ISSN: 1449-1907International Journal of Medical Sciences
Int J Med Sci 2011; 8(2):161-167. doi:10.7150/ijms.8.161 This issue Cite
Short Research Communication
Anticancer Activity of the PR Domain of Tumor Suppressor RIZ1
1. Drug Design and Discovery Research Group, College of Pharmacy and Nutrition, University of Saskatchewan, 110 Science Place, Saskatoon, SK S7N 5C9, Canada
2. Vaccine and Infectious Disease Organization, University of Saskatchewan, 120 Veterinary Road, Saskatoon, SK S7N 5E3, Canada
3. Saskatchewan Structural Sciences Centre, University of Saskatchewan, 110 Science Place, Saskatoon, SK S7N 5C9, Canada
Abstract
Human tumor suppressor gene RIZ encodes two protein products, tumor suppressor RIZ1 and proto-oncoprotein RIZ2, which regulate cellular functions in a Yin-Yang fashion. The only structural difference between them is that RIZ2 lacks the N-terminal PR domain. In this study, we showed that RIZ1 mRNA expression level was elevated in stage IV of eight different types of cancer (stage III for prostate cancer), indicating that RIZ1 might play an important role in tumor metastasis, and the PR domain alone possessed anticancer activity.
Keywords: RIZ1, RIZ, Human tumor suppressor, tumor metastasis
Introduction
Yin-Yang regulation, which refers to dual complimentary opposite reactions, has been discovered in many biological and physiological regulations. Recently, human tumor suppressor gene RIZ was shown to function in a Yin-Yang fashion [1, 2]. RIZ, located on the distal short arm of chromosome 1 (1p36), encodes two different protein products, RIZ1 and RIZ2, using alternative promoters [1-10]. The expression levels for RIZ1 and RIZ2 are nearly the same among many different human tissues except for the testes [9]. RIZ2, a proto-oncoprotein, promotes cell division; whereas RIZ1, a tumor suppressor, arrests cells in the G2/M phase of the cell cycle and induces apoptosis [10-12]. An imbalance in the amount of RIZ1 and RIZ2 may be an important cause of cancer development [1]. In fact, silencing of RIZ1 expression, associated with increased RIZ2 expression, has been observed in various human cancers, such as hepatoma, leukemia, malignant lymphoma, breast cancer, colorectal cancer, and thyroid carcinoma [1, 4-8, 13, 14].
RIZ1 and RIZ2 share identical amino acid sequences except that RIZ2 lacks the N-terminal PR domain (PRDM2, ~200 amino acids) [9], which is a member of the PRDM (PRDI-BF1 and RIZ homology domain) family [15]. Thus, the PR domain is possibly responsible for the tumor suppressing activity of RIZ1. In vitro studies showed that PR interacted with a PRB (PR-binding) motif located in the C-terminal region of both RIZ1 and RIZ2, implicating the Yin-Yang fashion between RIZ1 and RIZ2 in cell regulations [6, 16, 17]. Seventeen types of PRDMs have been identified in humans [15, 18]. Most of them are located near the N-terminal portion of the proteins followed by zinc-fingers, and are involved in the regulation of cell division and differentiation [15, 18]. PRDM1, PRDM2, PRDM3, PRDM4, PRDM5, PRDM14 and PRDM16, which have been identified to be related to cancer, are also functional in a Yin-Yang fashion [2, 15, 18]. In addition, the PR domain was shown to possess H3K9 histone methyltransferase activity [19]. Since histone methylation has been proposed as an important epigenetic mechanism to suppress cancer, the histone methyltransferase activity of the PR domain may as well play a critical role for the tumor suppressing function of RIZ1 [15].
Despite recent rapid progress in RIZ1 studies, two important questions have not yet been addressed. The first question is whether the expression level for RIZ1 varies over the progression of cancer; and the second question is whether the PR domain alone possesses anticancer activity. In this study, we quantitatively analyzed the mRNA expression level of RIZ1 over the disease progression in eight different types of cancer, and evaluated the anticancer activity of the PR domain against human hepatoma HuH7 cells through both direct administration of recombinant His6-tagged PR and cDNA transfection. Our results showed that the PR domain alone exhibited anticancer activity.
Materials and Methods
Overexpression and purification of PR domain
The cloning, overexpression and purification of recombinant His6-tagget PR domain has been published previously [20].
