ISSN: 1449-1907International Journal of Medical Sciences
Int J Med Sci 2014; 11(1):65-79. doi:10.7150/ijms.7426 This issue Cite
Research Paper
Human Hepatic Progenitor Cells Express Hematopoietic Cell Markers CD45 and CD109
1. State Key Laboratory for Diagnosis and Treatment of Infectious Diseases, Collaborative Innovation Center for Diagnosis and Treatment of Infectious Diseases, The First Affiliated Hospital, School of Medicine, Zhejiang University. 79 Qingchun Rd., Hangzhou, 310003. China.
2. Department of Surgery, The First Affiliated Hospital, School of Medicine, Zhejiang University. 79 Qingchun Rd., Hangzhou, 310003. China.
3. Department of Pathology, The First Affiliated Hospital, School of Medicine, Zhejiang University. 79 Qingchun Rd., Hangzhou, China. 310003
Abstract
Objective: To clarify the precise characteristics of human hepatic progenitor cells (HPCs) for future cytotherapy in liver diseases.
Methods: Hepatic progenitor-like cells were isolated and cultured from the livers of patients who had undergone partial hepatectomy for various pathologies but displayed no sign of hepatic dysfunction. These cells were characterized by transcriptomic profiling, quantitative real-time PCR and immunocyto/histochemistry.
Results: Cultured HPCs contained polygonal, high nucleus/cytoplasm ratio and exhibited a global gene expression profile similar (67.8%) to that of primary hepatocytes. Among the genes with more than 20-fold higher expression in HPCs were a progenitor marker (CD90), a pentraxin-related gene (PTX3), collagen proteins (COL5A2, COL1A1 and COL4A2), cytokines (EGF and PDGFD), metabolic enzymes (CYBRD1, BCAT1, TIMP2 and PAM), a secreted protein (SPARC) and an endothelial protein C receptor (PROCR). Moreover, eight markers (ALB, AFP, CK8, CK18, CK19, CD90, CD117 and Oval-6) previously described as HPC markers were validated by qRT-PCR and/or immunocyto/histochemistry. Interestingly, human HPCs were also positive for the hematopoietic cell markers CD45 and CD109. Finally, we characterized the localization of HPCs in the canals of Hering and periportal areas with six previously described markers (Oval-6, CK8, CK18, CK19, CD90 and CD117) and two potential markers (CD45 and CD109).
Conclusion: The human HPCs are highly similar to primary hepatocytes in their transcriptional profiles. The CD45 and CD109 markers could potentially be utilized to identify and isolate HPCs for further cytotherapy of liver diseases.
Keywords: Human hepatic progenitor cell, Immunocytochemistry, Transcriptional profile
Introduction
Hepatocyte transplantation presents a promising alternative to orthotopic liver transplantation in patients with end-stage liver disease, including chronic viral hepatitis, hepatocellular carcinoma, and alcoholic liver disease (1-5). However, the supply of hepatocytes is limited by a shortage of donor organs and the lower proliferation of these cells in vitro (6-8). Therefore, many researchers are currently focused on developing alternative sources of cells (9, 10). Stem cells are promising resources for hepatocyte therapy to correct inborn errors of metabolism and bridge patients with different kinds of end-stage liver failure to transplantation or spontaneous recovery (10-13).
The liver displays an extensive regenerative capacity in response to loss of mass from injury or partial hepatectomy (1, 14). In most cases, hepatocytes can undergo several rounds of proliferation. However, this capacity is limited after extensive or chronic injury and/or when hepatocyte repopulation is impaired (1). Moreover, the cell types that participate in liver regeneration remain uncertain, although mature hepatocytes and stem cells might both be involved (15, 16). Hepatic progenitor cells (HPCs), the human counterpart to rodent oval cells, exist in portal or periportal areas and contribute to liver regeneration (17, 18). These cells emerge into the parenchyma, generally in the form of neoductules, and differentiate into mature hepatocytes and cholangiocytes to restore the damaged liver. However, the precise origin of these bipotent progenitors is still unclear (17, 19, 20). Bone marrow stem cells (BMSCs) can repopulate an injured liver, differentiate into mature hepatocytes in vitro, and participate in liver regeneration (8, 16, 21). BMSC transplantation has been used to improve a mouse model of tyrosinemia (9, 22) and an inborn defect in a hepatocytic enzyme (1, 4, 5) by reconstituting the architecture and function of the injured liver. Our previous studies have demonstrated that acute hepatic failure-derived rat BMSCs have a hepatic transcriptional profile, expressing many of the same genes expressed by hepatocytes (23, 24). These studies indicate that BMSCs are a potential source of HPCs for hepatocyte regeneration.
Species-specific gene expression has been characterized in human and animal HPCs (9, 17). In rodents (25-29), oval cells express markers of both hepatocyte and cholangiocyte lineages (such as albumin [ALB], cytokeratin 18 [CK18] and CK19) in addition to markers of both hematopoietic and mesenchymal stem cells (CD34, CD45 and CD90). In humans, HPCs with morphological features and immunohistological markers similar to those of oval cells have been characterized in normal liver and patients with chronic liver injury or submassive hepatic necrosis (30-36). Furthermore, these bipotent progenitor cells with characteristics similar to HPCs exist not only in human adult liver but also in human fetal liver (37, 38).
However, the precise characteristics of HPCs in humans are unknown, as are any gene expression differences between HPCs and primary hepatocytes. Therefore, studies on human HPCs are urgently needed. In this study, we isolated HPCs from the livers of patients who had undergone partial hepatectomy for various pathologies but displayed no signs of hepatic dysfunction. These cells were characterized by global gene expression profiling, quantitative real-time reverse-transcription polymerase chain reaction (qRT-PCR) and immunocyto/histochemistry.
Materials and Methods
Chemicals and reagents
Chemicals for cell isolation and culture including ITX mixture, nicotinamide and dexamethasone were purchased from Sigma Chemical Corporation (MO, USA). Dulbecco's modified Eagle's medium (DMEM), Iscove's modifi ed Dulbecco's medium (IMDM), penicillin, streptomycin, trypsin-ethylenediaminetetraacetic acid (EDTA), bovine serum albumin (BSA), collagenase IV and fetal bovine serum (FBS) were purchased from Invitrogen (CA, USA). Human hepatocyte growth factor (HGF) and epidermal growth factor (EGF) were from Cell Science (Canton, MA). Regular culture dishes and flasks were purchased from BD Falcon (NJ, USA), and immunohistochemistry reagents were purchased from Santa Cruz Biotechnology (CA, USA) and Bio-Rad Laboratories (CA, USA).
Isolation and culture of human hepatocytes and HPCs
Human liver tissue specimens (n=12) used in this study were harvested from therapeutic partial hepatectomies performed on patients with various pathologies (trauma-induced liver rupture, hepatic haemangioma, etc.) who exhibited no sign of hepatic dysfunction, hepatitis viruses (including A-E) infection and hepatic cancer. Informed consent was obtained from the patients and/or their relatives before the procedure. All experimental protocols in this study were approved by the Ethics Committee of the First Affiliated Hospital, School of Medicine, Zhejiang University. The method of primary hepatocyte isolation was previously described (39). Briefly, the liver specimens (10 to 20 g) were perfused with four different buffer solutions supplemented with EDTA, 0.5% dispase, 0.05% collagenase type IV and DNase I. The freshly isolated whole liver cell suspensions were centrifuged at 50 × g for 1 min at 4 °C. The pellet was resuspended in DMEM and centrifuged at 50 × g for 1 min at 4 °C. The supernatant and pellet were harvested separately. The pelleted hepatocytes were resuspended in PBS and centrifuged twice at 50 × g for 1 min at 4 °C. The purified hepatocytes were harvested, and total RNA was extracted to perform global gene expression profiling by RT-PCR and qRT-PCR. The supernatant portion containing the HPCs was centrifuged at 150 × g for 2 min at 4 °C. The pellet was resuspended in DMEM and centrifuged at 150 × g for 5 min at 4 °C. The pelleted cells were resuspended in DMEM with 10% FBS and purified by Percoll density gradient centrifugation. The harvested cells were cultured in a 10-mm plastic Petri dish in DMEM supplemented with 10% FBS, 2 mM L-glutamine, 100 U/ml penicillin and 100 µg/ml streptomycin. The cultures were maintained at 37 °C in a humidified incubator in a mixture of 95% air and 5% CO2. The medium was changed twice a week. After three or four days of culture, some contaminated hepatic cells showed a spindly shape including endothelial cells, hepatic stellate cells and kupffer cells were scratched out from the petri dish by cell scraper. The cells with a round or oval shape were remained to proliferate for further culture. After one week of culture, the culture medium containing FBS was replaced with hepatocyte serum-free medium. The fresh medium was changed twice a week. To harvest enough HPCs for further transcriptional profiling analysis, the cells were cultured for three weeks.
