Two TNF receptors. Immunol Today. 1992;13:151-3
3. Tartaglia LA, Goeddel DV, Reynolds C. et al. Stimulation of human T-cell proliferation by specific activation of the 75-kDa tumor necrosis factor receptor. J Immunol. 1993;151:4637-41
4. Rothe M, Sarma V, Dixit VM. et al. TRAF2-mediated activation of NF-kappa B by TNF receptor 2 and CD40. Science. 1995;269:1424-7
5. Carpentier I, Coornaert B, Beyaert R. Function and regulation of tumor necrosis factor receptor type 2. Curr Med Chem. 2004;11:2205-12
6. Changhui M, Tianzhong M, Zhongjing S. et al. Silencing of tumor necrosis factor receptor 1 by siRNA in EC109 cells affects cell proliferation and apoptosis. J Biomed Biotechnol. 2009;2009:760540
7. Hatata T, Higaki K, Nanba E. et al. Inhibition of nuclear factor-kappaB activity by small interfering RNA in esophageal squamous cell carcinoma cell lines. Oncol Rep. 2011;26:659-64
8. Wilson AG, Symons JA, McDowell TL. et al. Effects of a polymorphism in the human tumor necrosis factor alpha promoter on transcriptional activation. Proc Natl Acad Sci U S A. 1997;94:3195-9
9. Kroeger KM, Carville KS, Abraham LJ. The -308 tumor necrosis factor-alpha promoter polymorphism effects transcription. Mol Immunol. 1997;34:391-9
10. Huizinga TW, Westendorp RG, Bollen EL. et al. TNF-alpha promoter polymorphisms, production and susceptibility to multiple sclerosis in different groups of patients. J Neuroimmunol. 1997;72:149-53
11. Higuchi T, Seki N, Kamizono S. et al. Polymorphism of the 5'-flanking region of the human tumor necrosis factor (TNF)-alpha gene in Japanese. Tissue Antigens. 1998;51:605-12
12. Skoog T, van't Hooft FM, Kallin B. et al. A common functional polymorphism (C-->A substitution at position -863) in the promoter region of the tumour necrosis factor-alpha (TNF-alpha) gene associated with reduced circulating levels of TNF-alpha. Hum Mol Genet. 1999;8:1443-9
13. Sharma S, Sharma A, Kumar S. et al. Association of TNF haplotypes with asthma, serum IgE levels, and correlation with serum TNF-alpha levels. Am J Respir Cell Mol Biol. 2006;35:488-95
14. Lv K, Chen R, Cai Q. et al. Effects of a single nucleotide polymorphism on the expression of human tumor necrosis factor-alpha. Scand J Immunol. 2006;64:164-9
15. Cui G, Wang H, Li R. et al. Polymorphism of tumor necrosis factor alpha (TNF-alpha) gene promoter, circulating TNF-alpha level, and cardiovascular risk factor for ischemic stroke. J Neuroinflammation. 2012;9:235
16. Kim S, Moon SM, Kim YS. et al. TNFR1 promoter -329G/T polymorphism results in allele-specific repression of TNFR1 expression. Biochem Biophys Res Commun. 2008;368:395-401
17. Sainz J, Salas-Alvarado I, Lopez-Fernandez E. et al. TNFR1 mRNA expression level and TNFR1 gene polymorphisms are predictive markers for susceptibility to develop invasive pulmonary aspergillosis. Int J Immunopathol Pharmacol. 2010;23:423-36
18. Park TJ, Kim HJ, Kim JH. et al. Associations of CD6, TNFRSF1A, and IRF8 polymorphisms with risk of inflammatory demyelinating diseases. Neuropathol Appl Neurobiol. 2013Aug;39(5):519-30
19. Correa GT, Bandeira GA, Cavalcanti BG. et al. Association of -308 TNF-alpha promoter polymorphism with clinical aggressiveness in patients with head and neck squamous cell carcinoma. Oral Oncol. 2011;47:888-94
20. Tahara T, Shibata T, Nakamura M. et al. Effect of IL-1beta and TNF-alpha polymorphisms on the prognosis and survival of gastric cancer patients. Clin Exp Med. 2011;11:211-7