Quantitative analysis of the RIZ1 mRNA expression level
The protocol used to analyze mRNA expression levels of selected target genes using Origene qPCR Cancer Survey Panels (Rockville, Maryland, USA) has been reported previously [21]. Briefly, the primers and the TaqMan probe for gene RIZ encoding RIZ1 were designed and synthesized by Applied Biosystems (Carlsbad, California, USA) based on the internally transcribed spacer (ITS) region. The TaqMan probe was labeled with FAM at 5'-end and non-fluorescent quencher at 3'-end. The RIZ1 mRNA expression levels were measured against the Origene TissueScan Cancer Survey Panel 96-I (twelve patients for each of the eight selected types of cancer) using quantitative RT-PCR on an Applied Biosystems 7300 Real-Time PCR System. The RIZ1 mRNA expression was averaged in each disease stage and normalized to an internal control, β-actin. The fold-difference in mRNA expression at each disease stage was determined by comparison to expression levels in normal patients (stage 0, expression level set as 1). Unpaired t-test with Welch's correction between the RIZ1 mRNA expression levels in normal and cancer patients for each cancer type was performed with GraphPad Prism 4.0 (GraphPad Software, San Diego, California, USA).
Cell culture of HuH7 cell line
Human hepatoma HuH7 were cultured in 6-well cell culture plates in Dulbecco's Modified Eagle's Medium with 10% fetal bovine serum and 1% gentamicin in a humidified, 5% CO2 atmosphere at 37°C. The cell cultural media were changed every 2-3 days. The HuH7 cells were subcultured using 0.25% trypsin, 0.53 mM EDTA solution before reaching 100% confluence.
Anticancer activity of PR domain by direct administration
All cell line experiments were undertaken in triplicate. The purified recombinant His6-tagged PR domain (> 90% purity) was directly administered into the cell cultural media with final concentrations of 1 μg/mL, 2 μg/mL, and 3 μg/mL, respectively, after the HuH7 cells reached 80-90% confluence. Tris-HCl buffer was used as the blank control. The cells were treated for 24 hr before the cell death rate was examined by the trypan blue method. The cells were stained by 0.01% trypan blue for 10 min and then examined under a microscope. At least 100 cells were counted for each treatment. Statistical analysis was performed using GraphPad InStat (GraphPad Software, San Diego, California, USA).
Anticancer activity of PR domain by cDNA transfection
The cloning of the PR domain (residues 13-193) has been reported previously [20]. Plasmid DNA harvested from positively transformed Escherichia coli DH5α cells was digested by restriction endonucleases BamHI and XhoI. The digested DNA fragment was subsequently sub-cloned into the expression vector pcDNA/HisMax C (Invitrogen, Burlington, Ontario, Canada) at the BamHI and XhoI sites. The constructed plasmid pcDNA/HisMax/PR vector was transfected into HuH7 cells (~ 1 million) by the calcium phosphate precipitation method [22]. Transfection with the empty pcDNA/HisMax C vector was used as a control. The expression of the PR domain in transfected HuH7 cells was confirmed by Western blot described below; and the cell death rate was examined by the trypan blue method after 24 and 48 hr of transfections, respectively.
Western blot of PR domain
The expression of the PR domain was examined after 24 hr of transfection with pcDNA/HisMax/PR and the empty vector, respectively. Supernatants (30 μg) of the cell lysates were separated by SDS-PAGE and transferred onto nitrocellulose membranes. The membranes were incubated with anti-Histidine tag or anti-β-actin antibodies overnight at 4 °C before being blotted with anti-rabbit IgG conjugated with horseradish peroxidase (1: 3000), for 1 hr at room temperature. The blotting results were scanned using a Li-Cor Odyssey scanner (Li-Cor, Lincoln, Nebraska, USA).
