ISSN: 1449-1907International Journal of Medical Sciences
Int J Med Sci 2017; 14(12):1189-1196. doi:10.7150/ijms.20721 This issue Cite
Research Paper
Association of Surfactant Protein with Expression of Hoxa5 and Hoxb5 in Rabbit Fetal Lung
Department of Pediatrics, Kyung Hee University School of Medicine, Seoul, Korea
Abstract
Hox genes regulate organ formation and identity of the embryo, and expressed in specific temporo-spatial patterns in the developing embryo. We compared the expression levels of the Hoxa5, Hoxb5, surfactant protein (SP)-A, and SP-B genes in immature and mature rabbit fetal lung tissues, and to uncover roles for Hoxa5, Hoxb5, SP-A, and SP-B. Cesarean sections were performed after rabbits were divided into two groups of 30-31 days of gestation (term group, n = 24) and 26-27 days of gestation (preterm group, n = 24). mRNA levels of Hoxa5, Hoxb5, SP-A, and SP-B were compared by quantitative reverse transcriptase polymerase chain reaction, and protein expression of Hoxa5 and Hoxb5 was compared by western blot analysis. Fetal lung tissue histology was observed by hematoxylin and eosin (H&E) staining. The relative expression ratios of SP-A and SP-B mRNA in the term to preterm groups were 2.45:1 and 2.94:1, respectively. Hoxb5 mRNA and protein levels decreased in the term group, with a relative expression ratio of 0.48:1 and 0.50:1, however, Hoxa5 mRNA and protein levels increased in the term group with a relative expression ration of 2.99:1 and 2.33:1, respectively, for the term to preterm groups. Moreover, a significant positive correlation was found between the expression of Hoxa5 and SP-A, SP-B in the term group. Hoxa5 gene may be essential for the expression of SP-A and SP-B in term rabbits. The Hoxb5 gene may be an important factor for lung maturation in preterm rabbits.
Keywords: Hox genes, lung development, surfactant protein-A, surfactant protein-B, quantitative reverse transcription polymerase chain reaction, western blot.
Introduction
Hox genes play a pivotal role in the formation of body structure, while controlling the formation of organs, and they are related to the expression of the other genes [1, 2]. In mammals, including humans, there are four Hox clusters (A, B, C, and D), which were formed by duplications of a single ancestral cluster. The 39 Hox genes within these clusters fall into 13 paralogous groups based on sequence similarity. Hox genes specify the identity of the body segments and structures where they are expressed, and mutations result in homeotic transformation to some other fate [3-5].
Airway branching is one of many processes required for lung development. Airways appear to set arterial and smooth muscle patterns, and signals from the airways likely have a decisive effect on airway cell differentiation and alveolus formation. Therefore, the airway branching program is a pivotal stage of full lung development [6]. From a morphogenetic point of view, such as proximal-distal, anterio-posterior, and left-right stereotypic patterns of airway branching, Hox genes might play an important role in branching morphogenesis, epithelial cell fate, and differentiation of the conducting airway epithelium [7, 8].
In fact, Hox gene mutations cause serious functional and structural respiratory problems in animal models. Hoxa1 and Hoxa3 mutant mice die after birth owing to pulmonary failure, and Hoxa5 homozygous mutants die after birth because of improper tracheal and lung morphogenesis. Mutations in Hoxa5 and Hoxb5 are associated with the occurrence of bronchopulmonary sequestration, congenital cystic adenomatoid malformation tissue, primary pulmonary hypertension, and emphysema [9-13].
Despite growing evidence linking Hox genes to lung embryogenesis and development, information regarding the expression of Hox genes in lung maturation, such as the production of surfactant protein (SP) is still lacking. In the present study, we surveyed the correlation of expression between the Hox genes and SP-A, SP-B in preterm and term rabbit fetal lungs. The abundant Hox genes identified in normal lung, Hoxa5 and Hoxb5, were the subjects of this expression study.
