ISSN: 1449-1907International Journal of Medical Sciences
Int J Med Sci 2018; 15(1):69-76. doi:10.7150/ijms.21113 This issue Cite
Research Paper
Lentivirus-mediated shRNA Targeting CNN2 Inhibits Hepatocarcinoma in Vitro and in Vivo
1. Department of Biochemistry and Molecular Biology, Guangxi Medical University, 22 Shuangyong Road, Nanning 530021, the Guangxi Zhuang Autonomous Region, China;
2. Guangxi Key Laboratory of Biological Targeting Diagnosis and Therapy Research, Guangxi Medical University, 22 Shuangyong Road, Nanning 530021, the Guangxi Zhuang Autonomous Region, China.
Abstract
Objective: Hepatocellular carcinoma (HCC) is one of the most common malignant tumors with a high rate of mortality. Our previous study shows the expression of calponin 2 (CNN2) is up-regulated in hepatocellular carcinoma tissues, especially in metastatic ones. To better understand the role of CNN2 in HCC, RNA interference (RNAi) was used to explore its role in tumor growth and metastasis.
Methods: Lentivirus-mediated CNN2-shRNA was transfected into SK-hep-1 cells, and the efficacy of CNN2 expression, cell migration, invasion, proliferation and cell cycles were evaluated by quantitative real-time polymerase chain reaction (qRT-PCR), Western blot (WB), Transwell assay, methyl thiazol tetrazolium assay and flow cytometry, respectively. SK-hep-1 cells transfected with Lentivirus-CNN2 shRNA were xenografted in Balb/C nude mice to explore the effect of CNN2-shRNA in tumor growth. Xenograft tumor tissues were examined for their histopathology, cell apoptosis, the expression of total protein and their corresponding phosphorylated protein of MEK1/2, ERK1/2, AKT, by hematoxylin and eosin stain (H & E staining), TUNEL assay, immunohistochemical technique, respectively.
Results: Our research shows it is evident that CNN2 shRNA can effectively down-regulate the expressions of CNN2 mRNA and protein, inhibit cell proliferations, arrest cell cycles at the S phase and reduce cell migration and invasion. SK-hep-1 cells with CNN2 down-regulation have markedly attenuated tumor growth in nude mice. Xenograft tumor tissues have displayed typical tumor characteristics and no apoptosis is detected in shRNA group or in control group. No metastatic tumor was found in any group of nude mice. With CNN2 protein down-regulation, the protein of pMEK1/2 and pERK1/2 are effectively down-regulated, except pAKT, AKT, MEK1/2 and ERK1/2.
Conclusions: CNN2 plays an important role in tumor growth and metastasis, possibly through MEK1/2-ERK1/2 signaling pathway. Our study illustrate that CNN2 might be a potential target in HCC molecular target therapy.
Keywords: CNN2, shRNA, Hepatocellular Carcinoma, Molecular target therapy.
Introduction
Previous studies have shown that hepatocellular carcinoma (HCC) accounts for one-third of deaths caused by cancer worldwide, which makes it the most malignant tumors in the present time [1, 2]. Although surgery is the best approach to improve both the overall and recurrence-free survivals, most HCC patients are diagnosed at a late stage, missing the best time for carrying out surgeries [3]. In addition, because of the high incidence of vascular invasion and extra hepatic diffusion, about 70% of HCC patients will relapse within five years after treatments [4]. Therefore, it is urgent to explore new and more effective therapies for treating HCC. At present, more and more researchers devote themselves to seek appropriate metastasis-associated genes as the molecular targets for HCC therapy. By regulating the expressions of those genes, changes of cancer development can be evaluated [5-8].
CNN2 is one kind of actin cytoskeleton binding protein. It is a subtype of calponin protein family which expresses in smooth muscle cells as well as in non-muscle cells. A number of studies have found that CNN2 protein is closely related to cell proliferations and metastasis [9-11]. But the relationship between CNN2 and HCC development remains unclear.
The aim of our present research is to provide a new theoretical basis for molecular target therapy of HCC. In order to achieve this goal, functionalities of CNN2 shRNA on proliferation and metastasis of SK-hep-1 cells have been observed in vitro and in vivo.