Results and Discussion
RIZ1 mRNA expression
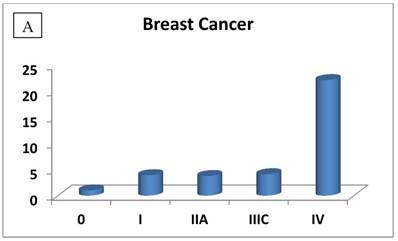
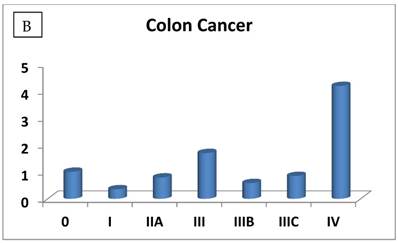
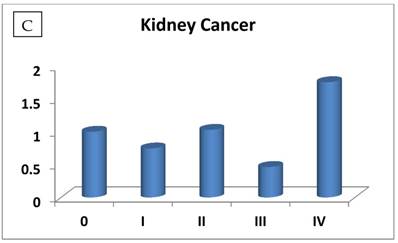
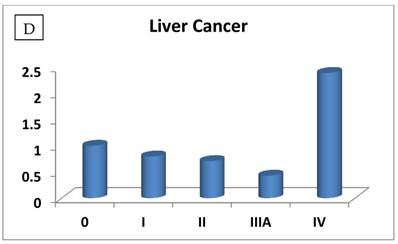
The RIZ1 mRNA expression level was elevated by an average of 3.1 fold in breast cancer (P=0.006) and decreased by 2 fold in thyroid cancer (P=0.028), respectively (Table 1). The change in RIZ1 mRNA expression was not statistically significant in colon, kidney, liver, lung, ovarian and prostate cancers. To obtain a preliminary impression on whether the expression of RIZ1 varies with cancer progression, we first compared the RIZ1 mRNA expression levels in late stages (III and IV combined) versus the early stages (I and II combined). No statistically significant variation was observed for any type of cancer (P value ranging from 0.35 to 0.94). Then, we compared the RIZ1 mRNA expression level at each individual cancer stage (Fig. 1), however, the small sample size associated with the panel (n=12 for each cancer type) precluded a complete analysis, one that must await a larger scale screening study. Nonetheless, we have gleaned valuable information from our limited analysis, which will provide an important reference for the design of future screenings.
The average fold difference (FD) in RIZ1 mRNA expression level in patients with cancer relative to patients without cancer
| Cancer Type | FD | P-value |
|---|---|---|
| Breast | 3.1 | 0.006 |
| Colon | 1.1 | 0.976 |
| Kidney | 0.6 | 0.070 |
| Liver | 0.8 | 0.384 |
| Lung | 1.3 | 0.768 |
| Ovary | 1.9 | 0.615 |
| Prostate | 0.7 | 0.351 |
| Thyroid | 0.5 | 0.028 |
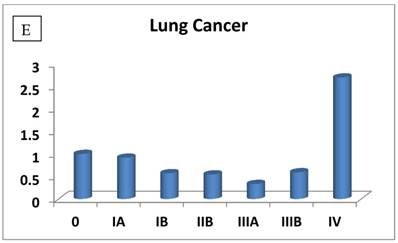
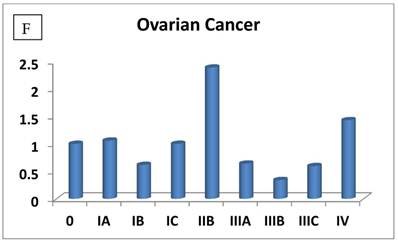
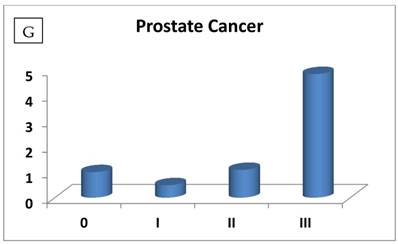
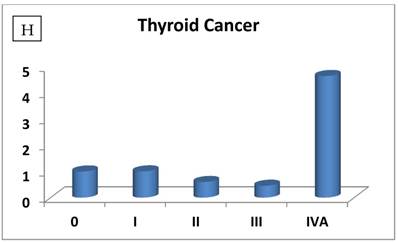
The most interesting discovery was that the RIZ1 mRNA expression was increased by 22.1, 4.2, 1.8, 2.4, 2.7, 1.4 and 4.7 folds at stage IV in breast, colon, kidney, liver, lung, ovarian and thyroid cancers, respectively, and 4.8 folds at stage III in prostate cancer (Fig. 1). Because cancer undergoes metastasis and spreads to other organs at late stages, we speculated that RIZ1 might play an important role in tumor metastasis, although increases in mRNA expression do not always translate proportionally into protein expression levels. Our speculation was in line with two previous observations [23, 24].