Materials and methods
Animal protocol
All procedures were approved by the Committee on Animal Research and Ethics of Kyung Hee University Hospital at Gangdong (KHNMC AP 2013-008). Twelve pregnant New Zealand white rabbits delivered their newborn rabbits by cesarean section after being anesthetized with 15 mg/kg of Zoletil® (tiletamine + zolazpam) injected into the auricular vein. A total of 48 fetuses were used, and grouped according to gestational age 26-27 days, corresponded to the canalicular stage of lung development (preterm group) and 30-31 days, which represented the alveolar stage (term group). We fixed the fetuses in a supine position, cut the trachea, inserted an 18-gauge angiocatheter into trachea, and positioned the tip of the angiocatheter above the carina with an elastic thread. The lungs then were ventilated several times manually using a syringe, and were dissected free and removed en bloc from the fetus. Next, we compared histological difference of lung tissues by staining with hematoxylin and eosin (H&E, Thermo-Scientific, Waltham, MA) to confirm the proper identification of groups (Fig. 1) and measured the aeration areas using NIH ImageJ program (Fig. 2). The lung tissues of newborn rabbits were dissected and frozen quickly in liquid nitrogen for total RNA and protein extraction (n = 24 of preterm group and n = 24 of term group).
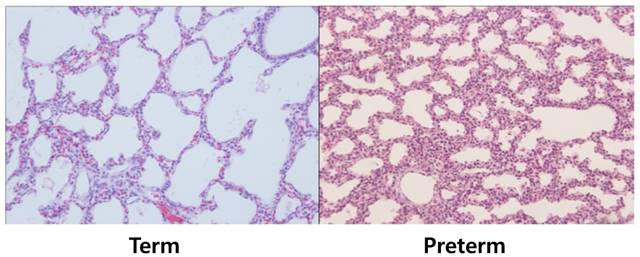
Histologic differences among two groups (hematoxylin and eosin stain, ×100). Lung tissues of term groups demonstrated better aerated features than preterm controls. For statistical analysis, alveolar areas were compared and revealed significant differences, as shown in Table 1.
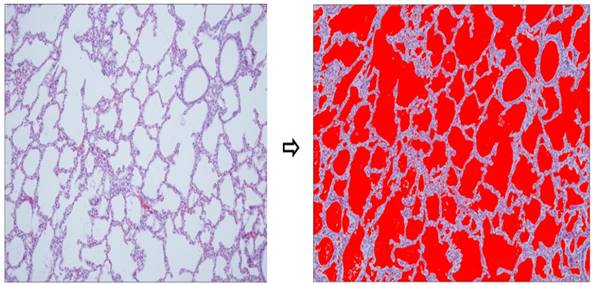
Aerated area percent using imageJ. Lung tissues of term group (n = 24) demonstrated significantly higher aerated area ratio than preterm controls (n = 24, p < 0.001).
RNA extraction
Total RNA was extracted from rabbit fetal lung tissues using an RNeasy Mini Kit (Qiagen, Valencia, CA, USA) according to the manufacturer's instructions. Briefly, samples that were frozen at -70°C were purified by RLT buffer (guanidine-thiocyanate-containing lysis buffer) and centrifuged at 15,000 rpm for 3 min at room temperature. The aqueous phase was carefully removed using a pipette. Seventy percent ethanol (1:1) was added to the tube, and mixed gently with a pipette. The solution (700 μL) was added to an RNeasy Spin Column and the column was centrifuged at 9,200 rpm (8,000 x g) for 15 s. The flow-through was discarded. Buffer RW1 (350 μL) was added to the RNeasy Spin Column, which then was centrifuged, and then the flow-through was discarded. A DNase I stock solution (10 μL) was added to 70 μL Buffer RDD, and the solution was mixed by inversion and then centrifuged briefly. The DNase I incubation mix (80 μL) then was added directly to the column membrane, which subsequently was incubated on the benchtop (20-30°C) for 15 min. Buffer RW1 (350 μL) was added to the RNeasy Spin Column, which was then centrifuged for 15 s at 8,000 x g, and the flow-through was discarded. A total of 250 μL ethanol (96-100%) was added to the diluted RNA, and mixed by pipetting. The sample was then transferred to an RNeasy Mini Spin Column placed in a 2 mL collection tube, which was centrifuged for 15 s at 8000 x g, and the flow-through was discarded. Buffer RPE (500 μL) was added to the RNeasy Spin Column, which was centrifuged for 15 s at 8000 x g to wash the membrane. After the flow-through was discarded, another 500 μL Buffer RPE was added to the RNeasy Spin Column, and the column was centrifuged for 2 min at 8000 x g to dry the membrane. The RNeasy Spin Column was placed in a new 1.5 mL collection tube, 30 μL RNase-free water was added directly to the spin column membrane, then the column was centrifuged for 1 min at 8000 x g to elute the RNA. The ratio of absorbance at 260 nm and 280 nm was used to assess the purity of RNA, which ranged from 1.8 to 2.0 when measured by a NanoDrop spectrophotometer (Thermo Fisher Scientific Inc., DE, USA).