Materials and Methods
Lentiviral vector encoding shRNA targeting CNN2
ShRNA was designed by Ambion company's online design tool. Lentiviral vector encoding CNN2 shRNA was synthesize and packaged by GenePhama company, Shanghai, China.
Cell culture and lentivirus infection
Kunming cell bank (Kunming, China) provided the SK-hep-1 cell line. Dulbecco's modified eagle's medium (DMEM) (Hyclone, USA) supplemented with 10% fetal bovine serum (FBS) (Gibco, USA) was used to culture SK-hep-1 cells at 37℃ in 5% CO2 humidified atmosphere. 5×105 cells/well were seeded into six-well plate and lentiviral vectors with CNN2 shRNA sequence or NC shRNA sequence were added at a multiplicity of infection (MOI) of 50 into the plate when the cells reached 70% confluency. The medium was replaced after 24 h of infection and the efficiency of transfection was evaluated by inverted fluorescence microscope. Then the cells were screened with puromycin (2 μg/ml) for three days.
Animals
The Experimental Animal Center of Guangxi Medical University provided Balb/C nude mice. All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Quantitative real-time PCR
SK-hep-1 cells transfected with shRNA and SK-hep-1 cells transfected with NC shRNA were collected and total RNA kit (Omega, USA) was used for total RNA extraction. PrimeScriptTM RT reagent Kit with gDNA Eraser (TAKARA, Japan) was used for cDNA synthesis. SYBR Premix Ex TaqTM Ⅱ (TAKARA, Japan) was used for qRT-PCR. All the procedures were carried out according to each manufacturer's instructions. Primer sequences for CNN2 are 5'- GGTCAAGGCCATATCCCAATAC-3' (sense) and 5'-GGCATAGAAACCACAAACTGCTC-3' (anti-sense). Primer sequences for GAPDH are 5'-TGCACCACCAACTGCTTAGC-3' (sense) and 5'-GGCATGGACTGTGGTCATGAG-3' (anti-sense). Each PCR consisted of 40 cycles (15s at 95℃, 30s at 55℃, and 30s at 72℃). 2-ΔΔCt method was used to analyze the difference of mRNA expression between the two groups.
Western blot
Cells were lysed by Radio-Immunoprecipitation Assay (RIPA) lysis solution (Beyotime, China) mixed with 1mM phenylmethyl sulfonylfluoride (PMSF) (Beyotime, China). Total protein concentration was measured by bicinchoninic acid (BCA) protein assay kit (Beyotime, China) with bovine serum albumin as the standard protein. Protein samples were mixed with loading buffer and bathed in 100℃ water for 5 min. Then the mixture were loaded and separated on 12% SDS-polyacrylamide gels (Beyotime, China) followed by electrotransferation using polyvinylidene difluoride (PVDF) membranes (Bio-rad, CA). Nonspecific binding sites were blocked with 5% nonfat milk. Α-Tublin was used as internal reference. PVDF membranes were incubated with primary antibodies (Santa, USA), then with secondary Horseradish Peroxidase (HRP)-conjugated antibodies (TransGen Biotech, China). After treated with electrochemiluminescence (ECL) analysis kit (PPLYGEN, China), the proteins on the membranes can be seen by developing the film. Then, the grey values were analyzed by software image J.
Cell migration and invasion assays
Cells of each group were collected and diluted to about 1.5×105 Cells/ml by serum free DMEM. 200 μl cell suspension were added into the upper chamber and 800 μl DMEM mixed with 10% FBS was added into the lower chamber. In invasion assay, matrigel (BD, USA) was diluted by serum free DMEM (1:8) and added into the upper chamber before cell seeding. The chambers (Corning Costar, USA) were incubated at 37℃ in carbon dioxide cell incubator for 24 h. The medium was discarded and the cells on the inner surface of membrane were swabbed by cotton buds. The cells on the lower surface of membrane were dyed with 0.1% crystal violet for 10 minutes and then washed twice using ddH2O. Cells were counted by 200× magnification light microscope (Olympus, Japan) in five random fields. The average number of cells can evaluate the abilities of migration and invasion.