Morphological characterization
The morphological features of the cultured HPCs were observed under phase-contrast microscopy (ECLIPSE TS100, Nikon, Tokyo, Japan) during the entire culture period, and cultures were evaluated on the basis of cell shape and size, colony formation, cell density, cytoplasm to nucleus ratio and colony size. Images were acquired and processed using digital imaging software (NIS-Elements F3.0).
Bipotential differentiation
To characterization of bipotential differentiation ability, HPCs were cultured in a commercially available adipogenic differentiation medium (Cambrex, Lonza, Walkersville) for adipocytes differentiation. On day 10, the differentiated cells were stained with Oil Red O. For hepatogenic differentiation, HPCs were cultured in serum-free hepatocyte differentiation medium (IMDM supplemented with 5 ng/mL hHGF, 100 μM dexamethasone, 50 mg/mL ITX + premix, 100 U/mL penicillin, and 100 μg/mL streptomycin). On day 10, the morphology of differentiated cells were characterized by phase-contrast microscopy. For cholangiogenic differentiation, HPCs were cultured in cholangiocyte differentiation medium (IMDM supplemented with 10% FBS, 50 mg/mL ITX + premix, 10 mM nicotinamide, 5 ng/mL hHGF, 5 ng/mL EGF, 100 μM dexamethasone, 100 U/mL penicillin, and 100 μg/mL streptomycin). On day 10, the differentiated cells were characterized by immunocytochemistry with the human cholangiocyte-specific marker CK19.
Microarray analysis
For transcriptional profiling of HPCs, the high-density microarray GeneChip Human Genome U133 Plus 2.0 Array (Affymetrix, CA, USA) was used to characterize mRNA expression following the manufacturer's instructions. Briefly, total RNA was extracted from cultured HPCs and primary hepatocytes using an RNeasy Mini Kit (Qiagen, CA, USA) following the manufacturer's protocol. Single- and double-stranded cDNA was synthesized from qualified total RNA samples using the GeneChip 3' IVT Express Kit (Affymetrix). Three samples of total RNA (200 ng) pooled from five different specimens of each cell type were used for cDNA synthesis. After double-stranded cDNA cleanup and a quality check, in vitro transcription was performed using the GeneChip IVT Labeling Kit (Affymetrix) to produce biotin-labeled cRNA from the cDNA. The Human Genome U133 Plus 2.0 arrays were hybridized at 45 °C for 16 h at 60 rpm with a hybridization cocktail containing 15 µg of fragmented and labeled cRNA in a hybridization oven (Affymetrix). The arrays were then washed and stained with streptavidin-phycoerythrin (SAPE, Molecular Probes, OR, USA). All hybridization assay methods, including the preparation of solutions, were performed according to the manufacturers' instructions. The distribution of fluorescent material on the array was detected using a 3000 7G GeneChip Scanner (Affymetrix). Microarray Suite version 5.0 and GeneChip Operating Software supplied by Affymetrix were used for gene expression analysis. The pairwise Pearson correlation coefficient was used to analyze global gene expression similarities between human HPCs and hepatocytes. The specimens were hierarchically clustered by inputting the probe sets/genes that exhibited a greater than 20-fold change between HPCs and hepatocytes.
RT-PCR analysis
To characterize the molecular properties of HPCs, we used RT-PCR to analyze selected genes that are known to be markers of HPCs and hepatocytes (30, 31, 33-36) (albumin [ALB], α-fetoprotein [AFP], cytokeratin 8 [CK8], CK18, CK19, CD34, CD45, CD90, CD117, tyrosine aminotransferase [TAT], glutamine synthase [GS], and glucose-6-phosphate dehydrogenase [G6PD]). The RT-PCR protocol was previously described (23). Briefly, 1 µg of the total RNA used in the microarray analysis was reverse transcribed to cDNA using the QuantiTect Reverse Transcription Kit (Qiagen, Valencia, CA) according to the manufacturer's instructions, and then the cDNA was amplified by PCR using previously published primers for markers of HPCs and hepatocytes (Table 1). The PCR conditions varied according to the primers used. GAPDH (glyceraldehyde phosphate dehydrogenase) was used as an internal control.
Primers used for human HPC gene amplification
| Gene | NCBI accession number | Length | Primer | |
|---|---|---|---|---|
| ALB | NM_000477.5 | 164 | F | 5'-CTGAGCAAAGGCAATCAACA-3' |
| R | 5'-CACAGTCTGCTGAGGTTGGA-3' | |||
| AFP | NM_001134.1 | 248 | F | 5'-AGCTTGGTGGTGGATGAAAC-3' |
| R | 5'-CCCTCTTCAGCAAAGCAGAC-3' | |||
| CK8 | NM_002273.2 | 131 | F | 5'-TCATAGACAAGGTACGGTTCC-3' |
| R | 5'-GCCTAAGGTTGTTGATGTAGC-3' | |||
| CK18 | NM_199187.1 | 164 | F | 5'-GAGCTGCTCCATCTGTAGGG-3' |
| R | 5'-CACAGTCTGCTGAGGTTGGA-3' | |||
| CK19 | NM_002276.4 | 138 | F | 5'-CATGAAAGCTGCCTTGGAAGA-3' |
| R | 5'-TGATTCTGCCGCTCACTATCAG-3' | |||
| CD34 | NM_001773.2 | 206 | F | 5'-TGCATGTGCAGACTCCTTTC-3' |
| R | 5'-GAGGACAAGGCTGAGGTCTG-3' | |||
| CD45 | NM_002838.3 | 158 | F | 5'-ACGAAGCTCTTAGCGTCAGG-3' |
| R | 5'-CTCTCGGGTGGAGTCTTCTG-3' | |||
| CD90 | NM_006288.3 | 185 | F | 5'-GACAGCCTGAGAGGGTCTTG-3' |
| R | 5'-CCCAGTGAAGATGCAGGTTT-3' | |||
| CD117 | NM_000222.2 | 218 | F | 5'-TGCTTCACAGAAGACCATGC-3' |
| R | 5'-GTGACCAACATGGAGTCGTG-3' | |||
| TAT | NM_000353.1 | 236 | F | 5'-TAGCTTCTAGGGGTGCCTCA-3' |
| R | 5'-AGCCATTGTGGACAACATGA-3' | |||
| GS | NM_002065.4 | 194 | F | 5'-TTGCAAGTCATCCTGCAAAG-3' |
| R | 5'-TGATCCTAAGCCCATTCCTG-3' | |||
| G6PD | NM_000151.1 | 140 | F | 5'-CCTGTAACCTGTGAGACTGG-3' |
| R | 5'-ATTCAAGCACCGAAATCTGTAG-3' | |||
| GAPDH | NM_002046.3 | 113 | F | 5'-CTCTCTGCTCCTCCTGTTCG-3' |
| R | 5'-ACGACCAAATCCGTTGACTC-3' | |||
Note: ALB, albumin; AFP, α-fetoprotein; CK8, cytokeratin 8; CK18, cytokeratin 18; CK19, cytokeratin 19; CD34, cluster of differentiation 34; CD45, cluster of differentiation 45; CD90, also known as Thy-1; CD117, also known as c-kit or stem cell factor receptor; TAT, tyrosine aminotransferase; GS, glutamine synthase; G6PD, glucose-6-phosphate dehydrogenase; and GAPDH, glyceraldehyde phosphate dehydrogenase.
Quantitative real-time RT-PCR analysis
We used qRT-PCR to confirm the mRNA expression levels of the following genes: 12 previously published marker genes for HPCs and hepatocytes, genes with a greater than 20-fold up-regulation in HPCs according to the microarray data, and cluster of differentiation (CD) antigen genes that showed between a 2-fold and a 20-fold up-regulation in HPCs or that were positive in HPCs but negative in hepatocytes. The total RNA used in used in the microarray analysis was reverse transcribed, and cDNA was synthesized using a QuantiTect Reverse Transcription Kit. The qRT-PCR reactions were performed with Platinum® Sybr® Green qPCR SuperMix-UDG (Invitrogen) as previously described (24). Thermal cycling was performed in the ABI 7900HT Fast Real-Time PCR system (Applied Biosystems, CA, USA) under the following conditions: 50 °C for 2 min, 95 °C for 10 min, and 35 cycles whose condition depended on the primers used (Tables 2 and 3). The target genes were assayed in five individual specimens and in triplicate on each plate. On each plate, GAPDH was used to normalize differences in cDNA concentration between samples.