The first observation was that RIZ1 up-regulates insulin-like growth factor-binding protein 2 (IGFBP2) and secreted glycoprotein SPARC (secreted protein, acidic and rich in cysteine) [23], both of which are highly expressed in malignant tissues and promote tumor metastasis [25-28]. The second observation was that RIZ1 may augment the expression of nuclear factor of activated T cell 1 (NFATc1) [24], which induces breast cancer cell invasion via cyclooxygenase-2 [29]. Therefore, we hypothesize that RIZ1 might possess dual functions during tumor progression, acting as a tumor suppressor to induce apoptosis in the early stages and a tumor promoter to induce metastasis in late stages. If this hypothesis were proved valid, the net result of RIZ1's apparently conflicting function at early and late stages of cancer would be decreasing local cancer cell population, an apparently unsuccessful self-protective mechanism of human body in battling against cancer. Further characterization including metastatic studies is needed to prove this hypothesis.
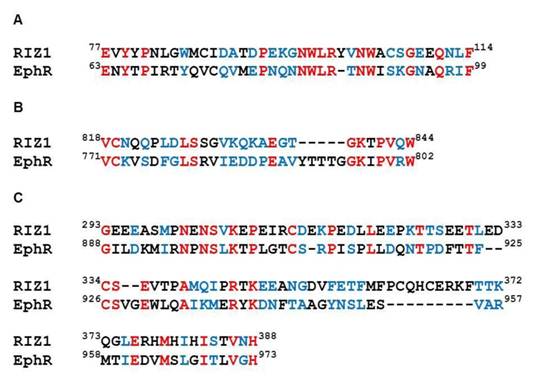
Moreover, we observed amino acid sequence similarities in three regions between RIZ1 and ephrin receptor (EphR) during sequence analysis (Fig. 2). The sequence identity and homology were 34.2% and 68.4%, 37.0% and 81.5%, and 22.9% and 53.1% in the three aligned regions, respectively (Fig. 2). In EphR, residues 63-99 are located in a region responsible for extracellular ligand binding, residues 771-802 are located in a region responsible for intracellular tyrosine kinase activity, and residues 924-973 are located in an essential motif for dimerization of receptor tyrosine kinases [30]. Since EphR is often overexpressed in cancer and involved in metastasis [31, 32], it is definitely worth further exploration on whether overexpression of RIZ1 could produce any effect on the ephrin signalling pathways.
Relative fold-difference in RIZ1 mRNA expression at different disease stages of human breast (A), colon (B), kidney (C), liver (D), lung (E), ovary (F), prostate (G) and thyroid (H) cancers. RIZ1 mRNA expression was screened in the Origene TissueScan Cancer Survey Panel 96-I and normalized to the internal control, β-actin in the different patients. The fold-difference in mRNA expression at each disease stage was determined by comparison to expression levels in normal patients (stage 0, expression level set as 1).
Amino acid sequence alignment between RIZ1 and ephrin receptor (EphR) in the extracellular ligand binding region, residues 63-99 (A), the intracellular tyrosine kinase region, residues 771-802 (B), and the essential motif for dimerization of receptor tyrosine kinases, residues 924-973 (C). Identical and homologous residues are shown in red and blue, respectively. The sequence identity and homology between RIZ1 and EphR were 34.2% and 68.4% in the residues 63-99 region, 37.0% and 81.5% in the residues 771-802 region, and 22.9% and 53.1% in the residues 924-973 region, respectively.
The second interesting observation in this study was that the RIZ1 mRNA expression is elevated in all stages through the progression of breast cancer (Fig. 1A). This is in contrary to the previous studies that RIZ1 was under-expressed in breast cancer [4]. However, the current result is consistent with a study on cancer stem cells that only 11-35% breast cancer cells sustained tumor growth through cell division [33], implicating that RIZ1 was likely overexpressed and acted as a tumor suppressor to inhibit breast cancer cell growth during disease progression. As for the other types of cancer, the RIZ1 mRNA level was down-regulated from stage I to III (except prostate cancer), which is consistent with the previous studies [1, 5-8]. Since either stage III of colon cancer or stage IIB of ovarian cancer has only one elderly patient (80 years and older), the RIZ1 mRNA expression increase may not be necessarily representative for the disease stage.