Cell proliferation assay
96-well plates (3000 cells/well) were used to seed the cells and incubated for 4 days (0, 24, 48, 72, 96 h). 10μl 3-(4,5-dimethyl-2-thiazolyl)-2,5-diphenyl-2-H-tetrazolium bromide (MTT) (Solarbio, China) was then added to each well. After 4 h of incubation in cell incubator, MTT was discarded and DMSO was added to dissolve formazan. The absorbance of formazan was measured daily by microplate reader at 490nm. Each assay was performed in triplicate.
Flow cytometry analysis of cell cycle
Cells were collected and washed by cold PBS for twice followed by fixing in 70% ice cold ethanol at 4℃ overnight. The cells were then centrifuged and re-suspended by 400µl cold PBS containing 20µl RNase A Solution (Vazyme Biotech, China). The mixture was kept in water bath at 37℃ for half-hour. A 400µl PI Staining Solution (Vazyme Biotech, China) was then added. After incubation at 4℃ for 30min, the mixture was analyzed using flow cytometry (Roche, Switzerland).
Xenografts in nude mice
The 4-6 weeks old, 15-20g weight Balb/C nude mice were randomly divided into two groups. 12 mice per group and half of the mice are male and half are female in each group. SK-hep-1 cells transfected with shRNA and SK-hep-1 cells transfected with NC shRNA were inoculated into the right armpit of nude mice (1×106 cells/mouse) subcutaneously. Tumor volumes were measured once a week by vernier calipers and calculated using formula: 1/2 × length × width2. The nude mice were sacrificed and dissected to observe the presence of metastatic tumors in organs at the fourth week. Xenotransplanted tumors were isolated and fixed in 4% paraformaldehyde (Solarbio, China). Paraffin was used to embed tumor tissues.
H & E staining and Tunel assay
Paraffin sections were dewaxed, hydrated. Cells were stained by hematoxylin (Nobleryder, China), counterstained by eosin (Nobleryder, China) and mounted by neutral balata. Cells Apoptosis were evaluated by Tunel Assay Kit (Roche, Switzerland) according to the manufacturer's instructions.
Immunohistochemistry
Paraffin sections were dewaxed and hydrated. Antigen were retrieved by citrate buffer and blocked with 3% H2O2 at room temperature for 15min. Lowlenthal serum was used to block nonspecific binding sites for 15min. Slides were incubated with diluted primary antibody (1:600) at 4°C overnight followed by secondary antibody for 20min at room temperature, and then incubated in streptavidin-horseradish peroxidase (SA-HRP) solution for 20min. The slides were stained by diaminobenzidine (DAB) (ZSGB-BIO, China) and hematoxylin and then mounted by neutral balata. Pictures were taken by 200× magnification light microscope (Olympus, Japan). The results were analyzed using software imagepro plus.
Statistical analysis
In this study, the SPSS 16.0 software was adopted for statistical analyses. The obtained data were expressed as mean ± standard deviation (SD). Student's t-tests were used for comparisons among groups and the P-value of less than 0.05 was considered having statistical difference.
Results
CNN2 shRNA significantly down-regulates the expression of CNN2 in SK-Hep-1 cells
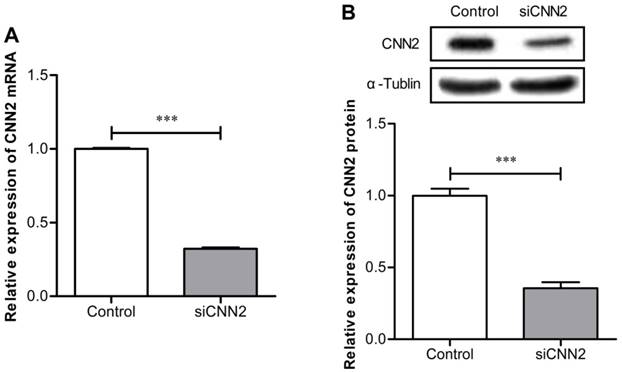
The expression of CNN2 mRNA has been evaluated by qRT-PCR. As can been seen in Figure 1A, comparing with control group, the expression of CNN2 mRNA is significantly down-regulated in SK-hep-1 cells transfected with shRNA (P < 0.01). The expression of CNN2 protein have been evaluated by Western Blot. As shown in Figure 1B, when comparing with the control group, expression of CNN2 protein is markedly down-regulated in SK-hep-1 cells transfected with shRNA (P < 0.001). The above results indicate that lentiviral vector with CNN2 shRNA sequence can efficiently inhibit the expression of CNN2.