Primer design
Primers for Hoxa5 and Hoxb5 were designed from the genomic sequences for Hoxa5 and Hoxab5 of domestic rabbit (GenBank; accession no. NW_003159267.1) using the software Primer 3 (http://frodo.wi.mit.edu/primer3/) and designed four primer pairs for each. The genes were amplified by polymerase chain reaction (PCR): Hoxa5 (sense: 5′-CGT GCT TCT ATT CCG ATA AG-3′; antisense: 5′-TGC TGT CGG GTT TGT ACT G-3′); Hoxb5 (sense: 5′-CCT GTA CCA GAG CCC CTT C-3′; antisense: 5′-CCT CGT CCG CCT TGT C-3′). We used commercial primers for SP-A and SP-B. The size of the PCR products for Hoxa5, Hoxb5, SP-A, SP-B were 117, 110, 133, and 208 bp, respectively, and the annealing temperature were 60.0˚C, 62.0˚C, 57.0˚C, and 55.0˚C, respectively.
Quantitative reverse transcription PCR (qRT-PCR) experiments
qRT-PCR was performed to quantitate mRNA expression of SP-A, SP-B, Hoxa5, and Hoxb5 in rabbit lungs using SYBR® Green I, according to the manufacturer's instructions. Reactions were carried out using Power SYBR Green PCR Master Mix (Applied Biosystems, Warrington, UK) on the StepOne Plus Real-Time PCR System (Applied Biosystems, Foster City, CA, USA) by denaturation at 95°C for 10 min, followed by 35 cycles at 56.5°C for 20 s and 72°C for 30 s. Melting curve analyses were performed to verify the amplification specificity. Relative quantification of gene expression was performed using the ΔΔ-CT method using StepOne Software 2.0 (Applied Biosystems). Reactions were analyzed in triplicate and mRNA expression levels were normalized to 18S. Relative quantification of mRNA expression was performed using the 2-∆∆CT method.
Western blot analysis for Hoxa5/b5
Extracted preterm and term rabbit fetal lung tissues were lysed in RIPA buffer (150 mm NaCl, 0.1% SDS, 1% triton X-100, 2 mM EDTA, 1% sodium deoxycholate, Tris-HCl pH 7.5) supplemented with a proteinase inhibitor cocktail tablet (R&D system, USA) and centrifuged at 12,000 rpm for 15 min at room temperature. The top layer was transferred to a new tube and then placed on ice. A NanoDrop spectrophotometer was used to determine the concentration of purified protein samples. Each sample was measured twice, and the average was calculated. Protein samples were diluted into 5X SDS loading buffer, heated to 95°C for 5 min, rapidly cooled, and then stored in a freezer. Protein extracts were separated on 8% sodium dodecyl sulfate-polyacrylamide gels and transferred to PVDF membranes (Pall Life Science, transfer membrane 0.45 um). The membranes were incubated with primary human antibodies specific for Hox (Abcam®, Cambridge, UK), since rabbit antibodies are not yet commercially available. Secondary antibodies were conjugated with peroxidase-linked mouse antibody (1:2000 dilution; Santa Cruz Biotechnology Inc., Santa Cruz, CA, USA) at 4°C after water bath with Tris-buffered saline (TBS; 0.05M TBS, 0.05% Tween 20). β-actin was used as a loading control. The protein expression levels of Hoxa5 and Hoxb5 were analyzed with the NIH ImageJ program, and were expressed as fold changes compared to the levels of preterm rabbits.
Statistical analysis
Statistical analysis was performed using the Student's t‑test, Spearman's correlation coefficient and linear regression.
Results
Characteristics of fetal rabbits and morphological comparison of rabbit lung tissue
Newborn rabbits were delivered from twelve pregnant New Zealand white rabbits by cesarean section at the gestational age of 26.3 ± 0.6 days (preterm group) and 30.1 ± 0.3 days (term group). The birth weights of the two groups were 40.4 ± 5.9 g and 63.6 ± 9.9 g, respectively. Aerated area ratios of the two groups were 0.36 (preterm group) and 0.59 (term rabbit). These results showed statistically significant differences between the two groups (Table 1). Lung tissues of term group demonstrated significantly higher aerated area ratio than preterm group.
