ISSN: 1449-1907International Journal of Medical Sciences
Int J Med Sci 2016; 13(12):963-969. doi:10.7150/ijms.16381 This issue Cite
Research Paper
Decreased Tumor Suppressor Candidate 3 Predicts Poor Prognosis of Patients with Esophageal Squamous Cell Carcinoma
1. Department of Radiation Oncology, Shandong Provincial Qianfoshan Hospital, Shandong University, Jinan 250014, P. R.China.
2. Medical Research Center, Shandong Provincial Qianfoshan Hospital, Shandong University, Jinan, 250014, P. R.China.
3. Department of Traditional Chinese Medicine, Shandong Provincial Qianfoshan Hospital, Shandong University, Jinan, 250014, P. R.China.
4. Department of Radiation Oncology, Shandong Cancer Hospital, Jinan, 250117, P. R.China.
5. Department of Oncology, Shandong University of Traditional Chinese Medicine Jinan, 250355, P. R.China.
Abstract
TUSC3 was recently identified as a potential tumor suppressor gene in a variety of human malignancies. However, no data are currently available regarding the expressions of TUSC3 in esophageal cancer (ESCC).The purposes of this study was to investigated the expressions of TUSC3 in ESCC tissues and assess the relationship between TUSC3 levels and clinico-pathological characteristics of ESCC patients. TUSC3 protein expressions were evaluated by immunohistochemistry (IHC) on tissue microarray slides in esophageal cancer, which included 95 esophageal squamous carcinoma specimens (ESCC), and 75 normal esophageal mucosa (NEM). We found that TUSC3 in ESCC was significant lower than that in NEM (P=0.000). According to multi-clinical classifications, TUSC3 level varied significantly with TNM stage, T stage, and N stage (p<0.001, p=0.0368, p<0.0001, respectively). Univariate analysis showed that gender, TNM stage, T stage, N stage, TUSC3 expression were prognostic factors for survival. Multivariate analysis showed that in our study, only TUSC3 expression was independent prognostic factors for ESCC. Our results indicated for the first time, a combined analysis of TUSC3 expressions as well as the clinical variables will help predict the prognosis of ESCC patients. Further large-sample validation and functional analysis should be performed to evaluate its potential prognostic and therapeutic values for ESCC patients.
Keywords: Tumor suppressor candidate 3 (TUSC3), Esophageal squamous cell carcinoma (ESCC), Biomarker, Overall survival (OS), Prognosis.
Introduction
Esophageal cancer is the 8th most frequently diagnosed cancer and the 6th most common cause of cancer-mortality worldwide[1]. Esophageal cancers are classified as esophageal adenocarcinoma (EAC) and esophageal squamous cell carcinoma (ESCC) according to histological type in clinical practice. Particularly, ESCC accounts for 95% of all esophageal cancers in China and the five-year survival rate is low, due to its late diagnosis[2]. The majority of patients present with the advanced stage, at which point ESCC patients are unable to undergo a radical treatment[3]. ESCC is extremely aggressive and often results in a dismal prognosis. An improved understanding of ESCC is urgently needed to identify novel biomarker and effective therapeutic strategies for eshophagus cancer patients.
Tumor suppressor candidate 3 (TUSC3), a novel tumor suppressor gene, originally has been known to be responsible for autosomal recessive mental retardation for several years[4-6]. Only recently was TUSC3 identified as a tumor suppressor gene when it was found deleted in a variety of human malignancies[7, 8]. The protein is localized in the endoplasmic reticulum and encodes a subunit of the endoplasmic reticulum-bound oligosaccharyl transferase (OST) complex, which is primarily responsible for protein N-linked glycosylation[9]. Studies showed that disfunction or deletion of TUSC3 exert its oncological effects as a modulator by inhibiting glycosylation efficiency and consequently inducing the endoplasmic reticulum stress and cell malignant transformation[10-13]. However, no data are currently available regarding the expressions of TUSC3 in ESCC. In the present study, we investigated the expressions of TUSC3 in ESCC and the relationship between TUSC3 expressions and the clinico-pathological parameters of ESCC patients, with an emphasis on prognostic factors that correlate with its survival time.