Knockdown of CNN2 inhibits cell migration and invasion
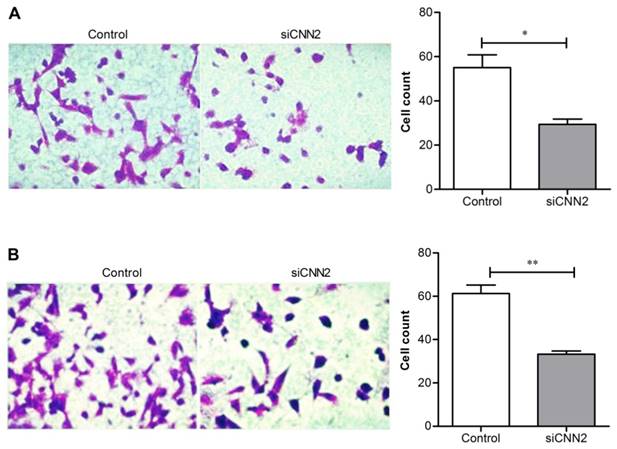
Transwell assay has been used to evaluate cell migration and invasion. Figure 2A shows that comparing with control group, the migration ability of cells transfected with CNN2 shRNA is significantly inhibited (29.33 ± 4.16 versus 55.00 ± 10.10, P < 0.05). Figure 2B shows that comparing with control group, the invasion ability of cells transfected with CNN2 shRNA is significantly inhibited (33.33 ± 2.52 versus 61.33 ± 6.66, P < 0.01). The above results indicate that lentiviral vectors with CNN2 shRNA sequence can effectively inhibit the migration and invasion abilities of SK-hep-1.
Knockdown of CNN2 inhibits cell proliferation
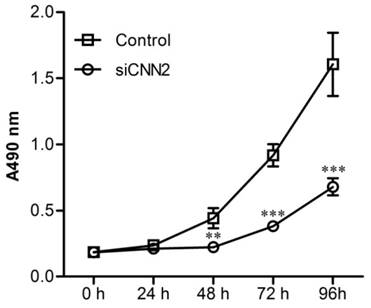
Cell proliferations have been analyzed using MTT assay. Figure 3 shows that after incubation in 96-well plates for 48 h, 72 h and 96 h, cells in siCNN2 group are significantly less than those in control group. The above results indicate that lentiviral vectors with CNN2 shRNA sequence effectively inhibit the proliferation abilities of SK-hep-1.
Knockdown of CNN2 blocks cell cycle at S-phase
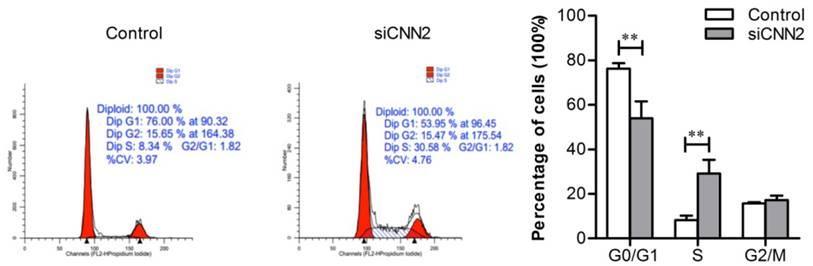
In order to understand whether CNN2 shRNA inhibits cell proliferations through regulating cell cycles, we have employed flow cytometry to analyze the distributions of cell cycles. As shown in Figure 4, the percentage of S phase cells in siCNN2 group is higher than those in control group (P < 0.01) and the percentage of G0/G1 phase cells in siCNN2 group is lower than those in control group (P < 0.01). The above results indicate that lentiviral vectors with CNN2 shRNA sequence inhibit proliferation through blocking cell cycles at S-phase.