Characteristics of fetal rabbits
| Preterm group (n = 24) | Term group (n = 24) | p - value | |
|---|---|---|---|
| Gestational age (days) | 26.1 ± 0.6 | 30.3 ± 0.3 | < 0.01 |
| Birth weight (gram) | 40.4 ± 5.9 | 63.6 ± 9.9 | < 0.01 |
| Aerated area ratio | 0.36 ± 0.12 | 0.59 ± 0.07 | < 0.01 |
Comparison of mRNA expressions of SP-A/B in rabbit lung tissue
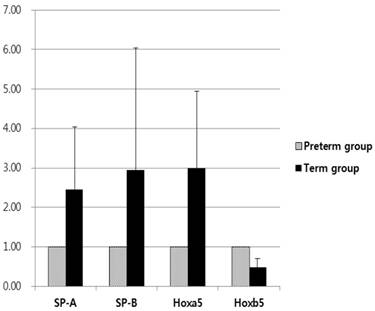
When the mRNA levels of SP-A and SP-B of the preterm group were set to a value of 1, the relative ratio of mRNA levels in the term group significantly increased to 2.45 and 2.94, respectively (p < 0.01, Fig. 3). There were no differences in the mRNA levels of SP-A and SP-B when considering birth order or location of lung tissues in fetal rabbits from the same mother. SP-A and SP-B expression in the term group was higher than that of the preterm group.
Comparison of mRNA expression and protein levels of Hoxa5/b5 in rabbit lung tissue
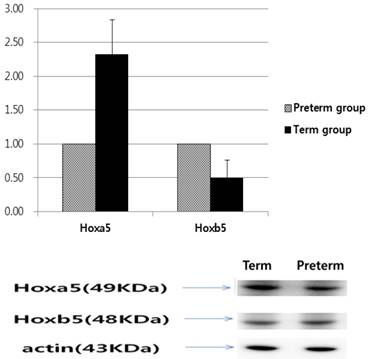
Based on qRT‑PCR to measure mRNA levels, Hoxa5 mRNA levels of the term group were 2.99 times more than that of the preterm group; however, Hoxb5 mRNA levels of the term group were 0.48 times less than that of the preterm group (p < 0.01, Fig. 3). The relative protein levels of Hoxa5 and Hoxb5 were tested by western blot analysis. When the protein levels of Hoxa5 and Hoxb5 of the preterm group were set to a value of 1, the relative ratio of protein levels of Hoxa5 was increased to 2.33 in the term group, while that of Hoxb5 significantly decreased to 0.50 in the term group (p < 0.01, Fig. 4). The protein levels of Hoxa5 were higher, and Hoxb5 were lower in the term group, corresponding with the changes in mRNA levels. Hoxb5 mRNA and protein levels were lower in the term group, but Hoxa5 mRNA and protein levels were higher in the term group than preterm group.
Relative expression levels of SP-A, SP-B, Hoxa5 and Hoxb5 mRNA were measured by quantitative real-time reverse transcription-polymerase chain reaction for quantification of mRNA expression. The expression of SP-A, SP-B, and Hoxa5 was 2.45, 2.94, and 2.99 times more in term group than in the preterm group (all p < 0.01). Hoxb5 was decreased 0.48 times less in term group than in preterm group (p < 0.01). Changes in gene expression were calculated using the 2-∆∆CT method.
Western blot analysis of Hoxa5 and Hoxb5. Hoxa5 expression 2.33 higher in the term group (p < 0.01), while Hoxb5 expression was 0.50 lower in the term group than in the preterm group (p < 0.01).
Correlation analysis between expression levels of SP-A/B and the Hoxa5/b5 genes
We found statistical correlations between the mRNA levels of SP‑A and SP‑B, and between Hoxa5 and Hoxb5 through Spearman's rank correlation coefficient, regardless of the group. In the term group, Hoxa5 positively correlated with SP-A and SP-B, but Hoxb5 insignificantly correlated with SP-A and SP-B. There were no correlations between the mRNA levels of these genes in the preterm group (Table 2).