Immunocytochemical characterization of HPCs
After the microarray and qRT-PCR analyses, seven previously reported marker proteins for HPCs and hepatocytes and two potential markers found in this study were measured by immunocytochemistry to characterize the HPC phenotype. HPCs were analyzed directly in the culture dishes by immunofluorescence and staining with 3,3'-diaminobenzidine (DAB) with a FITC-conjugated or biotinylated secondary antibody and streptavidin-HRP. The culture medium was removed, and the cells were rinsed with PBS three times and fixed with 4% paraformaldehyde for 30 min at RT. The HPCs were then washed three times with PBS and permeabilized with 80% methanol for 15 min at -20 °C. Following the PBS washes, the cells were incubated with a blocking solution (PBS with 5% FBS) for 30 min at RT. The cells were then incubated overnight with primary antibodies (Table 4), followed by incubation with nine fluorochrome-coupled goat anti-mouse or mouse anti-goat IgG antibodies at 37°C for 1 hr. Between each antibody incubation step, the cells were washed three times with PBS. Omission of the primary antibodies or substitution with nonimmune goat, mouse, and rabbit IgG was used as control where appropriate. The fluorescently labeled cells were visualized by confocal laser scanning microscopy.
Primers used for human HPC gene amplification
| Gene | NCBI accession number | Gene Title | Length | Primer | |
|---|---|---|---|---|---|
| PTX3 | NM_002852.3 | pentraxin-related gene, rapidly induced by IL-1 beta | 180 | F | TCAGGCTTTCCTCAGCATTT |
| R | CCAACACTGCAGACCAGAGA | ||||
| COL5A2 | NM_000393.3 | collagen, type V, alpha 2 | 177 | F | GGAAATGTGGGCAAGACTGT |
| R | TTGATGGTGGTGCTCATTGT | ||||
| COL1A1 | NM000088.3 | collagen, type I, alpha 1 | 214 | F | CCAAATCTGTCTCCCCAGAA |
| R | TCAAAAACGAAGGGGAGATG | ||||
| EFEMP1 | NM_004150.3 | EGF-containing fibulin-like extracellular matrix protein 1 | 212 | F | CAGGACACCGAAGAAACCAT |
| R | GTTTCCTGCTGAGGCTGTTC | ||||
| DKK3 | NM_013253.4 | dickkopf homolog 3 (Xenopus laevis) | 238 | F | TTCATCCAGCAGTGTTGCTC |
| R | GGTGTGGGGTAGTGGAGAGA | ||||
| ACTA2 | NM_001613 | actin, alpha 2, smooth muscle, aorta | 222 | F | TTCAATGTCCCAGCCATGTA |
| R | GAAGGAATAGCCACGCTCAG | ||||
| SLC16A4 | NM_004696 | solute carrier family 16, member 4 (monocarboxylic acid transporter 5) | 198 | F | TGGGATGGGACTGACTTTTC |
| R | CCATGTGCAGACAAACTGCT | ||||
| TPBG | NM_006670.4 | trophoblast glycoprotein | 185 | F | CTGGCTCAAGGAAACAGAGG |
| R | TAGCGCCTATCAGGGCTAAA | ||||
| MAP1B | NM_005909.3 | microtubule-associated protein 1B | 204 | F | ACTAAGCACCACCCATCCTG |
| R | GGAGATGGGACTTCCACTGA | ||||
| GJA1 | NM_000165.3 | gap junction protein, alpha 1, 43kDa | 249 | F | ATGAGCAGTCTGCCTTTCGT |
| R | TCTGCTTCAAGTGCATGTCC | ||||
| CYBRD1 | NM_024843.3 | cytochrome b reductase 1 | 150 | F | GTCACACGGCTCATACATGG |
| R | CTACACCCCACTCCAGCAAT | ||||
| BCAT1 | NM_005504.5 | branched chain aminotransferase 1, cytosolic | 192 | F | CACACAGCAGGAAGAGGTGA |
| R | CTCATAAGGAGCGCATAGCC | ||||
| ARMCX2 | NM_014782.5 | armadillo repeat containing, X-linked 2 | 156 | F | TCTGCTCTGGACACAGTTGG |
| R | TATTGCAGAAGCCATTGCAG | ||||
| SPARC | NM-003118.2 | secreted protein, acidic, cysteine-rich (osteonectin) | 172 | F | GTGCAGAGGAAACCGAAGAG |
| R | TCATTGCTGCACACCTTCTC | ||||
| ID4 | NM_001546.2 | inhibitor of DNA binding 4, dominant negative helix-loop-helix protein | 224 | F | ATGGGATGAGGAAATGCTTG |
| R | TGGAGGAAGGAAAGCAGAAA | ||||
| PROCR | NM_006404 | protein C receptor, endothelial (EPCR) | 211 | F | GGTGTGGCTGTAGGCATCTT |
| R | CTCCCCTCCCTCAAATCTTC | ||||
| PDGFD | NM_025208 | platelet derived growth factor D | 172 | F | GTGGAGGAAATTGTGGCTGT |
| R | CGTTCATGGTGATCCAACTG | ||||
| COL4A2 | NM_001846.2 | collagen, type IV, alpha 2 | 204 | F | AAGGAATCATGGGCTTTCCT |
| R | CTCTGGCACCTTTTGCTAGG | ||||
| TIMP2 | NM_003255.4 | TIMP metallopeptidase inhibitor 2 | 223 | F | TGATCCACACACGTTGGTCT |
| R | TTTGAGTTGCTTGCAGGATG | ||||
| PAM | NM_000919.2 | peptidylglycine alpha-amidating monooxygenase | 194 | F | TTGCTCTTTGCAGTGAATGG |
| R | CACACGGTGTTGGTATGAGC | ||||
Primers used for human HPC gene amplification
| Gene | NCBI accession number | Length | Primer | |
|---|---|---|---|---|
| CD24 | NM_013230.2 | 188 | F | AACTAATGCCACCACCAAGG |
| R | CCTGTTTTTCCTTGCCACAT | |||
| CD44 | NM_000610.3 | 233 | F | AGCAACCAAGAGGCAAGAAA |
| R | GTGTGGTTGAAATGGTGCTG | |||
| CD99 | NM_002414.3 | 209 | F | CTGGGCGGATGATGTTTACT |
| R | TCGATGGACACGTGATTTGT | |||
| CD109 | NM_133493.3 | 193 | F | TGTCTCCTTCCCACATCCTC |
| R | CAGCTTCTTTCCCAAACTGC | |||
| CD200 | NM_005944.5 | 153 | F | TGACCCAGCCCTATTTTACG |
| R | GGGAAAAGTTAACGCATGGA | |||
| CD320 | NM_016579.2 | 153 | F | GGTCCCTGGACACTCCCTAT |
| R | TTTAATCGCCACCCTCAGAC | |||
Antibodies used in immunocyto/histochemistry
| Antigen | Isotype | Dilution | Manufacturer | |
|---|---|---|---|---|
| Immunocytochemistry | Immunohistochemistry | |||
| Oval-6 | Monoclonal mouse IGg1 | 1:100 | 1:200 | Santa Cruz Biotechnology |
| ALB | Monoclonal goat IGg1 | 1:500 | 1:500 | Bethyl Laboratories |
| CK 8 | Monoclonal mouse IGg1 | 1:200 | 1:200 | Santa Cruz Biotechnology |
| CK 18 | Monoclonal mouse IGg1 | 1:200 | 1:200 | Santa Cruz Biotechnology |
| CK 19 | Monoclonal mouse IGg2a | 1:200 | 1:200 | Santa Cruz Biotechnology |
| CD45 | Monoclonal mouse IGg1 | 1:100 | 1:200 | Santa Cruz Biotechnology |
| CD90 | Polyclonal rabbit IGg | 1:400 | 1:400 | Santa Cruz Biotechnology |
| CD109 | Polyclonal rabbit or goat IGg | 1:100 | 1:200 | Santa Cruz Biotechnology |
| CD117(c-Kit) | Polyclonal rabbit or goat IGg | 1:100 | 1:300 | Santa Cruz Biotechnology |
Immunohistochemical localization of HPCs in human liver tissue
To validate potential HPC markers expressed in liver tissue, the same antibodies (except for ALB, because albumin is co-expressed in hepatocytes and HPCs) used for immunocytochemistry were used to detect the localization of HPCs in human liver tissue. Liver segments (1 to 2 g) were fixed with formalin and embedded in paraffin before sectioning. The sections (5-μm thick) were applied to poly-L-lysine-coated slides. Following an inactivation of endogenous peroxidases with 3% H2O2 for 10 min at RT, the sections were incubated overnight with primary antibodies (Table 4). Control experiments included omission of the primary antibodies and substitution of the primary antibodies with nonimmune rabbit or mouse IgG. Subsequently, the sections were washed with PBS three times and incubated with the appropriate secondary antibodies at 37°C for 1 hr. After the reaction with DAB Chromogen, the sections were rinsed with distilled water, counterstained with hematoxylin, and mounted with Histomount solution (Invitrogen), and the labeled cells were visualized with an inverted microscope (Nikon, Eclipse E200, Tokyo, Japan), and digital images were captured using Nis-elements F 3.0 software. Omission of the primary antibody or substitution with an unrelated IgG served as a negative control.
Statistical analysis
Only those genes detected as present or absent in triplicate arrays were reported as expressed or non-expressed genes. The data were analyzed by the chi-square test using SPSS software version 16.0. The level of significance for all statistical analyses was P < 0.05.