The effect of shRNA on CNN2 knockdown. (A) Expressions of CNN2 mRNA are significantly down-regulated in SK-hep-1 cells treated with CNN2 shRNA compared with control group. (B) Expressions of CNN2 protein are significantly down-regulated in SK-hep-1 cells treated with CNN2 shRNA compared with control group (***P < 0.001).
The effects of CNN2 knockdown on cell migration and invasion. (A) The migration ability is significantly weakened in SK-hep-1 cells treated with CNN2 shRNA compared with control group. (B) The invasion ability is significantly weakened in SK-hep-1 cells treated with CNN2 shRNA compared with control group (*P < 0.05, **P < 0.01).
The effect of CNN2 knockdown on cell proliferation. The proliferation ability is significantly weakened in SK-hep-1 cells treated with CNN2 shRNA compared with control group (**P < 0.01, ***P < 0.001).
Knockdown of CNN2 inhibits xenotransplanted tumor growth of nude mice
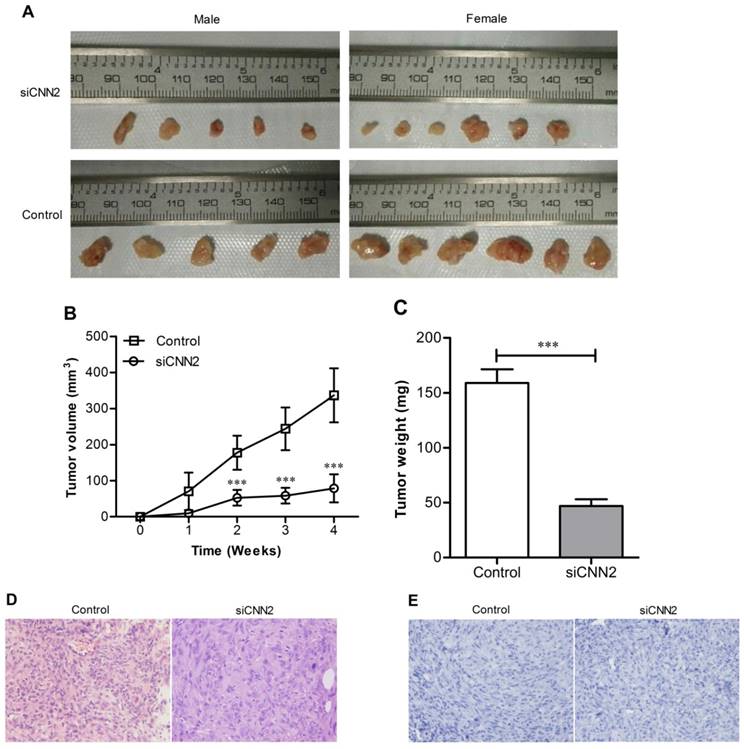
Tumors have been formed at the site of inoculation within 7 to 14 days in all tested mice except one male nude mice in siCNN2 group. Tumor volumes and weights were measured once a week. Tumor volumes in siCNN2 group are lower than those in control group (siCNN2 group 78.98 ± 39.2 mm3, control group 336.51 ± 74.7 mm3, P < 0.001) (Figure 5A and 5B). There has been no difference of mice weights between siCNN2 group and control group. Mice have been sacrificed at the fourth week and tumor weights were measured. As shown in Figure 5C, tumor weights in siCNN2 group are lower than in control group (siCNN2 group 47.00 ± 21.28 mg, control group 159.00 ± 43.1 mg, P < 0.001). Mice were all dissected but no metastatic tumor was found in mice organs such as lung and liver. H & E staining have shown typical characteristics of tumor, that is, the nuclei are large, deeply stained and cells are closely arranged (Figure 5D). There is no cell apoptosis being detected (Figure 5E). The above results indicate that lentiviral vectors with CNN2 shRNA sequence can inhibit xenotransplanted tumor growth, which means that down-regulation of CNN2 may be beneficial to the treatment of liver cancer.