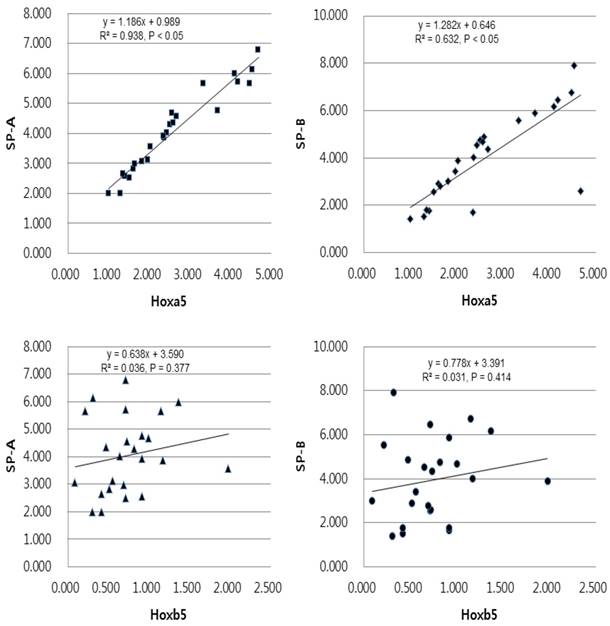
In linear regression analysis between the mRNA levels of Hoxa5 and Hoxb5 and those of SP-A and SP-B, statistically insignificant correlations were found in preterm group (Fig. 5). In term group, however, the regression coefficient among SP-A, SP-B and Hoxa5 were [0.938, p < 0.05, SP-A mRNA expression = 0.989 + 1.186 x Hoxa5 mRNA expression], [0.632, p < 0.05, SP-B mRNA expression = 0.646 + 1.282 x Hoxa5 mRNA expression] and among SP-A, SP-B and Hoxb5 were [0.036, p = 0.377, SP-A mRNA expression = 3.590 + 0.638 x Hoxb5 mRNA expression], [0.031, p = 0.414, SP-B mRNA expression = 3.391 + 0.778 x Hoxb5 mRNA expression] (Fig. 6). We confirmed that changes in Hoxa5 and Hoxb5 mRNA levels do not influence those of SP-A and SP-B in the preterm group, but mRNA levels of Hoxa5 increase those of SP-A and SP-B in the term group.
Pearson's correlation coefficient among SP-A, SP-B, and Hoxa5, Hoxb5 mRNA expression in quantitative real-time reverse transcription-polymerase chain reaction (qRT-PCR)
| Item | SP-A | SP-B | Hoxa5 | Hoxb5 |
|---|---|---|---|---|
| (preterm/term) | (preterm/term) | (preterm/term) | (preterm/term) | |
| SP-A | 1.00/1.00 | |||
| SP-B | 0.705*/0.804* | 1.00/1.00 | ||
| Hoxa5 | 0.165/0.988* | 0.067/0.820* | 1.00/1.00 | |
| Hoxb5 | 0.189/0.274 | -0.096/0.207 | 0.305/0.288 | 1.00/1.00 |
*p < 0.05
Univariate linear regression analysis of the mRNA expression levels of SP-A, SP-B and Hoxa5, Hoxb5 in the preterm group. The equation in the preterm group was [SP-A expression = 0.955 + 0.127 x Hoxa5 expression (r2 = 0.017, and p = 0.542)], [SP-B expression = 1.057 - 0.049 x Hoxa5 expression (r2 = 0.002, and p = 0.857)], [SP-A expression = 0.852 + 0.156 x Hoxb5 expression (r2 = 0.075, and p = 0.197)], [SP-B expression = 1.064 - 0.040 x Hoxb5 expression (r2 = 0.004, and p = 0.765)].
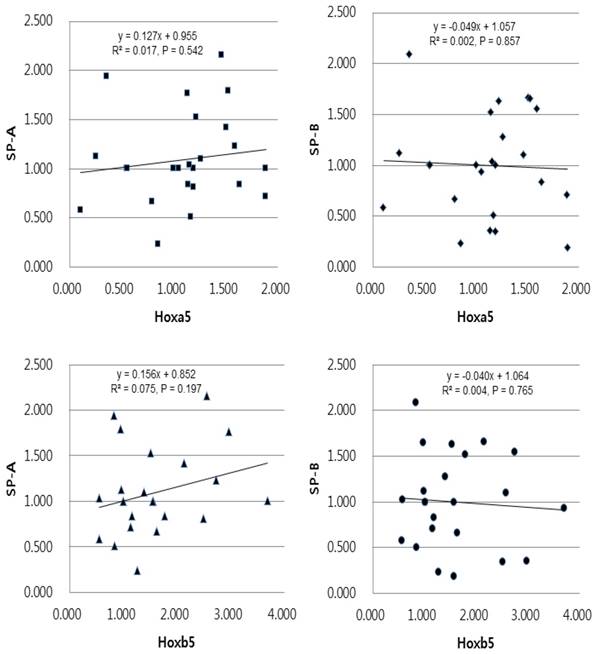
Univariate linear regression analysis of the mRNA expression levels of SP-A, SP-B and Hoxa5, Hoxb5 in the term group. The equation in the term group was [SP-A expression = 0.989 + 1.186 x Hoxa5 expression (r2 = 0.938, and p < 0.05)], [SP-B expression = 0.646 + 1.282 x Hoxa5 expression (r2 = 0.632, and p < 0.05)], [SP-A expression = 3.590 + 0.638 x Hoxb5 expression (r2 = 0.036, and p = 0.377)], [SP-B expression = 3.391 + 0.778 x Hoxb5 expression (r2 = 0.031, and p = 0.414)].