21. Hong Y, Ge Z, Jing C. et al. Functional Promoter -308G>A Variant in Tumor Necrosis Factor alpha Gene Is Associated with Risk and Progression of Gastric Cancer in a Chinese Population. PLoS One. 2013;8:e50856
22. Lee EB, Jeon HS, Yoo SS. et al. Polymorphisms in apoptosis-related genes and survival of patients with early-stage non-small-cell lung cancer. Ann Surg Oncol. 2010;17:2608-18
23. Kuwahara A, Yamamori M, Fujita M. et al. TNFRSF1B A1466G genotype is predictive of clinical efficacy after treatment with a definitive 5-fluorouracil/cisplatin-based chemoradiotherapy in Japanese patients with esophageal squamous cell carcinoma. J Exp Clin Cancer Res. 2010;29:100
24. Okuno T, Tamura T, Yamamori M. et al. Favorable genetic polymorphisms predictive of clinical outcome of chemoradiotherapy for stage II/III esophageal squamous cell carcinoma in Japanese. Am J Clin Oncol. 2007;30:252-7
25. Sakaeda T, Yamamori M, Kuwahara A. et al. VEGF G-1154A is predictive of severe acute toxicities during chemoradiotherapy for esophageal squamous cell carcinoma in Japanese patients. Ther Drug Monit. 2008;30:497-503
26. Tamura T, Kuwahara A, Yamamori M. et al. VEGF -634C/G genotype is predictive of long-term survival after treatment with a definitive 5-fluorouracil/cisplatin-based chemoradiotherapy in Japanese patients with esophageal squamous cell carcinoma. Int J Med Sci. 2012;9:833-7
27. Miki I, Nakamura T, Kuwahara A. et al. THRB genetic polymorphisms can predict severe myelotoxicity after definitive chemoradiotherapy in patients with esophageal squamous cell carcinoma. Int J Med Sci. 2012;9:748-56
28. Moxley G, Posthuma D, Carlson P. et al. Sexual dimorphism in innate immunity. Arthritis Rheum. 2002;46:250-8
29. Ohtsu A, Boku N, Muro K. et al. Definitive chemoradiotherapy for T4 and/or M1 lymph node squamous cell carcinoma of the esophagus. J Clin Oncol. 1999;17:2915-21
30. Kaneko K, Ito H, Konishi K. et al. Definitive chemoradiotherapy for patients with malignant stricture due to T3 or T4 squamous cell carcinoma of the oesophagus. Br J Cancer. 2003;88:18-24
31. Shen HM, Pervaiz S. TNF receptor superfamily-induced cell death: redox-dependent execution. Faseb J. 2006;20:1589-98
32. Soga Y, Nishimura F, Ohyama H. et al. Tumor necrosis factor-alpha gene (TNF-alpha) -1031/-863, -857 single-nucleotide polymorphisms (SNPs) are associated with severe adult periodontitis in Japanese. J Clin Periodontol. 2003;30:524-31
33. Matsuyama R, Togo S, Shimizu D. et al. Predicting 5-fluorouracil chemosensitivity of liver metastases from colorectal cancer using primary tumor specimens: three-gene expression model predicts clinical response. Int J Cancer. 2006;119:406-13
34. Chainy GB, Singh S, Raju U. et al. Differential activation of the nuclear factor-kappa B by TNF muteins specific for the p60 and p80 TNF receptors. J Immunol. 1996;157:2410-7
35. McFarlane SM, Pashmi G, Connell MC. et al. Differential activation of nuclear factor-kappaB by tumour necrosis factor receptor subtypes. TNFR1 predominates whereas TNFR2 activates transcription poorly. FEBS Lett. 2002;515:119-26
Author contact
Corresponding author: Midori Hirai, Ph.D., Department of Hospital Pharmacy, School of Medicine, Kobe University, Kobe 650-0017, Japan, Tel: +81-78-382-6640, Fax: +81-78-382-6676, e-mail: midorihac.jp.

 Global reach, higher impact
Global reach, higher impact