Knockdown of CNN2 inhibits the expression of phosphorylated MEK1/2 and phosphorylated ERK1/2
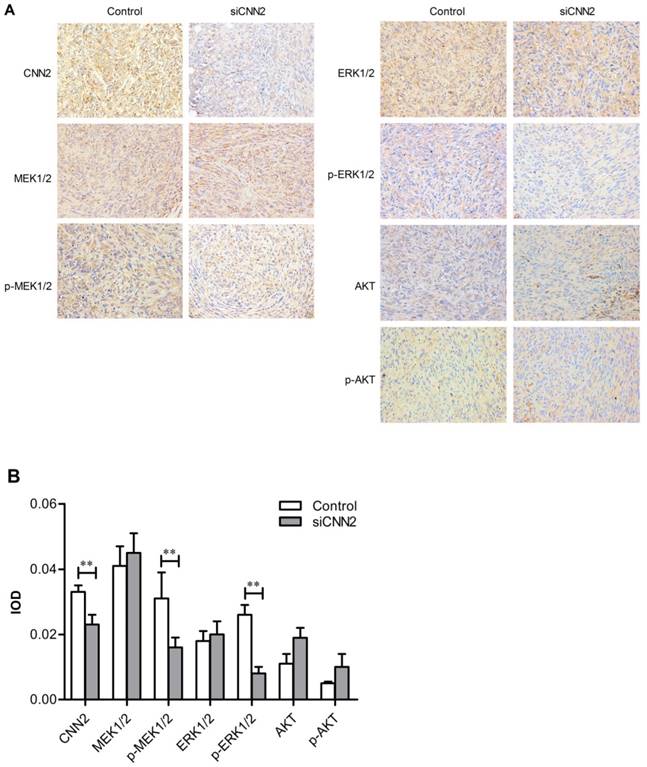
Immunohistochemical technique has been used to evaluate the expression of CNN2, MEK1/2, ERK1/2, AKT, phosphorylated MEK1/2, phosphorylated ERK1/2, and phosphorylated AKT. Pictures have been taken by 200× magnification light microscope (Figure 6A). As shown in Figure 6B, the expression of CNN2, phosphorylated MEK1/2 and phosphorylated ERK1/2 are down-regulated in siCNN2 group, while the others show no difference between siCNN2 group and control group (PCNN2 < 0.05, Pp-MEK1/2 < 0.05, P p-ERK1/2 < 0.01). The above results indicate that lentiviral vectors with CNN2 shRNA sequence may inhibit tumor growth through regulating pMEK1/2 and pERK1/2 expression.
Discussion
In this study, CNN2 in SK-hep-1 cells have been knockdown using lentivirus-mediated CNN2 shRNA. The expression of CNN2 mRNA and protein have been efficiently down-regulated. Meanwhile, cell migration, invasion, proliferation and xenotransplanted tumor growth have been evidently inhibited. From the results of immunohistochemistry, we know that CNN2 might be associated with MEK/ERK.
The effect of CNN2 knockdown on cell cycle. The distributions of G0/G1 phase are decreased and the distributions of S phase are increased in SK-hep-1 cells treated with CNN2 shRNA compared with control group (**P < 0.01).
The effect of CNN2 knockdown on xenograft tumor growth. (A) The tumor images represent excised tumors from each group. (B) Xenograft tumor growth was significantly inhibited in SK-hep-1 cells treated with CNN2 shRNA compared with control group. (C) The weight of tumor was lighter in SK-hep-1 cells treated with CNN2 shRNA compared with control group. (D) H&E staining of tissue section of tumor. (E) Tunel staining of tissue slices (***P < 0.001).