Discussion
New Zealand white rabbits come to full term 30-31 days after fertilization. Because lung development in rabbit is very similar to that in humans, the rabbit lung is considered a suitable model for studies of human lung development. In this study, we conducted an experiment using fetal lungs corresponding to the canalicular stage, at 26-27 days after fertilization, and to the alveolar stage at 30-31 days after fertilization [14-16].
The Hox genes, which are evolutionarily conserved, help to form the structure along the body axis and encode transcription factors that regulate complicated development processes in a number of organs. Hox gene expression may differ according to organ. Among the 13 Hox genes, (Hox1-Hox13), Hox1-Hox5 are involved in the development of the hindbrain, Hox4-Hox11 in the development of the spinal cord, and Hox1-Hox8 in the development of the lung [5, 17]. Volpe et al. reported that among Hox genes, Hoxb5 affects bronchial branching morphogenesis and the differentiation of airway epithelial cells in mice, and cases have been reported where abnormal Hoxb5 expression in humans caused bronchopulmonary sequestration [8, 12, 13]. Golpon et al. reported that Hoxa5 overexpression was associated with pulmonary emphysema [18]. In this way, there have been several reports on lung development and formation, but few studies have focused on SP.
This study compared SP-A and SP-B expression in pulmonary tissue of preterm and term rabbits. According to the data measured by qRT-PCR, mRNA expression was markedly higher in the term group than in the preterm group, and this result was similar to that of other studies. In their studies with rabbits, Durham et al. reported that longer gestational age was associated with higher SP-A, SP-B, and SP-C expression using northern blot analysis, and Grier et al., in studies with rats, reported that SP-B mRNA expression was higher in term fetuses [14, 19]. In this study, we found that the longer the gestational age, the higher the expression of SP-A and SP-B during lung maturation in term fetuses.
Most of previous studies on the roles of Hox genes in fetal lung maturity used rats and mice [19, 20]. However, no study has investigated how Hox genes are involved in human lung maturity, and how they are distributed and expressed. Furthermore, this is the only study that used rabbit lungs, which are similar to human lungs in terms of development process. In the study by Grier et al., 2009, using rats, Hoxa4 expression increased in the early period of lung development and then decreased in the late period, but Hoxa5 expression was continuous, without decreasing, in the late period. The study by Grier et al., and several other studies, also reported that the Hox B cluster, including Hoxb5, showed lower expression level in the late period of lung development compared to that in the early period [17, 19, 21]. In this study as well, Hoxa5 and Hoxb5 expression in pulmonary tissue were compared between the preterm group and the term group, using qRT-PCR and western blot. According to the results, Hoxa5 mRNA and protein expression were higher in the term group than in the preterm group; however, Hoxb5 mRNA and protein expression were lower in the term group. This suggests that Hoxa5 may be involved, in not only fetal lung formation during early pregnancy, but also in surfactant regulation during late pregnancy, and that Hoxb5 may be more involved in the early stage of lung formation than in later stages. Such a variation in Hox expression according to the stage of organ development is not limited to the lung. Tümpel et al. reported that in the hindbrain of rat fetuses, the Hoxa1 and Hoxb1 expression increased at the boundary between rhombomeres 3 and 4 at 7.5-8 days after sexual intercourse, Hoxa1expression decreased while Hoxb1 expression at rhombomere was maintained at 8-8.5 days after sexual intercourse, and Hoxa4, Hoxb4, and Hoxd4 expression at rhombomeres 6 and 7 increased at 8.5-9.5 days after sexual intercourse [22].
This study included a correlation analysis to detect association among the mRNA expression levels of SP-A, SP-B, Hoxa5, and Hoxb5, and the results indicated a strong, positive correlation among SP-A, SP-B, and Hoxa5 in the term group. With mutations in Hoxb5, mice survive well, without showing any major defects in the organs. However, with mutations in Hoxa5, abnormal formation of the organs and lungs occurs, which increases mortality, and a large range of phenotypes, including pulmonary emphysema, is seen [23-26]. This result is probably because Hoxa5 is able to compensate for, and mask any defects in, Hoxb5 [27, 28]. Accordingly, Hoxa5 is believed to play a central role in the lung development of rabbit fetuses.