Results
Morphological characteristics of HPCs
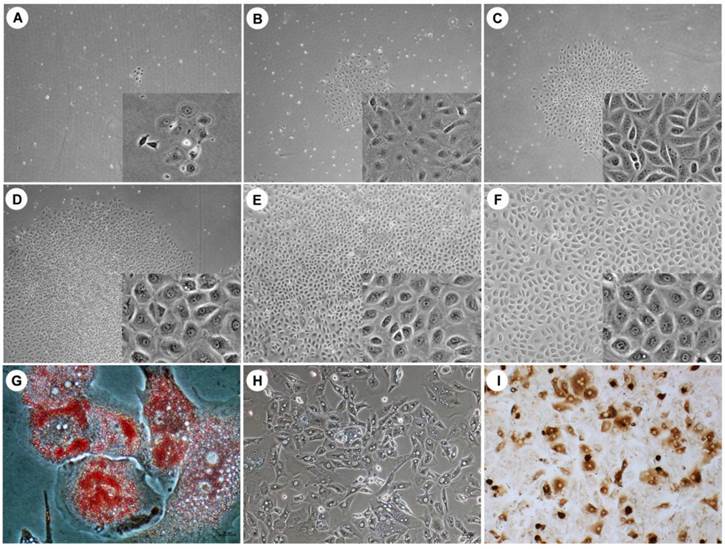
Many small, scattered and flattened colonies were observed under phase-contrast microscopy (Fig. 1) after an initial 7 days of culture. These loose colony cells displayed several morphologic characteristics of HPCs, such as a round or oval shape and a convex cytoplasm. These cells proliferated slowly and formed sharply bordered colonies (Fig. 1A-C). After 10 days of culture, the cells had a polygonal shape with a low cytoplasm/nucleus ratio and started to expand (Fig. 1D). The colony-forming cells reached confluence after 2 weeks (Fig. 1E, F). There was no evidence that the human hepatocytes were contaminated, as evaluated by phase-contrast microscopy. All cultured HPCs prepared from the different liver specimens exhibited similar morphologic characteristics.
Bipotential ability
The bipotential HPC characteristics were demonstrated via culture in multilineage differentiation conditions in vitro. The adipogenic differentiation of the HPCs was demonstrated by Oil Red O staining, and lipid droplets were visible in the differentiated adipocytes on day 10 after the induction of differentiation (Fig. 1G). Hepatogenesis was characterized by morphology. Under phase-contrast microscopy, the differentiated cells showed hepatocyte-like polygonal morphology with low cytoplasm/nucleus ratios (Fig. 1H). Cholangiogenic identification indicated that differentiated cholangiocytes were positive for CK19 (Fig. 1I). These results demonstrated that the isolated and cultured HPCs exhibited the classic bipotential progenitor cell characteristics.
Gene transcriptional profiles
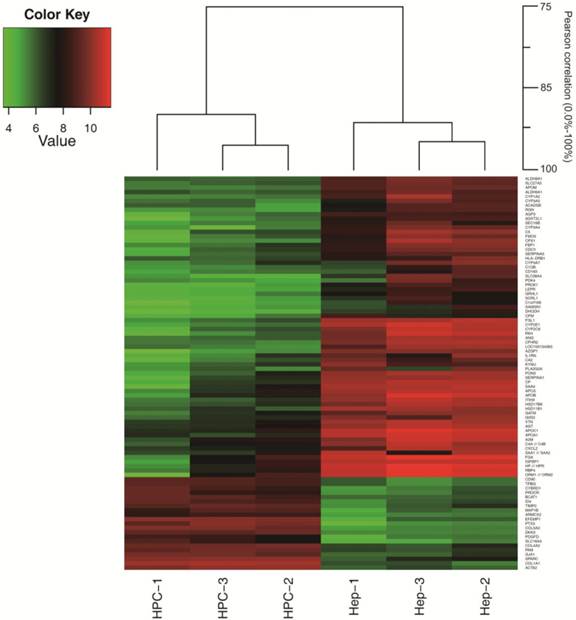
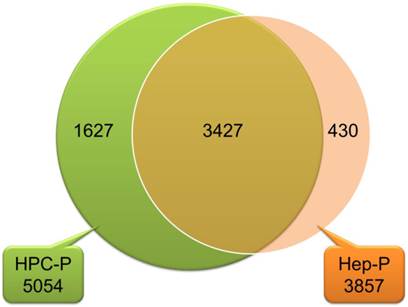
The Pearson correlation coefficient for comparing gene expression profiles of the same cell types, isolated from different liver specimens, was 0.90 to 0.96 for HPCs and 0.91 to 0.95 for hepatocytes (Fig. 2). These data indicate that there was great similarity in gene expression between the biological replicates. As shown in the Venn diagram in Fig. 3, HPCs and human hepatocytes share a similar transcriptome. Among the 5,054 probe sets/genes present in HPCs, 67.8% (3427/5054) of the probe sets/genes were also expressed in the primary hepatocytes. Conversely, when this similarity was represented as a percentage of the probe sets/genes in the primary hepatocytes that were also expressed in the HPCs, the similarity reached 88.9% (3427/3857). These values indicate a high degree of similarity between the gene transcriptional profiles of HPCs and primary hepatocytes.
Further analysis of global gene expression showed that 21 probe sets/genes (Table 5) were up-regulated by more than 20-fold in HPCs compared to primary hepatocytes. Among those genes were a progenitor cell marker (CD90, 39.3-fold), pentraxin-related gene (PTX3, 120.5-fold), collagen proteins (COL5A2, 88.4-fold; COL1A1, 80.9-fold; and COL4A2, 20.8-fold), cytokines (EGF, 74.1-fold and PDGFD, 21.6-fold), hepatocyte-related metabolic enzymes (CYBRD1, 27.3-fold; BCAT1, 26.2-fold; TIMP2, 20.7-fold and PAM, 20.4-fold), a secreted protein (SPARC, 23.8-fold) and the endothelial protein C receptor (PROCR, 22.1-fold). Among 12 previously reported marker genes for HPCs and hepatocytes (Table 6) that were analyzed in this study, 10 genes were expressed in both HPCs and hepatocytes, including CD90 (with the above-mentioned 39.3-fold relative up-regulation in HPCs) and four genes whose expression was more than 2-fold but less than 20-fold in HPCs relative to primary hepatocytes (CK19, 18.2-fold; CD45, 6.1-fold; CD117, 5.2-fold; and CK8, 2.8-fold). CK18 was expressed at the same level (1.0-fold) in both cells. In contrast, AFP and ALB were expressed at a lower level in HPCs than in hepatocytes (AFP, 0.3-fold and ALB, 0.2-fold). Three other mature hepatocyte marker genes (GS, TAT and G6PD) were absent in HPCs but present in hepatocytes. CD34 was absent in both cells. Moreover, CD109 genes were also expressed more than 2-fold (2.5-fold) in HPCs compared to hepatocytes. Five other surface marker genes (CD24, CD44, CD99, CD200 and CD320) were present in HPCs but absent in hepatocytes (Table 7).
The morphology of HPCs and HPCs differentiated into adipocytes, hepatocytes and cholangiocytes. Under phase-contrast microscopy, small and flattened colonies were observed after an initial 7-day culture (A, day 3; B, day 5; C, day 7). The colonies of loosely packed cells displayed a round or oval shape and a convex cytoplasm. After 10 days of culture, these cells exhibited a clear and larger cytoplasm, became polygonal and densely packed, and began to expand (D, day 10; E, day 15; F, day 20). After 10 days differentiation, differentiated adipocytes contained lipid droplets (G), differentiated hepatocytes exhibited a polygonal morphology with a low cytoplasm/nucleus ratio (H) and differentiated cholangiocyte were positive for CK19 (I). Original magnification: large view, A-D, 4x; E-F, 10x; small view, A-F, 20x; G, 40 x; H-I, 20 x.
A hierarchical cluster dendrogram of selected genes. The Pearson correlation coefficient was 0.90 to 0.96 for HPCs and 0.91 to 0.95 for hepatocytes. HPC, hepatic progenitor cells; Hep, hepatocytes.
A Venn diagram showing that HPCs and primary human hepatocytes had similar transcriptomes. Among the 5,054 probe sets/genes present in HPCs, 67.8% (3427/5054) probe sets/genes were also expressed in primary hepatocytes. Conversely, among 3,857 probe sets/genes expressed in primary hepatocytes, 88.9% (3427/3857) were also expressed in HPCs. HPC-P 5,054, total number of probe sets/genes present in HPCs; Hep-P3857, total number of probe sets/genes present in primary hepatocytes.