Anticancer activity of PR Domain
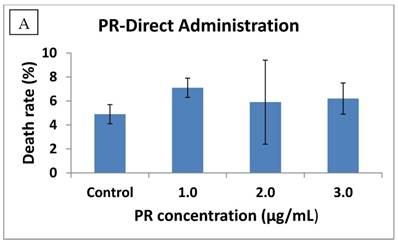
Transfection with full-length RIZ1 has been shown to suppress cancer cell growth in hepatoma and chronic myeloid leukemia [11, 23]. Since PR domain is the only structural difference between the two protein products (RIZ1 and RIZ2) of gene RIZ, we decided to examine whether the PR domain alone possessed any anticancer activity. Initially, we added purified His6-tagged PR domain (>90% purity) directly to the cultural media for human hepatoma HuH7 cells. The PR domain slightly increased the cell death rate for HuH7 cells at all three tested PR domain concentrations (Fig. 3A). Statistical significance was observed for the 1 µg/mL treatment (P=0.030), however, the increase in cell death rate was only marginal (< 2.5%). This indicated that it was unlikely that a receptor or transporter is present on the HuH7 cell membrane to actively mediate the translocation of PR domain into the cells.
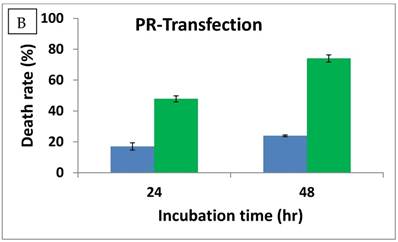
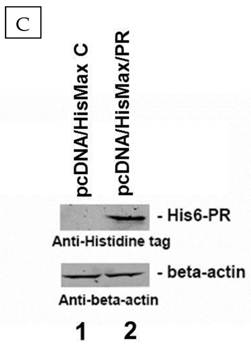
To ensure the presence of PR domain within the cells, we transfected HuH7 cells with a plasmid (pcDNA/HisMax/PR) expressing the PR domain. The PR domain expression was examined by Western blot after 24 hr of transfection using β-actin as a loading control. As shown in Fig. 3C, the PR domain was expressed in the HuH7 cells transfected with the pcDNA/HisMax/PR vector but not in the cells transfected with the empty pcDNA/HisMax C vector. The cell death rate for HuH7 cells was measured at 24 and 48 hr, respectively, after transfection. As shown in Fig. 3B, the cell death rate was increased from 17% to 48% after 24 hr of transfection and from 24% to 74% after 48 hr of transfection. Because either recombinant His6-tagged or GST-tagged PR domain is well folded in both prokaryotic Escherichia coli [20] and eukaryotic yeast cells (unpublished data), it is unlikely that the increased cell death was due to improper folding of RIZ1 in the tranfected HuH7 cells. Thus, the above observation suggested that the PR domain alone possesses anticancer activity. However, it is still unclear whether the anticancer activity of the PR domain is due to its methyltransferase activity or the interaction with the PRB motif. Further studies with a methyltransferase inhibitor specific for the PR domain may help to understand the exact mechanism for its anticancer function.
Anticancer activity of the PR domain against human hepatocarcinoma HuH7 cells either by direct admission (A) or transfection (B). The PR domain concentrations were 1 μg/mL, 2 μg/mL, and 3 μg/mL, respectively, in the direct administration. The cell death rate was measured by the trypan blue method. For the transfection, the HuH7 cells were transfected with the constructed viral vector pcDNA/HisMax/PR encoding the PR domain (shown in green) and the empty viral vector pcDNA/HisMax as a control (shown in blue). The cell death rate was also measured by the trypan blue method after 24 and 48 hr of transfection, respectively. The PR domain expression in the transfected HuH7 cells was confirmed by Western blot after 24 hr of transfection (C).
Conclusion
In this study, RIZ1 mRNA expression was shown to be up-regulated in stage IV of eight types of cancer (stage III in prostate cancer), indicating that RIZ1 might play an important role in tumor metastasis in late disease stages in addition to the previously reported tumor suppressor activity. The PR domain of RIZ1 possessed anticancer activity from the transfection studies. This finding leaves us an important question on the roles the other structural components of RIZ1 in its anticancer function.
Acknowledgements
This works was supported from a research grant from the Cancer Research Society of Canada. W. Sun would also like to acknowledge the College of Pharmacy and Nutrition, University of Saskatchewan for the awarding of a graduate scholarship.
Conflict of Interest
The authors declare that no conflict of interest exists.