Material and methods
Tissue samples
Tissue microarray slides were purchased from Shanghai Outdo Biotech Co., LTD, Shanghai, China. The slides included 95 esophageal squamous carcinoma specimens, 75 normal esophageal mucosa(NEM) tissue specimens. The detailed clinical-pathologic characteristics of patients with esophageal cancer are listed in Table 1. All patients were clinically staged (TNM staging, tumor nodes metastasis staging) according to the seventh edition of the American Joint Committee on Cancer (AJCC) system for esophageal cancer[14]. The pathological differentiated degrees are defined as follows: 1, High-differentiation carcinoma; 2, Medium-differentiation carcinoma; and 3, Low-differentiation. The degree of differentiation for the tumors in each of the patients was evaluated by two pathologists.
Immunohistochemistry assay
Immunohistochemistry (IHC) staining was performed directly on the tissue slides. Briefly, after incubation for 2 hours at 56°C, the slides were dewaxed with xylene and rehydrated through graded alcohols (100%, 90%, 70% and 50% alcohol; 5 minutes each). Endogenous peroxidase activity was blocked with 3% H2O2 for 15 minutes. For antigen retrieval, sections were incubated in sodium citrate buffer (0.01 M, pH 6.0) for 20 minutes in a household microwave oven (600W). Then, the slides were incubated with 10% normal goat serum to block nonspecific binding sites. Thereafter, the slides were incubated with the TUSC3 goat polyclonal antibody (Santa Cruz, USA, 1:100 final dilution) overnight at 4°C. After washing, the bio-labeled secondary antibody, rabbit anti-goat IgG (ZSGB-Bio, China), was applied at a 1:200 dilution for 40 minutes at 37°C. The sections were then stained with diaminobenzidine (DAB). Finally, the sections were counterstained with hematoxylin and eosin, dehydrated with graded alcohol and mounted using neutral gum. A digital pathology system for stained cells scoring was performed by Aperio ImageScope (Aperio Technologies, Inc., Vista, CA).
Immunoreactivity was observed in the cytoplasm of cells and the scoring was based on cytoplasmic staining. Immunoreactivity for TUSC3 expressions was independently evaluated by two pathologists from the Qianfoshan hospital and categorized according to the immunoreactive score (IRS): IRS = SI (staining intensity) × PP (percentage of positively stained cells). SI was determined as 0 (negative), 1 (weak), 2 (moderate) or 3 (strong). PP was scored as 0 (negative), 1 (<25% of the cells), 2 (25%-50% of the cells), 3 (50%-75% of the cells), or 4 (>75% of the cells). A final score was then calculated by multiplying the above two scores. Additionally, all of the specimens were divided into two groups showing negative or positive expressions by using an IRS of 6 as the cut-off value.
Basic Characteristics of Patients.
| N(%) | Positive | Negative | Positive rate(%) | χ2 | pa | |
|---|---|---|---|---|---|---|
| Age(yrs) | ||||||
| Median(range) | 65(48-81) | 0.066 | 0.797 | |||
| <65 | 43(45.2) | 13 | 30 | 30.2 | ||
| ≥65 | 52(54.8) | 17 | 35 | 32.7 | ||
| Gender | 3.020 | 0.082 | ||||
| Male | 71(74.7) | 19 | 52 | 26.8 | ||
| Female | 24(25.3) | 11 | 13 | 45.8 | ||
| Location | 0.649 | |||||
| Upper | 5 ( 5.3) | 2 | 3 | 40.0 | ||
| Mid-lower | 90 (94.7) | 28 | 62 | 31.1 | ||
| TNM Stageb | 17.306 | 0.000※ | ||||
| I+II | 45(47.4) | 24 | 23 | 53.3 | ||
| III | 50(52.6) | 6 | 44 | 12.0 | ||
| LNMd | 11.823 | 0.001※ | ||||
| negative | 42(44.2) | 21 | 21 | 50.0 | ||
| positive | 53(55.8) | 9 | 44 | 17.0 | ||
| Differentiation gradec | 2.284 | 0.131 | ||||
| 1 | 31(32.6) | 13 | 18 | 41.9 | ||
| 2+3 | 64(67.4) | 17 | 47 | 26.6 |
aPositive rates of TUSC3 expression were compared by Fisher exact test;
bTNM staging is defined according to the seventh edition of the tumornodemetastasis classification for malignant tumors.