In another correlation analysis, moreover, the expression levels of Hox genes and SP genes did not correlate with each other in the preterm group, but in the term group, Hoxa5 was positively correlated with SP-A and SP-B, but Hoxb5 was not correlated with SP genes. In linear regression analysis, as well, the Hoxa5 and Hoxb5 expression did not affect SP-A, -B mRNA expression in the preterm group, while in the term group, increased Hoxa5 expression correlated with increased the SP-A and SP-B expression. This confirmed that Hoxa5 does not affect SP expression in immature lungs, but is involved in SP expression in mature lungs and affects late lung maturity. While Hoxb5 does not affect SP expression in the term group.
The underlying mechanism has not yet been fully elucidated, but recent advances in deciphering the epigenetic status of loci in developing embryos, as well as steric tissues in the nuclear space, have provided considerable information [29]. Hox genes play important roles in regulating biological processes, including organ morphogenesis and maturation. However, little is known about how Hox genes accomplish their function of sculpting variations on a basic segmental shape, many remain to be identified [18, 30]. Several Hox realizator genes have been identified that demonstrate how the Hox gene carries out the function of sculpting deformation in its basic segmental shape, but many genes should be identified. The number of Hox morphological effector genes will go through a known development process until almost all genes that mediate cell adhesion, cleavage, and migration are directly regulated by the Hox protein [31].
To the best of our knowledge, this is the first report describing interactions between the Hox genes and SP-A, SP-B in rabbit fetal lungs. Attempts to understand the role of Hox genes in normal lung development will be enhanced by the identification of their downstream target genes. Whereas Hoxa5 gene upregulates the SP-A and SP-B expression in this study. More research is also needed to explore the role of Hox genes in developmental processes. Furthermore, immunohistochemical studies should be performed to determine the site of Hox expression in fetal pulmonary tissue, and studies to determine the onset and duration of Hox expression during lung development in the gestation period.
Abbreviations
SP: surfactant protein; qRT-PCR: quantitative reverse transcription PCR.
Author contribution
Research conception and design: Chung SH, Bae CW; Data acquisition: Chung SH and Bae CW; Data analysis and interpretation: Chung SH; Drafting of the manuscript: Chung SH; Critical revision of the manuscript: Chung SH; Approval of the final manuscript: all authors.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Pearson JC, Lemons D, McGinnis W. Modulating Hox gene functions during animal body patterning. Nat Rev Genet. 2005;6:893-904
2. Garcia-Fernàndez J. The genesis and evolution of homeobox gene clusters. Nat Rev Genet. 2005;6:881-92
3. Kessel M, Gruss P. Murine developmental control genes. Science. 1990;249:374-9
4. Favier B, Dollé P. Developmental functions of mammalian Hox genes. Mol Hum Reprod. 1997;3:115-31
5. Philippidou P, Dasen JS. Hox genes: choreographers in neural development, architects of circuit organization. Neuron. 2013;80:12-34
6. Metzger RJ, Klein OD, Martin GR, Krasnow MA. The branching programme of mouse lung development. Nature. 2008;453:745-50
7. Warburton D, Bellusci S, De Langhe S, Del Moral PM, Fleury V, Mailleux A. et al. Molecular mechanisms of early lung specification and branching morphogenesis. Pediatr Res. 2005;57:26R-37R
8. Volpe MV, Martin A, Vosatka RJ, Mazzoni CL, Nielsen HC. Hoxb-5 expression in the developing mouse lung suggests a role in branching morphogenesis and epithelial cell fate. Histochem Cell Biol. 1997;108:495-504
9. Lufkin T, Dierich A, LeMeur M, Mark M, Chambon P. Disruption of the Hox-1.6 homeobox gene results in defects in a region corresponding to its rostral domain of expression. Cell. 1991;66:1105-19
10. Chisaka O, Capecchi MR. Regionally restricted developmental defects resulting from targeted disruption of the mouse homeobox gene hox-1.5. Nature. 1991;350:473-9