Genes with >20-fold up-regulation in HPCs
| Probe ID | Gene | Representative Public ID | Fold* | Gene Title |
|---|---|---|---|---|
| 206157_at | PTX3 | NM_002852 | 120.5 ±13.6 | pentraxin-related gene, rapidly induced by IL-1 beta |
| 221729_at | COL5A2 | AL575735 | 88.4 ±10.5 | collagen, type V, alpha 2 |
| 1556499_s_at | COL1A1 | BE221212 | 80.9 ±6.2 | collagen, type I, alpha 1 |
| 201842_s_at | EFEMP1 | AI826799 | 74.1 ±8.2 | EGF-containing fibulin-like extracellular matrix protein 1 |
| 214247_s_at | DKK3 | AU148057 | 57.2 ±7.42 | dickkopf homolog 3 (Xenopus laevis) |
| 200974_at | ACTA2 | NM_001613 | 55.0 ±6.5 | actin, alpha 2, smooth muscle, aorta |
| 213869_x_at | CD90 | AL558479 | 39.3 ±4.2 | Thy-1 cell surface antigen |
| 205234_at | SLC16A4 | NM_004696 | 35.3 ±4.9 | solute carrier family 16, member 4 (monocarboxylic acid transporter 5) |
| 203476_at | TPBG | NM_006670 | 29.2 ±4.3 | trophoblast glycoprotein |
| 212233_at | MAP1B | AL523076 | 28.9 ±4.2 | microtubule-associated protein 1B |
| 201667_at | GJA1 | NM_000165 | 27.7 ±3.3 | gap junction protein, alpha 1, 43kDa |
| 222453_at | CYBRD1 | AL136693 | 27.3 ±3.9 | cytochrome b reductase 1 |
| 226517_at | BCAT1 | AL390172 | 26.2 ±3.7 | branched chain aminotransferase 1, cytosolic |
| 203404_at | ARMCX2 | NM_014782 | 24.1 ±3.5 | armadillo repeat containing, X-linked 2 |
| 200665_s_at | SPARC | NM_003118 | 23.8 ±4.2 | secreted protein, acidic, cysteine-rich (osteonectin) |
| 209291_at | ID4 | AW157094 | 22.5 ±4.6 | inhibitor of DNA binding 4, dominant negative helix-loop-helix protein |
| 203650_at | PROCR | NM_006404 | 22.1 ±4.01 | protein C receptor, endothelial (EPCR) |
| 219304_s_at | PDGFD | NM_025208 | 21.6 ±3.9 | platelet derived growth factor D |
| 211964_at | COL4A2 | X05610 | 20.8 ±4.2 | collagen, type IV, alpha 2 |
| 224560_at | TIMP2 | BF107565 | 20.7 ±3.1 | TIMP metallopeptidase inhibitor 2 |
| 202336_s_at | PAM | NM_000919 | 20.4 ±3.3 | peptidylglycine alpha-amidating monooxygenase |
*Fold increase in hepatic progenitor cells vs. hepatocytes. The fold change between the two groups was calculated from the normalized average mRNA expression level in each group.
Microarray data validated by qRT-PCR: the fold change of mRNA expression between HPCs and hepatocytes
| Gene | Microarray | qRT-PCR | Gene Title | ||
|---|---|---|---|---|---|
| HPC | Hep | Fold (M ± SD) a | Fold (M ± SD) b | ||
| CD90 | P | P | 39.3 ±4.2 | 21.5 ± 4.8 | also known as Thy-1 |
| CK19 | P | P | 18.2 ± 2.5 | 12.5 ± 2.4 | cytokeratin 19 |
| CD45 | P | P | 6.1 ± 1.5 | 3.2 ± 0.5 | CD45 |
| CD117 | P | P | 5.2 ± 1.3 | 2.5 ± 1.4 | also known as c-kit, stem cell factor receptor |
| CK8 | P | P | 2.8 ± 0.5 | 2.1 ± 0.2 | cytokeratin 8 |
| CK18 | P | P | 1.0 ± 0.3 | 2.3 ± 0.1 | cytokeratin 18 |
| AFP | P | P | 0.3 ± 0.1 | 0.1 ± 0.01 | a-fetoprotein |
| ALB | P | P | 0.2 ± 0.01 | 0.1 ± 0.01 | albumin |
| GS | A | P | N/A | A/A | glutamine synthase |
| TAT | A | P | N/A | A/A | tyrosine aminotransferase |
| G6PD | A | P | N/A | A/A | glucose-6-phosphate dehydrogenase |
| CD34 | A | A | N/A | A/A | CD34 |
a: The fold change in mRNA expression between HPCs and hepatocytes as determined by microarray analysis was calculated from three chips in each group. b: The fold change in mRNA expression between HPCs and hepatocytes as measured by real-time RT-PCR; the results of qRT-PCR are representative of five individual samples, and PCR was performed in triplicate for each sample. A, absent; P, present; N/A, not applicable; A/A, absent from both kinds of cells.
Microarray data validated by qRT-PCR: the fold change of mRNA expression between HPCs and hepatocytes
| Gene | Microarray | qRT-PCR | ||
|---|---|---|---|---|
| HPC | Hep | Fold (M ± SD) a | Fold (M ± SD) b | |
| CD24 | P | A | N/A | A/A |
| CD44 | P | A | N/A | A/A |
| CD99 | P | A | N/A | A/A |
| CD109 | P | P | 2.5 ± 0.6 | 3.4 ± 0.5 |
| CD200 | P | A | N/A | A/A |
| CD320 | P | A | N/A | A/A |
a: The fold change in mRNA expression between HPCs and hepatocytes as determined by microarray analysis was calculated from three chips in each group. b: The fold change in mRNA expression between HPCs and hepatocytes as measured by real-time RT-PCR; the qRT-PCR results represent five individual samples, and PCR was performed in triplicate for each sample. A, absent; P, present; N/A, not applicable; A/A, absent from both kinds of cells.
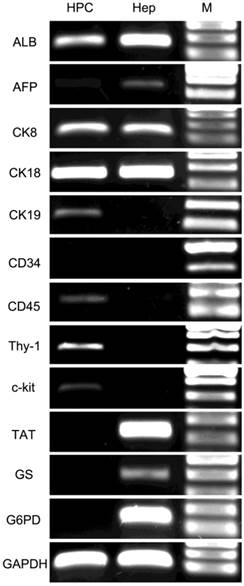
RT-PCR showed that the colony-forming cells were positive for several previously reported marker genes for HPCs and hepatocytes (Fig. 4). The hepatocyte marker genes ALB, CK8 and CK18 exhibited strong expression in both cells, whereas AFP was expressed weakly in HPCs. The cholangiocyte marker genes CK7 and CK19 were expressed at an intermediate level in HPCs but were negative in hepatocytes. The progenitor marker gene CD90 exhibited strong expression, and CD45, CD117 and vimentin (VIM) exhibited intermediate expression in HPCs but were negative in hepatocytes. The hematopoietic cell marker CD34 and mature hepatocyte markers TAT, GS and G6PD were negative in HPCs and strongly positive in hepatocytes. These results were consistent with the microarray findings.
RT-PCR analysis of HPC-specific genes. HPCs were positive for ALB, AFP (weakly), CK8, CK18, CK19, CD45, CD90 and CD117 but negative for CD34, TAT, GS and G6PD. Lane HPC, samples from HPCs; Lane Hep, samples from primary hepatocytes. Lane M, DNA marker.
Validation of mRNA expression of selected genes by qRT-PCR
To validate the results of the microarray analysis, 21 enriched genes with a greater than 20-fold up-regulation, 12 HPC and hepatocyte marker genes described in the literature and six surface marker genes present in HPCs were further analyzed by qRT-PCR. The validation experiment for the 21 HPC-enriched genes showed similar results of microarray (Table 8) in that these genes experienced a 6.5-fold to 78.1-fold up-regulation. Similarly, for the 12 previously described HPC and hepatocyte markers (Table 6), six presented a 2.1-fold to 21.5-fold up-regulation in HPCs (CD90, 21.5-fold; CK19, 12.5-fold; CD45, 3.2-fold; CD117, 2.5-fold; CK8, 2.1-fold; CK18, 2.3-fold), two were weakly expressed in HPCs compared to hepatocytes (AFP, 0.1-fold; ALB, 0.1-fold), and 4 (CD34, TAT, GS and G6PD) were absent in HPCs. The validation of six surface marker genes (Table 7) showed that CD109 presented a 3.4-fold up-regulation in HPCs compared to hepatocytes, while the other five genes (CD24, CD44, CD99, CD200 and CD320) were absent in both cell types. These data indicate a high degree of consistency between the microarray and qRT-PCR analyses.