cDifferentiated degree was evaluated by two pathologists from Qianfoshan Hosptital. The pathological differentiated degrees are defined as follows: 1, High-differentiated carcinoma; 2, Medium-differentiated carcinoma; 3, Low-differentiated carcinoma.
dLNM: lymph node metastasis.
※ P<0.05
Statistical analysis
The SPSS 13.0 software was used for the statistical analyses. Levels of TUSC3 expressions were compared using a rank sum test. Comparisons of positive rate between two groups were performed using a Fisher exact test and the Spearman correlation method was used to evaluate the association of scores. The significance of correlations between clinical pathological parameters (age, gender, TNM-stages, T-stage, N-stage, differentiated degree, and mass location) and IRS of TUSC3 were determined using Fisher's exact test. All reported P values were two-sided, and P < 0.05 was considered statistically significant. Survival analysis and curves were established according to the Kaplan-Meier method and were compared using the log-rank test. Cox's regression was used to perform the multivariate survival analysis. Overall survival(OS) was defined as the elapsed time from the initial treatment date to the death or to the patients last visit. The initial recurrence was categorized as locoregional recurrence or distant metastasis depending on the location of the recurred lesion. A receiver operating characteristics (ROC) curve analysis was performed to assess the cut-off for TUSC3 levels in patients with ESCC and NEM. The area under curve (AUC) and p-values were evaluated. Results were considered to be statistically significant with a P value <0.05.
Results
Patients characteristics
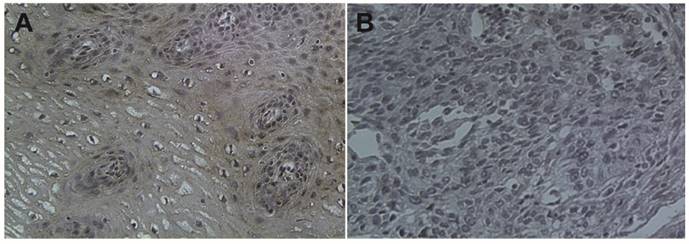
The baseline characteristics of the 95 patients are shown in Table 1. All patients were postoperated for ESCC and no patients are classified in TNM stage IV. The basic characteristics of the patients are shown in Table 1. The median age of ESCC patients was 65 (range, 48-81 years). No significant difference was observed in gender or age between normal controls and patients. There was no difference of TUSC3 expression between upper and mid-lower esophageal cancer (p=0.649, Table 1). Regarding the TNM staging, a significant increase in TUSC3 expressions could be observed in patients withⅠ+Ⅱstage compared with III stage patients (p=0.000, Table 1). When the Lymph node metastasis were considered, analysis revealed a marked decrease in TUSC3 expressions in patients with lymph node metastasis positive (LNM+) compared with patients with lymph node metastasis negative (LNM-) (p=0.001, Table 1). Additionally, the positive rate of TUSC3 expressions in patients with differentiated degree 1 showed no difference with those in patients with differentiated degree 2+3 (p=0.131, Table 1). The representative immunohistochemistry assay is shown in Fig.1 (magnification 40×).
Detection of TUSC3 level in ESCC patients and healthy controls
The positive rate of TUSC3 in NEM (normal esophageal mucosa) group and ESCC group were 96%, 31.6% respectively (Table 2). The positive rate of TUSC3 in NEM group was significantly higher than that in ESCC group (P=0.000; Table 2).