The CNN2 has been first isolated from bovine aortic smooth muscles and chicken gizzard [12]. In 1996, human CNN2, a neutral protein, which has high homology with the mouse CNN2, has been discovered by Hiroaki Masuda. It exists in smooth muscle cells, and also exists in non muscle cells such as fibroblasts, endothelial cells and keratinocytes [13]. The C-terminal tandem repeats of CNN2 have been demonstrated to participate in actin binding, indicating that CNN2 plays a role in regulating cell movement [14]. Current studies have indicated that CNN2 is up-regulated in a variety of cancers, such as cutaneous squamous cell carcinoma, rectal cancer and breast cancer [15-17]. Tianxing Ji and Debald M have detected CNN2 content in the serum of breast cancer patients, breast benign disease patients and healthy women, found that CNN2 content in the serum of breast cancer patients is the highest [18, 19]. Tianxing Ji has also demonstrated that combination of CNN2, CA15-3 and CEA may increase the diagnostic accuracy of breast cancer. However, another study has reported that CNN2 expression in prostate cancer cells is down-regulated, and its expression in high metastatic prostate cancer cells is lower than those in low metastatic prostate cancer cells [20].That means CNN2 has been proved to inhibit cell proliferation and metastasis in prostate cancer cells, contrary to the roles of CNN2 in hepatocellular carcinoma in our studies. It is speculated that CNN2 may play variable roles in different types of cancers, and this might explain the differences in CNN2 expression levels as well as the differences in cell biological behaviors.
It has been proved that the occurrence and development of HCC is correlated with RAS/RAF/MEK/ERK signal pathway, Wnt/β-catenin signal pathway, PI3K/AKT/mTOR signal pathway, etc [21-24]. Activations of those pathways will inhibit apoptosis and promote cell proliferation, angiogenesis, cell metastasis [25-28]. At the present, it is still little known about the relationship between CNN2 and signal pathway. Tang J's research has found that down-regulation of CNN2 expression can inhibit migration ability of endothelial cells in vascular formation [10]. The mechanisms may be that CNN2 can up-regulate the phosphorylation level of ERK1/2 in the MAPK pathway. In the present work, we have also proved that down-regulation of CNN2 expression decrease the expression of phosphorylated MEK1/2 and phosphorylated ERK1/2, while has no influence on phosphorylated AKT, AKT, MEK1/2, and ERK1/2. In conclusion, our research has first demonstrated that down-regulation of CNN2 by lentivirus-mediated shRNA can inhibit SK-hep-1 cells proliferation, metastasis and xenotransplanted tumor growth. Moreover, CNN2 might regulate the development of hepatocellular carcinoma by interacting with MAPK pathway. It seems that CNN2 may be a new potential target for HCC therapy.
Immunohistochemical results. (A) Representative images from immunohistochemical analysis. (B) Integrated optical density of CNN2, MEK1/2, p-MEK1/2, ERK1/2, p-ERK1/2, AKT and p-AKT (**P < 0.01).
Acknowledgements
This work is supported by National Natural Science Foundation of China (81572994, 81460432); Guangxi Nature Science Foundation (2015GXNSFDA139017, 2014GXNSFBA118206); Guangxi Science and Technology Research and Technology Development Project (15104001-7); Programs for Department of Education of Guangxi Zhuang Autonomous Region (2013ZD011); Guangxi Nanning Qingxiu District Science and Technology Research and Technology Development Project (2014S03).
Compliance with ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Ferlay J, Shin HR, Bray F. et al. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer. 2010;127:2893-2917
2. Chu KK, Cheung TT. Update in management of hepatocellular carcinoma in Eastern population. World J Hepatol. 2015;7:1562-1571
3. Song T. Recent advances in surgical treatment of hepatocellular carcinoma. Drug Discov Ther. 2015;9:319-330
4. Sakon M, Nagano H, Nakamori S. et al. Intrahepatic recurrences of hepatocellular carcinoma after hepatectomy: analysis based on tumor hemodynamics. Arch Surg. 2002;137:94-99