Microarray data validated by qRT-PCR: the fold change of mRNA expression between HPCs and hepatocytes.
| Gene | Microarray | qRT-PCR | Gene Title |
|---|---|---|---|
| Fold (M ± SD) a | Fold (M ± SD) b | ||
| PTX3 | 120.5 ±13.6 | 78.1±10.5 | pentraxin-related gene, rapidly induced by IL-1 beta |
| COL5A2 | 88.4 ±10.5 | 43.6 ±7.2 | collagen, type V, alpha 2 |
| COL1A1 | 80.9 ±6.2 | 41.1 ±6.1 | collagen, type I, alpha 1 |
| EFEMP1 | 74.1 ±8.2 | 46.2 ±8.3 | EGF-containing fibulin-like extracellular matrix protein 1 |
| DKK3 | 57.2 ±7.42 | 36.5 ±4.6 | dickkopf homolog 3 (Xenopus laevis) |
| ACTA2 | 55.0 ±6.5 | 48.2 ±4.3 | actin, alpha 2, smooth muscle, aorta |
| CD90 | 39.3 ±4.2 | 21.5 ±3.2 | Thy-1 cell surface antigen |
| SLC16A4 | 35.3 ±4.9 | 17.1 ±4.2 | solute carrier family 16, member 4 (monocarboxylic acid transporter 5) |
| TPBG | 29.2 ±4.3 | 16.7 ±2.8 | trophoblast glycoprotein |
| MAP1B | 28.9 ±4.2 | 18.1 ±3.4 | microtubule-associated protein 1B |
| GJA1 | 27.7 ±3.3 | 15.8 ±4.6 | gap junction protein, alpha 1, 43kDa |
| CYBRD1 | 27.3 ±3.9 | 15.1 ±2.9 | cytochrome b reductase 1 |
| BCAT1 | 26.2 ±3.7 | 9.6 ±3.1 | branched chain aminotransferase 1, cytosolic |
| ARMCX2 | 24.1 ±3.5 | 18.2 ±3.7 | armadillo repeat containing, X-linked 2 |
| SPARC | 23.8 ±4.2 | 19.0 ±4.2 | secreted protein, acidic, cysteine-rich (osteonectin) |
| ID4 | 22.5 ±4.6 | 7.4±1.8 | inhibitor of DNA binding 4, dominant negative helix-loop-helix protein |
| PROCR | 22.1 ±4.01 | 6.5±1.4 | protein C receptor, endothelial (EPCR) |
| PDGFD | 21.6 ±3.9 | 6.9±2.1 | platelet derived growth factor D |
| COL4A2 | 20.8 ±4.2 | 15.7±3.3 | collagen, type IV, alpha 2 |
| TIMP2 | 20.7 ±3.1 | 12.7±2.4 | TIMP metallopeptidase inhibitor 2 |
| PAM | 20.4 ±3.3 | 13.2±2.8 | peptidylglycine alpha-amidating monooxygenase |
a: The fold change in mRNA expression between HPCs and hepatocytes as determined by microarray analysis was calculated from three chips in each group. b: The fold change in mRNA expression between HPCs and hepatocytes as measured by real-time RT-PCR; the results of qRT-PCR are representative of five individual samples, and PCR was performed in triplicate for each sample.
Immunocyto/histochemistry expression analysis
Immunocytochemical characterization of cultured HPCs (Fig. 5) showed that these cells were consistently positive for the progenitor-specific markers Oval-6, CD90 and CD117; hepatocyte-specific markers CK8, CK18 and ALB; and cholangiocyte-specific markers CK19. Cultured HPCs were also positive for the hematopoietic cell markers CD45 and CD109 after 3 weeks of culture. The images of negative control are showed in the Fig. 6.
To characterize the localization of HPCs within normal liver tissue, the same markers used for immunocytochemistry were analyzed in individual liver specimens from which HPCs and primary hepatocytes had been isolated. Immunohistochemical staining of the liver specimens (Fig. 7) indicated that most of the cells positive for 1 of the 8 selected markers were localized in periportal areas and the canal of Hering. The progenitor-specific markers Oval-6 (Fig. 7A), CD90 (Fig. 7F) and CD117 (Fig. 7H) labeled cells that were scattered in small ductular regions/canals of Hering, and these cells were not associated with bile ductule-like structures. The cholangiocyte-specific markers CK19 (Fig. 7C) labeled cells that were scattered in the periportal area, and some of these cells resembled bile ductule walls. Although the CK8 (Fig. 7D) and CK18 (Fig. 7B) stained weakly throughout most of each liver section, the cells highly expressing these two markers could easily be distinguished from surrounding hepatocytes. The cells that expressed the hematopoietic cell markers CD45 (Fig. 7E) and CD109 (Fig. 7G) appeared similar to the CK18-positive cells located in the periportal areas and the canals of Hering. CD45 (see Table 6 and Fig. 7E) and CD109 (see Table 7 and Fig. 7G) were commonly expressed in hepatocytes and progenitor-like cells. The images of negative control are showed in the Fig. 8.
Immunocytochemical staining showing that the colony-forming cells were consistently positive for the progenitor-specific markers Oval-6 (A), CD90 (F) and CD117 (H); the hepatocyte-specific markers ALB (I), CK18 (B) and CK8 (D); and the cholangiocyte-specific markers CK19 (C). These cells were also positive for two hematopoietic cell markers CD45 (E) and CD109 (G). Original magnification, 20x.
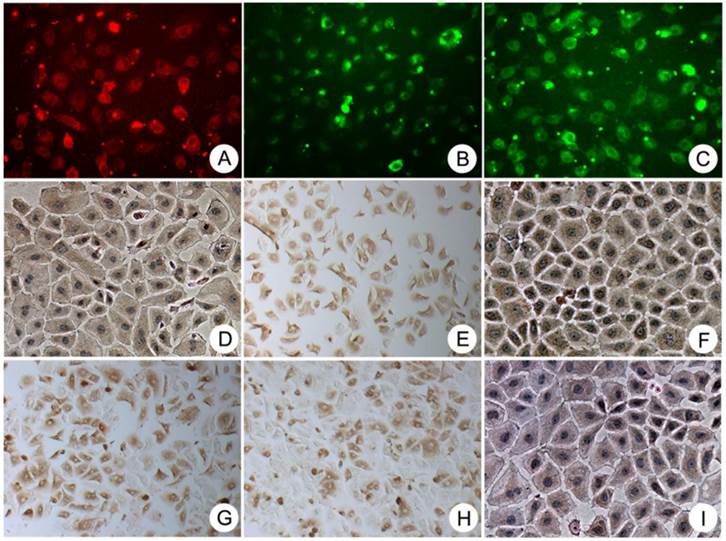
The negative control of immunocytochemical staining. A-I were Oval-6, CK18, CK19, CK8, CD45, CD90, CD109, CD117 and ALB respectively. Original magnification, 20x.
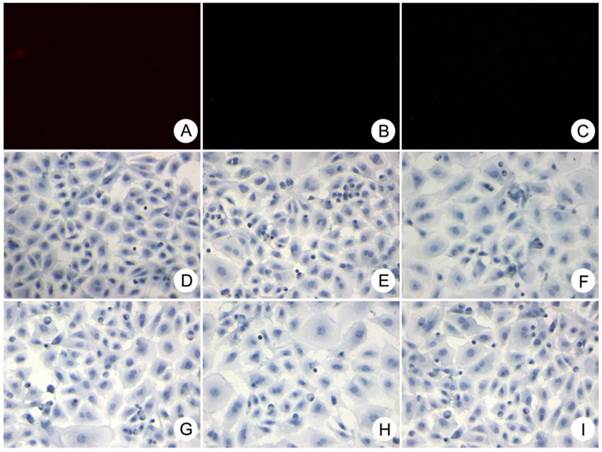
Immunohistological staining showing that cells positive for the progenitor-specific markers Oval-6 (A), CD90 (F) and CD117 (H) were scattered in the small ductular region/canals of Hering and that these cells were not associated with bile ductule-like structures. Cells positive for the cholangiocyte-specific marker CK19 (C) were scattered in the periportal area, and some of these cells resembled the walls of bile ductules. CK8 (D) and CK18 (B) were weakly expressed in hepatocytes and strongly expressed in HPCs. The cells expressing hematopoietic cell markers CD45 (E) and CD109 (G) were located in the periportal areas and the canals of Hering. Black arrow: specific marker positive HPCs. White arrow in C: specific marker positive cells resembled bile ductule wall. White arrow in B and D: specific marker positive hepatocytes. Original magnification: 20x.