Comparison of TUSC3 expression between NEM and ESCC.
| Number | TUSC3 expression | Positive rate (%) | χ2 | pa | ||
|---|---|---|---|---|---|---|
| + | - | |||||
| NEM | 75 | 72 | 3 | 96 | 0.000 | 0.000 |
| ESCC | 95 | 30 | 65 | 31.6 | ||
a Fisher exact test was used to analyze the positive rates of TUSC3 expression.
bESCC, esophageal cancer; NEM, normal esophageal mucosa.
P < 0.05 was considered statistically significant.
Immunohistochemical staining of TUSC3 in human esophageal cancers. A. normal esophageal mucosa; B. esophageal squamous cell carcinoma.
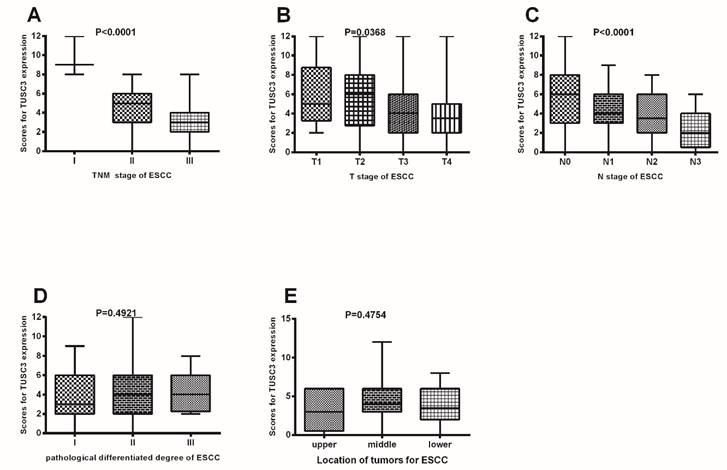
TUSC3 expressions were compared among different clinico-pathological groups. TUSC3 expressions were compared among different clinical TNM staging in ESCC patients (A); TUSC3 expressions were compared among the various T stages (B), N stages (C), pathological differentiated degrees (D) and mass location(E). The rank sum test was used to analyze the differences between groups.
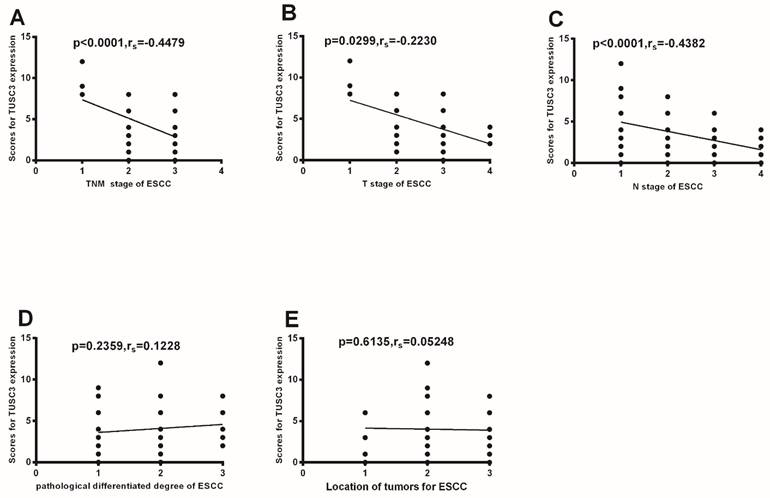
Association between TUSC3 expressions and the clinical-pathological characteristcs of ESCC
As shown in Figure 2, there were significant differences in TUSC3 expressions among patients with different TNM stages (Stage I to Stage III) in ESCC (P<0.0001 Fig. 2A). Additionally, significant differences in TUSC3 expressions were identified among the patients with different T stages (P=0.0368, Fig.2B) and N stages (P<0.0001, Fig.2C). However, no such differences were found among the patients with different pathological differentiated degrees (P=0.4921, Fig.2D) and mass locations(P=0.4754, Fig.2E).