References
1. Jiang G, Liu L, Buyse IM, Simon D, Huang S. Decreased RIZ1 expression but not RIZ2 in hepatoma and suppression of hepatoma tumorigenicity by RIZ1. Int J Cancer. 1999;83:541-6
2. Jiang GL, Huang S. The yin-yang of PR-domain family genes in tumorigenesis. Histol Histopathol. 2000;15:109-17
3. Buyse IM, Takahashi EI, Huang S. Physical mapping of the retinoblastoma interacting zinc finger gene RIZ to D1S228 on chromosome 1p36. Genomics. 1996;34:119-21
4. He L, Yu JX, Liu L, Buyse IM, Wang MS, Yang QC, Nakagawara A, Brodeur GM, Shi YE, Huang S. RIZ1, but not the alternative RIZ2 product of the same gene, is underexpressed in breast cancer, and forced RIZ1 expression causes G2-M cell cycle arrest and/or apoptosis. Cancer Res. 1998;58:4238-44
5. Chadwick RB, Jiang GL, Bennington GA, Yuan B, Johnson CK, Stevens MW, Niemann TH, Peltomaki P, Huang S, de la Chapelle A. Candidate tumor suppressor RIZ is frequently involved in colorectal carcinogenesis. Proc Natl Acad Sci USA. 2000;97:2662-7
6. Sasaki O, Meguro K, Tohmiya Y, Funato T, Shibahara S, Sasaki T. Altered expression of retinoblastoma protein-interacting zinc finger gene, RIZ, in human leukaemia. Br J Haematol. 2002;119:940-8
7. Lal G, Padmanabha L, Smith BJ, Nicholson RM, Howe JR, O'Dorisio MS, Domann FE Jr. RIZ1 is epigenetically inactivated by promoter hypermethylation in thyroid carcinoma. Cancer. 2006;107:2752-9
8. Piao GH, Piao WH, He Y, Zhang HH, Wang GQ, Piao Z. Hyper-methylation of RIZ1 tumor suppressor gene is involved in the early tumorigenesis of hepatocellular carcinoma. Histol Histopathol. 2008;23:1171-5
9. Liu L, Shao G, Steele-Perkins G, Huang S. The retinoblastoma interacting zinc finger gene RIZ produces a PR domain-lacking product through an internal promoter. J Biol Chem. 1997;272:2984-91
10. Du Y, Carling T, Fang W, Piao Z, Sheu JC, Huang S. Hypermethylation in human cancers of the RIZ1 tumor suppressor gene, a member of a histone/protein methyltransferase superfamily. Cancer Res. 2001;61:8094-9
11. Jiang GL, Huang S. Adenovirus expressing RIZ1 in tumor suppressor gene therapy of microsatellite-unstable colorectal cancers. Cancer Res. 2001;61:1796-8
12. Deng Q, Huang S. PRDM5 is silenced in human cancers and has growth suppressive activities. Oncogene. 2004;17:4903-10
13. Mori N, Morosetti R, Spira S, Lee S, Ben-Yehuda D, Schiller G, Landolfi R, Mizoguchi H, Koeffler HP. Chromosome band 1p36 contains a putative tumor suppressor gene important in the evolution of chronic myelocytic leukemia. Blood. 1998;92:3405-9
14. Tam W, Gomez MF, Chadburn A, Knowles DM, Houldsworth J. Lack of A563G (I188V) missense mutation in RIZ/ PR in human diffuse large B-cell lymphomas. Genes Chromosomes Cancer. 2007;46:416-8
15. Kim KC, Huang S. Histone methyltransferases in tumor suppression. Cancer Biol Ther. 2003;2:491-9
16. Huang S, Shao G, Liu L. The PR domain of the Rb-binding zinc finger protein RIZ1 is a protein binding interface and is related to the SET domain functioning in chromatin-mediated gene expression. J Biol Chem. 1998;273:15933-9
17. Canote R, Du Y, Carling T, Tian F, Peng Z, Huang S. The tumor suppressor gene RIZ in cancer gene therapy (review). Oncol Rep. 2002;9:57-60
18. Watanabe Y, Toyota M, Kondo Y, Suzuki H, Imai T, Ohe-Toyota M, Maruyama R, Nojima M, Sasaki Y, Sekido Y, Hiratsuka H, Shinomura Y, Imai K, Itoh F, Tokino T. PRDM5 identified as a target of epigenetic silencing in colorectal and gastric cancer. Clin Cancer Res. 2007;13:4786-94