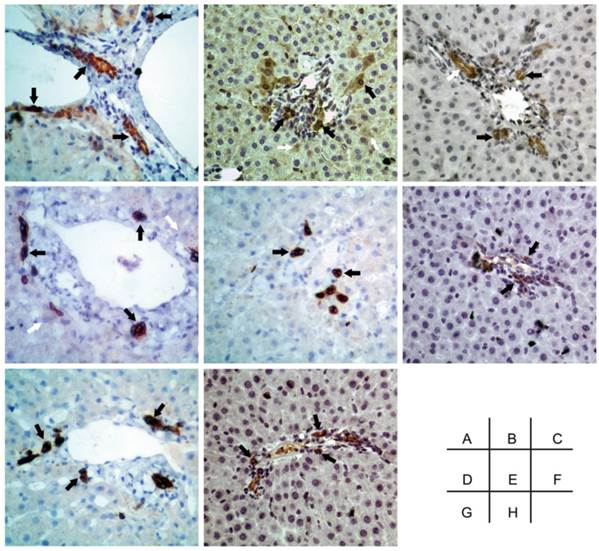
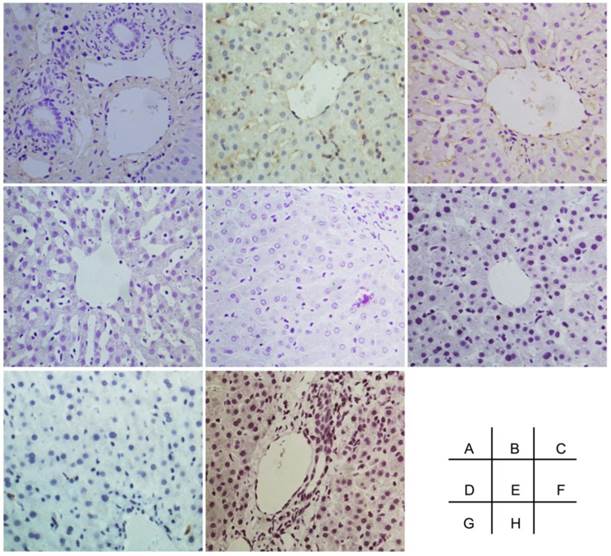
The negative control of immunohistochemical staining. A-I were Oval-6, CK18, CK19, CK8, CD45, CD90, CD109 and CD117 respectively. Original magnification, 20x.
Discussion
Numerous studies have shown that HPCs mainly participate in liver regeneration following acute and chronic hepatic damage (11, 16, 40). Although many reports have demonstrated that progenitor cell populations can be isolated from injured rodent and dog livers (25, 41-43), only a few research groups have successfully isolated HPCs from adult human liver (30-36). The distinction between hepatocytes and HPCs in human liver is controversial.
A summary of previously published phenotypic characteristics of human HPCs is shown in Table 9. A total of 38 potential marker genes have been identified in human HPCs in seven publications. Herrera et al. (30) isolated and characterized a population of human liver stem cells that were positive for CD29, CD44, CD73, CD90, VIM, nestin, ALB, AFP, CK8 and CK18 and negative for CK19, CD34, CD45, CD117 and CD133. Schmelzer et al. (32) characterized the phenotypes of pluripotent human hepatic progenitors, which were positive for CK19, NCAM, EpCAM, CLDN3 and ALB and negative for AFP and adult liver-specific proteins. Dan et al. (31) described multipotent progenitor cells isolated from human fetal liver that expressed CD34, CD90, CD117, EpCAM, c-met, SSEA-4, VIM, CK18 and CK19 but not ALB or AFP. Duret et al. (33) characterized a population of nonparenchymal epithelial cells from adult human liver that expressed AAT, CK7, CK18 and CK19 as well as a low level of ALB but not REX1, AFP, CD34, CD90, CD117 or Oval-6. Terrace et al. (34) described HPCs isolated from developing human liver that were positive for CK18, CK19, ALB, E-cadherin, dlk/pref-1, CD90, CD34 and VIM, and these cells were located in the portal vein endothelium. Stachelscheid et al. (35) and Jozefczuk et al. (36) isolated and characterized adult human liver progenitors from ischemic liver tissue; these cells were positive for EpCAM, CK7, CK8, CK18, CK19, VIM, CYP2B6, ALB, A1AT, γ-GT and HNF4 but negative for AFP and CD90.
In this study, the HPCs isolated from adult human liver exhibited morphological characteristics similar to those described previously (33). These cells possessed bipotential differentiation characteristics formed colonies after 3 days and presented polygonal, densely packed cytoplasm after 10 days in culture. Global gene expression analysis of these cells revealed a high similarity (67.8%) to primary hepatocytes. Compared with primary hepatocytes, 21 genes up-regulated more than 20-fold in HPCs were detected, including the progenitor marker CD90, four genes related to hepatocyte metabolism (CYBRD1, BCAT1, TIMP2 and PAM) and two cytokine genes (EGF, PDGFD). The HPC-enriched genes detected by microarray analysis were validated by qRT-PCR, with consistent results. Further analysis showed that most previously reported marker genes for HPCs (ALB, AFP, CK8, CK18, CK19, CD90, CD117 and Oval-6) were positive in our cultured HPCs. Moreover, two hematopoietic cell markers CD45 and CD109 were detected in HPCs. Five cell-surface markers and three mature hepatocyte-specific genes were not detected in HPCs by microarray or qRT-PCR analysis. The potential markers mentioned above were also confirmed at the protein level by immunocyto/immunohistochemistry, with consistent results. Although the morphology of the cells expressing Oval-6 and CD90 (Fig 6A and F) in human livers seems different from that of the cells expressing CD45 and CD109 (Fig 6E and G). It is possible that these cells were at the different stages of growth and development.
In summary, our data show that HPCs isolated from adult human liver are highly similar to cultured primary hepatocytes in their transcriptional profiles, and those gene expression data are in accordance with immunohistological staining of hepatocytes in liver sections. In addition to eight established marker genes/proteins (ALB, AFP, CK8, CK18, CK19, CD90, CD117 and Oval-6), the novel markers CD45 and CD109 can potentially be utilized to identify and isolate HPCs for further cytotherapy of liver diseases. Based on our findings, future studies should be conducted to validate these two novel HPC-specific markers and their roles in hepatocyte development and liver regeneration.
Phenotypic characteristics of human HPCs in comparison with described markers in publications
| Reference | Positive | Negative |
|---|---|---|
| Dan et al. (31) | CK18, CK19, CD34, CD90, CD117, EpCAM, Vimentin c-Met, SSEA-4 | ALB, AFP |
| Herrera et al. (30) | CK8, CK18, CD29, CD44, CD73, CD90, Vimentin, Nestin, ALB, AFP | CK19, CD34, CD45, CD117, CD133 |
| Schmelzer et al. (32) | CK19, NCAM, EpCAM, CLDN3, ALB (low) | AFP |
| Duret et al. (33) | CK7, CK18, CK19, ALB (low), AAT | REX1, AFP, CD34, CD90, CD117, OV-6 |
| Terrace et al. (34) | CK18, CK19, CD34, CD90, ALB, E-cadherin, dlk/pref-1, Vimentin | |
| Stachelscheid et al. (35) | CK7, CK8, CK18, CK19, EpCAM, Vimentin, CYP2B6, ALB (intermediate), A1AT (intermediate), γ-GT, HNF4 (low) | AFP, CD90 |
| Jozefczuk et al. (36) | CK7, CK8, CK18, CK19, CD24, CD133, EpCAM, Vimentin, CYP1B1, ABCC4, ANXA3, ANXA1, CLDN4, DDR1, GABRP, ITGB4, MUC1, S100A3, ALB (intermediate), A1AT (intermediate), HNF4 (low) | AFP, CD90 |
| Li, et al. (this study) | CK8, CK18, CK19, CD45, CD90, CD109, CD117, OV-6, ALB, AFP, PTX3, COL5A2, COL1A1, EFEMP1, DKK3, ACTA2, SLC16A4, TPBG, MAP1B, GJA1, CYBRD1, BCAT1, ARMCX2, SPARC, ID4, PROCR, PDGFD, COL4A2, TIMP2, PAM | CD24, CD34, CD44, CD99, CD200, D320, TAT, GS, G6PD |
Acknowledgements
This work was supported by the Zhejiang Provincial and National Natural Science Foundation of China (LR13H030001, 81271708), Chinese High Tech Research and Development (863) Program (2013AA020102), and National S&T Major Project (2012ZX10004503-006, 2012ZX10002004-001).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Fox IJ, Roy-Chowdhury J. Hepatocyte transplantation. J Hepatol. 2004;40:878-886
2. Soltys KA, Soto-Gutierrez A, Nagaya M, Baskin KM, Deutsch M, Ito R, Shneider BL. et al. Barriers to the successful treatment of liver disease by hepatocyte transplantation. J Hepatol. 2010;53:769-774
3. Dong HH, Xiang S, Chen XP, Liang HF, Zhang W, Jing K, Zhang WG. et al. The epithelial-mesenchymal transition promotes transdifferentiation of subcutaneously implanted hepatic oval cells into mesenchymal tumor tissue. Stem Cells Dev. 2009;18:1293-1298
4. Fitzpatrick E, Mitry RR, Dhawan A. Human hepatocyte transplantation: state of the art. J Intern Med. 2009;266:339-357
5. Lee SW, Wang X, Chowdhury NR, Roy-Chowdhury J. Hepatocyte transplantation: state of the art and strategies for overcoming existing hurdles. Ann Hepatol. 2004;3:48-53
6. Ikeda E, Yagi K, Kojima M, Yagyuu T, Ohshima A, Sobajima S, Tadokoro M. et al. Multipotent cells from the human third molar: feasibility of cell-based therapy for liver disease. Differentiation. 2008;76:495-505