Correlations of TUSC3 levels with clinical-pathological characteristcs of ESCC
Further analysis shows TUSC3 expressions were negatively correlated with clinical TNM staging (P<0.0001; rs=-0.4479, Fig.3A). Additionally, TUSC3 expressions were negatively correlated with T stages (P=0.0299; rs=-0.2230, Fig.3B), and N stages (P<0.0001; rs=-0.4382, Fig.3C). However, there were no correlations between TUSC3 expressions and pathological differentiated degrees (P=0.2359, Fig.3D), mass location (P=0.6135, Fig.3E).
Univariate and multivariate analysis of prognostic factors for ESCC
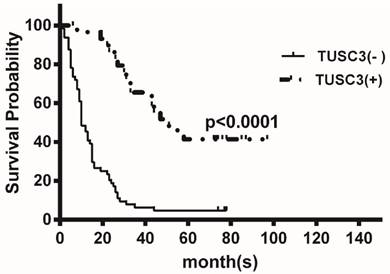
At the median follow-up of 16 months (range1-97months), the median OS time were 16 months. TUSC3 expression was divided into two groups according to the ROC analysis and defined ≥6 as positive expression, providing the best discrimination between patients and controls regarding optima values of sensitivity and specificity. Kaplan-Meier analysis revealed that low TUSC3 expression was associated with shorter overall survival (P<0.0001; Figure 4).
Univariate analysis showed four parameters were found to be independent prognostic factors: TNM stage, T stage, N stage, and TUSC3 expression. Multivariate analysis showed that in our study, only TUSC3 expression was independent prognostic factors for ESCC (Table 3).
Discussion
ESCC comprises 60-70% of all cases of esophageal cancer, which accounts for 5% of all cancer deaths worldwide[1, 15]. By the time the first symptoms appear, such as difficulty swallowing, ESCC has already well progressed. Therefore, there is an urgent need for reliable predictors and indicators of diagnosis and prognosis for ESCC.
TUSC3 has been recognized as a novel tumor suppressor gene involved in multiple-tumor development. The clinical characteristics of TUSC3 expression levels in several human tumors have been determined. Marta[10] found that TUSC3 plays a role in metastasis and that loss of TUSC3 is negatively related with lymph node metastasis in larynx and pharynx squamous cell carcinomas. Dietmar et al [11]found that TUSC3 loss may facilitate tumor growth. Ahmed et al[12] found that TUSC3 is involved in spermatogenesis in the testis and plays a role in normal prostate development. Peter Horak et al[13] found that TUSC3 expression is frequently lost in prostate cancer cell lines, leading to increased proliferation, migration and invasion of cancer cells. Our previous study showed TUSC3 was involved in the development of SCLC (Oncology letters accepted, data not shown). However, relatively little is known about the significance of TUSC3 expressions in ESCC patients. To the best of our knowledge, this is the first study that analyzes the relationship between the TUSC3 expression levels and the clinical characteristics of ESCC patients.
Univariate and multivariate analysis of the association of prognosis with clinicopathological characteristics and TUSC3 expression in ESCC patients.
| Univariate | Multivariate | |||||
|---|---|---|---|---|---|---|
| characteristics | HR | 95%CI | p | HR | 95%CI | p |
| Sex (male vs, female) | 1.730 | 0.998-3.002 | 0.051 | |||
| Age (<65 vs. ≥65) | 0.850 | 0.547-1.327 | 0.479 | |||
| TNM Stage(AJCC) (I+II vs. III) | 2.339 | 1.461-3.745 | 0.000※ | 2.061 | 0.641-6.628 | 0.225 |
| T Stage (1+2 vs. 3) | 1.901 | 1.199-3.016 | 0.006※ | 1.526 | 0.668-3.486 | 0.317 |
| N Stage (negative vs. positive) | 1.199 | 1.199-3.016 | 0.006※ | 0.580 | 0.194-1.736 | 0.330 |
| TUSC3 (negative vs. positive) | 0.193 | 0.109-0.340 | 0.000※ | 0.203 | 0.110-0.374 | 0.000※ |
| Differentiation grade (1 vs. 2+3) | 0.952 | 0.595-1.523 | 0.837 | |||
CI: confidence interval; HR :hazard ratio.