19. Derunes C, Briknarová K, Geng L, Li S, Gessner CR, Hewitt K, Wu S, Huang S, Woods VI Jr, Ely KR. Characterization of the PR domain of RIZ1 histone methyltransferase. Biochem Biophys Res Commun. 2005;333:925-34
20. Sun W, Geyer CR, Yang J. Cloning, expression, purification and crystallization of the PR domain of human retinoblastoma protein-binding zinc finger protein 1 (RIZ1). Int J Mol Sci. 2008;9:943-50
21. Chen L, Ling B, Alcorn J, Yang J. Analysis of the expression of human N-myristoyltransferase 1 (hNMT-1) in cancers. Open Biomarkers J. 2009;2:6-10
22. Graham FL, van der Eb AJ. A new technique for the assay of infectivity of human adenovirus 5 DNA. Virology. 1973;52:456-67
23. Pastural E, Takahashi N, Dong WF, Bainbridge M, Hull A, Pearson D, Huang S, Lowsky R, DeCoteau JF, Geyer CR. RIZ1 repression is associated with insulin-like growth factor-1 signaling activation in chronic myeloid leukemia cell lines. Oncogene. 2007;26:1586-94
24. Noman AS, Koide N, Iftakhar-E-Khuda I, Dagvadorj J, Tumurkhuu G, Naiki Y, Komatsu T, Yoshida T, Yokochi T. Retinoblastoma protein-interacting zinc finger 1 (RIZ1) participates in RANKL-induced osteoclast formation via regulation of NFATc1 expression. Immunol Lett. 2010;131:166-9
25. Miyake H, Hara I, Yamanaka K, Muramaki M, Gleave M, Eto H. Introduction of insulin-like growth factor binding protein-2 gene into human bladder cancer cells enhances their metastatic potential. Oncol Rep. 2005;13:341-5
26. Wang H, Arun BK, Wang H, Fuller GN, Zhang W, Middleton LP, Sahin AA. IGFBP2 and IGFBP5 overexpression correlates with the lymph node metastasis in T1 breast carcinomas. Breast J. 2008;14:261-7
27. Porte H, Chastre E, Prevot S, Nordlinger B, Empereur S, Basset P, Chambon P, Gespach C. Neoplastic progression of human colorectal cancer is associated with overexpression of the stromelysin-3 and BM-40/SPARC genes. Int J Cancer. 1995;64:70-5
28. Wang CS, Lin KH, Chen SL, Chan YF, Hsueh S. Overexpression of SPARC gene in human gastric carcinoma and its clinic-pathologic significance. Br J Cancer. 2004;91:1924-30
29. Yiu GK, Toker A. NFAT induces breast cancer cell invasion by promoting the induction of cyclooxygenase-2. J Biol Chem. 2006;281:12210-7
30. Stapleton D, Balan I, Pawson T, Sicheri F. The crystal structure of an Eph receptor SAM domain reveals a mechanism for modular dimerization. Nat Struct Biol. 1999;6:44-9
31. Brantley-Sieders DM, Fang WB, Hwang Y, Hicks D, Chen J. Ephrin-A1 facilitates mammary tumor metastasis through an angiogenesis-dependent mechanism mediated by EphA receptor and vascular endothelial growth factor in mice. Cancer Res. 2006;66:10315-24
32. Pasquale EB. Eph receptors and ephrins in cancer: bidirectional signalling and beyond. Nat Rev Cancer. 2010;10:165-80
33. Dalerba P, Cho RW, Clarke MF. Cancer stem cells: models and concepts. Annu Rev Med. 2007;58:267-84
Author contact
![]() Corresponding author: Jian Yang, Drug Design and Discovery Research Group, College of Pharmacy and Nutrition, University of Saskatchewan, 110 Science Place, Saskatoon, SK S7N 5C9, Canada. Tel: 306-966-6361; Fax: 306-966-6377; E-mail: jian.yangca
Corresponding author: Jian Yang, Drug Design and Discovery Research Group, College of Pharmacy and Nutrition, University of Saskatchewan, 110 Science Place, Saskatoon, SK S7N 5C9, Canada. Tel: 306-966-6361; Fax: 306-966-6377; E-mail: jian.yangca
Received 2010-9-3
Accepted 2011-2-15
Published 2011-2-21