7. Brezillon N, Kremsdorf D, Weiss MC. Cell therapy for the diseased liver: from stem cell biology to novel models for hepatotropic human pathogens. Dis Model Mech. 2008;1:113-130
8. Zern MA. Cell transplantation to replace whole liver transplantation. Gastroenterology. 2009;136:767-769
9. Forbes S, Vig P, Poulsom R, Thomas H, Alison M. Hepatic stem cells. J Pathol. 2002;197:510-518
10. Kakinuma S, Nakauchi H, Watanabe M. Hepatic stem/progenitor cells and stem-cell transplantation for the treatment of liver disease. J Gastroenterol. 2009;44:167-172
11. Stutchfield BM, Rashid S, Forbes SJ, Wigmore SJ. Practical barriers to delivering autologous bone marrow stem cell therapy as an adjunct to liver resection. Stem Cells Dev. 2010;19:155-162
12. Strain AJ, Crosby HA, Nijjar S, Kelly DA, Hubscher SG. Human liver-derived stem cells. Semin Liver Dis. 2003;23:373-384
13. Hashemi SM, Soleimani M, Zargarian SS, Haddadi-Asl V, Ahmadbeigi N, Soudi S, Gheisari Y. et al. In vitro differentiation of human cord blood-derived unrestricted somatic stem cells into hepatocyte-like cells on poly(epsilon-caprolactone) nanofiber scaffolds. Cells Tissues Organs. 2009;190:135-149
14. Jochheim-Richter A, Rudrich U, Koczan D, Hillemann T, Tewes S, Petry M, Kispert A. et al. Gene expression analysis identifies novel genes participating in early murine liver development and adult liver regeneration. Differentiation. 2006;74:167-173
15. Moreno R, Martinez-Gonzalez I, Rosal M, Farwati A, Gratacos E, Aran JM. Characterization of mesenchymal stem cells isolated from the rabbit fetal liver. Stem Cells Dev. 2010;19:1579-1588
16. Dalakas E, Newsome PN, Boyle S, Brown R, Pryde A, McCall S, Hayes PC. et al. Bone marrow stem cells contribute to alcohol liver fibrosis in humans. Stem Cells Dev. 2010;19:1417-1425
17. Zhang Y, Bai XF, Huang CX. Hepatic stem cells: existence and origin. World J Gastroenterol. 2003;9:201-204
18. Haridass D, Narain N, Ott M. Hepatocyte transplantation: waiting for stem cells. Curr Opin Organ Transplant. 2008;13:627-632
19. Wang J, Clark JB, Rhee GS, Fair JH, Reid LM, Gerber DA. Proliferation and hepatic differentiation of adult-derived progenitor cells. Cells Tissues Organs. 2003;173:193-203
20. Coradeghini R, Guida C, Scanarotti C, Sanguineti R, Bassi AM, Parodi A, Santi PL. et al. A comparative study of proliferation and hepatic differentiation of human adipose-derived stem cells. Cells Tissues Organs. 2010;191:466-477
21. Gilchrist ES, Plevris JN. Bone marrow-derived stem cells in liver repair: 10 years down the line. Liver Transpl. 2010;16:118-129
22. Vassilopoulos G, Wang PR, Russell DW. Transplanted bone marrow regenerates liver by cell fusion. Nature. 2003;422:901-904
23. Li J, Tao R, Wu W, Cao H, Xin J, Guo J, Jiang L. et al. 3D PLGA scaffolds improve differentiation and function of bone marrow mesenchymal stem cell-derived hepatocytes. Stem Cells Dev. 2010;19:1427-1436
24. Li J, Tao R, Wu W, Cao H, Xin J, Guo J, Jiang L. et al. Transcriptional profiling and hepatogenic potential of acute hepatic failure-derived bone marrow mesenchymal stem cells. Differentiation. 2010;80:166-174
25. Dorrell C, Erker L, Lanxon-Cookson KM, Abraham SL, Victoroff T, Ro S, Canaday PS. et al. Surface markers for the murine oval cell response. Hepatology. 2008;48:1282-1291
26. Jelnes P, Santoni-Rugiu E, Rasmussen M, Friis SL, Nielsen JH, Tygstrup N, Bisgaard HC. Remarkable heterogeneity displayed by oval cells in rat and mouse models of stem cell-mediated liver regeneration. Hepatology. 2007;45:1462-1470
27. Arai M, Yokosuka O, Fukai K, Imazeki F, Chiba T, Sumi H, Kato M. et al. Gene expression profiles in liver regeneration with oval cell induction. Biochem Biophys Res Commun. 2004;317:370-376
28. Petersen BE, Grossbard B, Hatch H, Pi L, Deng J, Scott EW. Mouse A6-positive hepatic oval cells also express several hematopoietic stem cell markers. Hepatology. 2003;37:632-640
29. Engelhardt NV, Factor VM, Medvinsky AL, Baranov VN, Lazareva MN, Poltoranina VS. Common antigen of oval and biliary epithelial cells (A6) is a differentiation marker of epithelial and erythroid cell lineages in early development of the mouse. Differentiation. 1993;55:19-26
30. Herrera MB, Bruno S, Buttiglieri S, Tetta C, Gatti S, Deregibus MC, Bussolati B. et al. Isolation and characterization of a stem cell population from adult human liver. Stem Cells. 2006;24:2840-2850
31. Dan YY, Riehle KJ, Lazaro C, Teoh N, Haque J, Campbell JS, Fausto N. Isolation of multipotent progenitor cells from human fetal liver capable of differentiating into liver and mesenchymal lineages. Proc Natl Acad Sci U S A. 2006;103:9912-9917
32. Schmelzer E, Wauthier E, Reid LM. The phenotypes of pluripotent human hepatic progenitors. Stem Cells. 2006;24:1852-1858
33. Duret C, Gerbal-Chaloin S, Ramos J, Fabre JM, Jacquet E, Navarro F, Blanc P. et al. Isolation, characterization, and differentiation to hepatocyte-like cells of nonparenchymal epithelial cells from adult human liver. Stem Cells. 2007;25:1779-1790
34. Terrace JD, Currie IS, Hay DC, Masson NM, Anderson RA, Forbes SJ, Parks RW. et al. Progenitor cell characterization and location in the developing human liver. Stem Cells Dev. 2007;16:771-778
35. Stachelscheid H, Urbaniak T, Ring A, Spengler B, Gerlach JC, Zeilinger K. Isolation and characterization of adult human liver progenitors from ischemic liver tissue derived from therapeutic hepatectomies. Tissue Eng Part A. 2009;15:1633-1643
36. Jozefczuk J, Stachelscheid H, Chavez L, Herwig R, Lehrach H, Zeilinger K, Gerlach JC. et al. Molecular characterization of cultured adult human liver progenitor cells. Tissue Eng Part C Methods. 2010;16:821-834
37. Oertel M, Shafritz DA. Stem cells, cell transplantation and liver repopulation. Biochim Biophys Acta. 2008;1782:61-74
38. Navarro-Alvarez N, Soto-Gutierrez A, Kobayashi N. Stem cell research and therapy for liver disease. Curr Stem Cell Res Ther. 2009;4:141-146
39. Li J, Li L, Yu H, Cao H, Gao C, Gong Y. Growth and metabolism of human hepatocytes on biomodified collagen poly(lactic-co-glycolic acid) three-dimensional scaffold. ASAIO J. 2006;52:321-327
40. Herrera MB, Fonsato V, Gatti S, Deregibus MC, Sordi A, Cantarella D, Calogero R. et al. Human liver stem cell-derived microvesicles accelerate hepatic regeneration in hepatectomized rats. J Cell Mol Med. 2010;14:1605-1618
41. Arends B, Spee B, Schotanus BA, Roskams T, van den Ingh TS, Penning LC, Rothuizen J. In vitro differentiation of liver progenitor cells derived from healthy dog livers. Stem Cells Dev. 2009;18:351-358
42. Brooling JT, Campbell JS, Mitchell C, Yeoh GC, Fausto N. Differential regulation of rodent hepatocyte and oval cell proliferation by interferon gamma. Hepatology. 2005;41:906-915
43. Matsusaka S, Toyosaka A, Nakasho K, Tsujimura T, Sugihara A, Takanashi T, Uematsu K. et al. The role of oval cells in rat hepatocyte transplantation. Transplantation. 2000;70:441-446
Author contact
![]() Corresponding author: Lanjuan Li. Tel: 86-571-87236425; Fax: 86-571- 87236456 ; E-mail: ljliedu.cn
Corresponding author: Lanjuan Li. Tel: 86-571-87236425; Fax: 86-571- 87236456 ; E-mail: ljliedu.cn
Received 2013-8-16
Accepted 2013-12-11
Published 2013-12-21