※ P<0.05
Correlation between TUSC3 expressions and different clinico-pathological characteristics. Correlations between TUSC3 expressions and clinical TNM staging were analyzed in ESCC patients (A). The correlations between TUSC3 expressions and the different T stages (B) and N stages(C) were analyzed. Differentiated degrees(D) and mass location(E) were analyzed in ESCC. The Spearman correlation method was used to evaluate the association of scores.
Kaplan-Meier analysis of the correlation between TUSC3 expression and overall survival in ESCC patients. Patients with low TUSC3 expression had significantly shorter overall survival (P<0.0001).
Using tissue microarray technology, we found that there was significant difference between ESCC group and NEM group in TUSC3 expression rate (χ2 = 0.000, P = 0.000; Table 2). Furthermore, regarding the TNM stage of ESCC, significant differences in TUSC3 expressions were identified among patients with different TNM staging (Stage I to Stage III) in the ESCC patients (P<0.0001, Fig. 2A). Further correlation analysis also confirmed the above results (P<0.0001; rs=-0.4479, Fig.3A). As we all know, TNM stage is an index to reflect tumor progression in clinical practices[15].TUSC3 expression may be a useful predictor of the progression of ESCC.
To further confirm the above results, the TUSC3 expressions among different T stages and N stages was evaluated in ESCC patients, respectively. Significant differences in TUSC3 expressions were identified among the patients with different T stages(P=0.0368, Fig.2B).Also, analysis revealed a marked decrease in TUSC3 expressions in patients with lymph node metastasis positive (LNM+) compared with patients with lymph node metastasis negative (LNM-) (p=0.001, Table 1). Additionally, significant differences in TUSC3 expressions were identified among patients with different T stages (P=0.0368; Fig. 2B) and different N stages (N0, N1, N2, N3) (P=0.000; Fig. 2C) in the ESCC patients. Correlation analysis also identified a negative correlation between TUSC3 expressions and T stages (P=0.0299; rs=-0.2230, Fig. 3B) or N stages (P<0.0001; rs=-0.4382, Fig. 3C) in all ESCC tissues tested. The results suggest that lower TUSC3 expression may indicate more depth of tumor invasion and the higher probability of lymph nodes metastasis in ESCC patients. The results have guided significance to clinical practice. For an individual patient, a combined analysis of TUSC3 expressions as well as the clinical variables will help predict the incidence of lymph node metastasis and invasion degree for ESCC patients.
To date, there is little knowledge about the prognostic value of TUSC3 for human malignancies. We found that low TUSC3 expression group had a significantly poorer OS than those with high TUSC3 expression group (P<0.0001; Figure 4). Univariate and multivariate analyses showed that only TUSC3 expression was identified as an independent predictor for ESCC (p=0.000; Table 3).
The major limitation of our study is the relatively small number of samples included in our study that might result in imprecise evaluation of TUSC3 levels and its correlation with other clinical indices. Our further study will verify the above results using a larger sample size. In addition, the mechanism through which TUSC3 was involved in the development of ESCC needs to be further studied.
In conclusion, A combined analysis of TUSC3 expressions as well as the clinical variables will help predict the incidence of lymph node metastasis and invasion degree for ESCC patients. Notably, our findings provide the first evidence that a loss or reduce of TUSC3 may be associated with poorly prognosis of ESCC patients.