5. Husain SR, Han J, Au P. et al. Gene therapy for cancer: regulatory considerations for approval. Cancer Gene Ther. 2015;22:554-563
6. Khare S, Zhang Q, Ibdah JA. Epigenetics of hepatocellular carcinoma: Role of microRNA. World J Gastroenterol. 2013;19:5439-5445
7. Pogribny IP, Rusyn I. Role of epigenetic aberrations in the development and progression of human hepatocellular carcinoma. Cancer Lett. 2014;342:223-230
8. Giordano S, Columbano A. Met as a therapeutic target in HCC: facts and hopes. J Hepatol. 2014;60:442-452
9. Hossain MM, Hwang DY, Huang QQ. et al. Developmentally regulated expression of calponin isoforms and the effect of h2-calponin on cell proliferation. Am J Physiol. Cell Physiol. 2003;284:C156-C167
10. Tang J, Hu G, Hanai J. et al. A Critical Role for Calponin 2 in Vascular Development. J Biol Chem. 2006;281:6664-6672
11. Huang QQ, Hossain MM, Wu K. et al. Role of H2-calponin in Regulating Macrophage Motility and Phagocytosis. J Biol Chem. 2008;283:25887-25899
12. Winder SJ, Walsh MP. Calponin: thin filament-linked regulation of smooth muscle contraction. Cell Signal. 1993;5:677-686
13. Masuda H, Tanaka K, Takagi M. et al. Molecular cloning and characterization of human non-smooth muscle calponin. J Biochem. 1996;120:415-424
14. Gimona M, Mital R. The single CH domain of calponin is neither sufficient nor necessary for F-actin binding. J Cell Sci. 1998;111:1813-1821
15. Dang C, Gottschling M, Manning K. et al. Identification of dysregulated genes in cutaneous squamous cell carcinoma. Oncol Rep. 2006;16:513-519
16. Choi SY, Jang JH, Kim KR. Analysis of differentially expressed genes in human rectal carcinoma using suppression subtractive hybridization. Clin ExpMed. 2011;11:219-226
17. Debald M, Franken S, Heukamp LC. et al. Identification of specific nuclear structural protein alterations in human breast cancer. J Cell Biochem. 2011;112:3176-3184
18. Ji T, Ma F, Huo L. et al. Calponin-h2 is upregulated in the tissues and plasma of patients with breast cancer. Mol Med Rep. 2015;12:2886-2892
19. Debald M, Jin JP, Linke A. et al. Calponin-h2: a potential serum marker for the early detection of human breast cancer?. Tumor Biol. 2014;35:11121-11127
20. Moazzem Hossain M, Wang X, Raymond C. Bergan, et al. Diminished expression of h2-calponin in prostate cancer cells promotes cell proliferation, migration and the dependence of cell adhesion on substrate stiffness. FEBS Open Bio. 2014;4:627-636
21. Delire B, Stärkel P. The Ras/MAPK pathway and hepatocarcinoma: pathogenesis and therapeutic implications. Eur J Clin Invest. 2015;45:609-623
22. Vilchez V, Turcios L, Marti F. et al. Targeting Wnt/β-catenin pathway in hepatocellular carcinoma treatment. World J Gastroenterol. 2016;22:823-832
23. Wang Z, Zhang G, Wu J. et al. Adjuvant therapy for hepatocellular carcinoma: current situation and prospect. Drug Discov Ther. 2013;7:137-143
24. Choi KJ, Baik IH, Ye SK. et al. Molecular Targeted Therapy for Hepatocellular Carcinoma: Present Status and Future Directions. Biol Pharm Bull. 2015;38:986-991
25. Chen Y, Liu Q, Wu M. et al. GAB2 promotes cell proliferation by activating the ERK signaling pathway in hepatocellular carcinoma. Tumour Biol. 2016;37:11763-11773
26. Fan WH, Du FJ, Liu XJ. et al. Knockdown of FRAT1 inhibits hypoxia-induced epithelial-to-mesenchymal transition via suppression of the Wnt/β-catenin pathway in hepatocellular carcinoma cells. Oncol Rep. 2016;36:2999-3004
27. Sui Y, Zheng X, Zhao D. Rab31 promoted hepatocellular carcinoma (HCC) progression via inhibition of cell apoptosis induced by PI3K/AKT/Bcl-2/BAX pathway. Tumour Biol. 2015;36:8661-8670
28. Zhang XL, Jia Q, Lv L. et al. Tumorspheres derived from HCC cells are enriched with cancer stem Cell-like cells and present high chemoresistance dependent on the Akt pathway. Anticancer Agents Med Chem. 2015;15:755-763
Author contact
![]() Corresponding authors: Sufang Zhou; Email: zsf200000com. Yongxiang Zhao; Email: yongxiang_zhaocom
Corresponding authors: Sufang Zhou; Email: zsf200000com. Yongxiang Zhao; Email: yongxiang_zhaocom
Received 2017-5-20
Accepted 2017-9-11
Published 2018-1-1