11. Kappen C. Hox genes in the lung. Am J Respir Cell Mol Biol. 1996;15:156-62
12. Volpe MV, Archavachotikul K, Bhan I, Lessin MS, Nielsen HC. Association of bronchopulmonary sequestration with expression of the homeobox protein Hoxb-5. J Pediatr Surg. 2000;35:1817-9
13. Volpe MV, Pham L, Lessin M, Ralston SJ, Bhan I, Cutz E. et al. Expression of Hoxb-5 during human lung development and in congenital lung malformations. Birth Defects Res A Clin Mol Teratol. 2003;67:550-6
14. Durham PL, Nanthakumar EJ, Snyder JM. Developmental regulation of surfactant-associated proteins in rabbit fetal lung in vivo. Exp Lung Res. 1992;18:775-93
15. Korstanje R, O'Brien PC, Yang F, Rens W, Bosma AA, van Lith HA. et al. Complete homology maps of the rabbit (Oryctolagus cuniculus) and human by reciprocal chromosome painting. Cytogenet Cell Genet. 1999;86:317-22
16. Pringle KC. Human fetal lung development and related animal models. Clin Obstet Gynecol. 1986;29:502-13
17. Bogue CW, Gross I, Vasavada H, Dynia DW, Wilson CM, Jacobs HC. Identification of Hox genes in newborn lung and effects of gestational age and retinoic acid on their expression. Am J Physiol. 1994;266:L448-54
18. Golpon HA, Geraci MW, Moore MD, Miller HL, Miller GJ, Tuder RM. et al. HOX genes in human lung: altered expression in primary pulmonary hypertension and emphysema. Am J Pathol. 2001;158:955-66
19. Grier DG, Thompson A, Lappin TR, Halliday HL. Quantification of Hox and surfactant protein-B transcription during murine lung development. Neonatology. 2009;96:50-60
20. Packer AI, Mailutha KG, Ambrozewicz LA, Wolgemuth DJ. Regulation of the Hoxa4 and Hoxa5 genes in the embryonic mouse lung by retinoic acid and TGFbeta1: implications for lung development and patterning. Dev Dyn. 2000;217:62-74
21. Mollard R, Dziadek M. Homeobox genes from clusters A and B demonstrate characteristics of temporal colinearity and differential restrictions in spatial expression domains in the branching mouse lung. Int J Dev Biol. 1997;41:655-66
22. Tümpel S, Wiedemann LM, Krumlauf R. Hox genes and segmentation of the vertebrate hindbrain. Curr Top Dev Biol. 2009;88:103-37
23. Rancourt DE, Tsuzuki T, Capecchi MR. Genetic interaction between hoxb-5 and hoxb-6 is revealed by nonallelic noncomplementation. Genes Dev. 1995;9:108-22
24. Aubin J, Lemieux M, Tremblay M, Bérard J, Jeannotte L. Early postnatal lethality in Hoxa-5 mutant mice is attributable to respiratory tract defects. Dev Biol. 1997;192:432-45
25. Boucherat O, Chakir J, Jeannotte L. The loss of Hoxa5 function in mice promotes NOTCH-dependent goblet cell metaplasia in lung airways. Biol Open. 2012;1:677-91
26. Mandeville I, Aubin J, LeBlanc M, Lalancette-Hébert M, Janelle MF, Tremblay GM. et al. Impact of the loss of Hoxa5 function on lung alveogenesis. Am J Pathol. 2006;169:1312-27
27. McIntyre DC, Rakshit S, Yallowitz AR, Loken L, Jeannotte L, Capecchi MR. et al. Hox patterning of the vertebrate rib cage. Development. 2007;134:2981-9
28. Boucherat O, Montaron S, Bérubé-Simard FA, Aubin J, Philippidou P, Wellik DM. et al. Partial functional redundancy between Hoxa5 and Hoxb5 paralog genes during lung morphogenesis. Am J Physiol Lung Cell Mol Physiol. 2013;304:L817-30
29. Montavon T, Duboule D. Chromatin organization and global regulation of Hox gene clusters. Philos Trans R Soc Lond B Biol Sci. 2013;368:20120367
30. Lappin TR, Grier DG, Thompson A, Halliday HL. HOX genes: seductive science, mysterious mechanisms. Ulster Med J. 2006;75:23-31
31. Foronda D, de Navas LF, Garaulet DL, Sánchez-Herrero E. Function and specificity of Hox genes. Int J Dev Biol. 2009;53:1404-19
Author contact
![]() Corresponding author: Sung-Hoon Chung, Department of Pediatrics, Kyung Hee University Hospital at Gangdong, 892 Dongnam-ro, Gangdong-gu, Seoul 05278, Korea. Tel: +82-2-440-6287, Fax: +82-2-440-7175 E-mail: pedcnet
Corresponding author: Sung-Hoon Chung, Department of Pediatrics, Kyung Hee University Hospital at Gangdong, 892 Dongnam-ro, Gangdong-gu, Seoul 05278, Korea. Tel: +82-2-440-6287, Fax: +82-2-440-7175 E-mail: pedcnet
Received 2017-4-25
Accepted 2017-8-7
Published 2017-9-19