Abbreviations
TUSC3: Tumor suppressor candidate 3; ESCC: Esophageal squamous cell carcinoma; EAC: esophageal adenocarcinoma; NEM: normal esophageal mucosa; IHC: immunohistochemistry; OS: Overall survival.
Acknowledgements
This work was supported by grants from medical and health science and technology development plan of Shandong Province (2015WS0213), grants from Science Foundation of Shandong Province (ZR2011HQ010, ZR2015HM077, ZR2016HQ50) and National Natural Science Foundation of China (No.30901712; No.81301868).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Stewart B, Wild CP. World Cancer Report. IARC Press. Washington: District of Columbia 2014 Chapter 5.3
2. Chen W, He Y, Zheng R. et al. Esophageal cancer incidence and mortality in China. J ThoracDis. 2013;5(1):19-26
3. Sankaranarayanan R, Swaminathan R, Jayant K. et al. An overview of cancer survival in Africa, Asia, Caribbean and Central America: the case for investments in cancer health services. In IARC Press. Washington: District of Columbia. 2011 Chapter 32
4. Loddo S, Parisi V, Doccini V. et al. Homozygous deletion in TUSC3 causing syndromic intellectual disability: a new patient. Am J Med Genet A. 2013;161A:2084-2087
5. Garshasbi M, Kahrizi K, Hosseini M. et al. A novel nonsense mutation in TUSC3 is responsible for non-syndromic autosomal recessivemental retardation in a consanguineous Iranian family. Am J Med Genet A. 2011;155A:1976-1980
6. Garshasbi M, Hadavi V, Habibi H. et al. A defect in the TUSC3 gene is associated with autosomal recessive mental retardation. Am J Hum Genet. 2008;82:1158-1164
7. Bova GS, MacGrogan D, Levy A. et al. Physical mapping of chromosome 8p22 markers and theirhomozygous deletion in ametastatic prostate cancer. Genomics. 1996;35:46-54
8. MacGrogan D, Levy A, Bova GS. et al. Structure and methylation-associated silencing of a gene within a homozygously deleted region of human chromosome band 8p22. Genomics. 1996;35:55-65
9. Mohorko E, Owen RL, Malojčić G. et al. Structural Basis of Substrate Specificity of Human Oligosaccharyl Transferase Subunit N33/Tusc3 and Its Role in Regulating Protein N-Glycosylation. Structure. 2014;22:590-601
10. Guervós MA, Marcos CA, Hermsen M. et al. Deletions of N33, STK11 and TP53 are involved in the development of lymph node metastasis in larynx and pharynx carcinomas. Cell Oncol. 2007;29:327-334
11. Pils D, Horak P, Gleiss A, Sax C. et al. Five genes from chromosomal band 8p22 are significantly down-regulated in ovarian carcinoma: N33 and EFA6R have a potential impact on overall survival. Cancer. 2005;104:2417-2429
12. Khalid AM, Asano A, Hosaka YZ. et al. Tumor Suppressor Candidate TUSC3 Expression during Rat Testis Maturation. Biosci Biotechnol Biochem. 2013;77:2019-2024
13. Horak P, Tomasich E, Vaňhara P. et al. TUSC3 Loss Alters the ER Stress Response and Accelerates Prostate Cancer Growth in vivo. Sci Rep. 2014;4:3739-3747
14. Mönig SP, Hölscher AH. Clinical classification systems of adenocarcinoma of the esophagogastric junction. Recent Results Cancer Res. 2010;182:19-28
15. Conteduca V, Sansonno D, Ingravallo G. et al. Barrett's esophagus and esophageal cancer: an overview. International Journal of Oncology. 2012;41(2):414-424
Author contact
![]() Corresponding author: Dr. Juan Du, Zip code: 250014. Fax: 0531-82967114. E-mail address: sunnydujuancom.
Corresponding author: Dr. Juan Du, Zip code: 250014. Fax: 0531-82967114. E-mail address: sunnydujuancom.
Received 2016-6-4
Accepted 2016-10-7
Published 2016-11-25
